 |
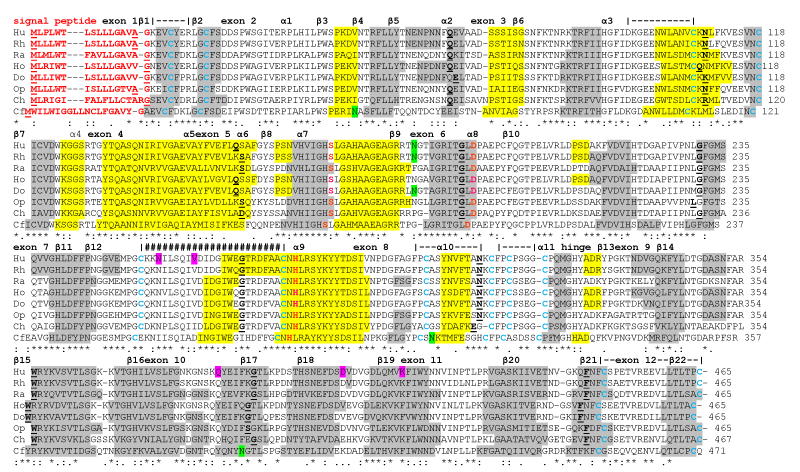
| Figure 1: Amino Acid Sequence Alignments for Vertebrate Pancreatic Lipase (PTL) Sequences. See Table 1 for sources of vertebrate PTL sequences; * shows identical residues for proteins; : one similar alternate residue; . two similar alternate residues; key PTL active site residues: Ser169, Asp193 and His280 (human PTL); N-signal peptide residues are in red; N-glycosylation residues are in shaded green; five disulphide bonds are represented as |----| with Cys residues in shaded blue; β-sheets (β1-β22) are numbered according to human PTL sequences [12] in shaded grey; α-helices (α1-10) are also numbered according to the human PTL structure and are in shaded yellow; |####| represents ‘lid’ region covering active site; colipase binding residues are in purple; hinge refers to region connecting the ‘lipase’ and ‘plat’ domains; bold underlined font shows residues corresponding to known or predicted exon start sites; exon numbers refer to human PTL gene coding exons. |