 |
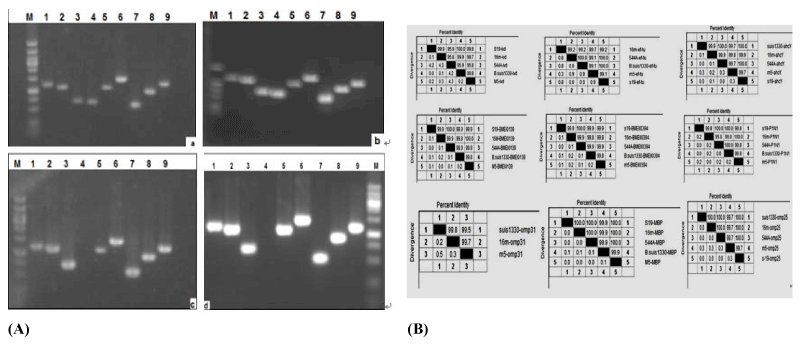
| Figure 3: PCR amplification and sequence distance analysis on the genes encoding identified proteins. A: The immunodominant proteins identified by LC-MS/MS were amplificated by PCR from Brucella melitensis 16M strain (a), M5 strain (b); Brucella abortus 544A strain (c) and S19 strain (d). M, DNA Marker; 1, maltose binding protein; 2, isovalery-CoA dehydrogenase; 3, 25kDa-outer membrane immunogenic protein precursor; 4, 31kDa-outer membrane immunogenic protein precursor; 5, glycerol trinitrate reductase; 6, succinyl-CoA synthetase subunit alpha; 7, peptidyl-prolyl cis-trans isomerase A; 8,S-adenosyl-L-homocysteine hydrolase; 9, Protein translation elongation factor EF-TU. B: All the PCR products of immunodominant proteins were inserted into PMD-18T vector. Following sequencing, Sequence Distances among these sequences were analyzed through DNAMAN, and the gene homologies ranging from 95%-99% among five Brucella strains. |