 |
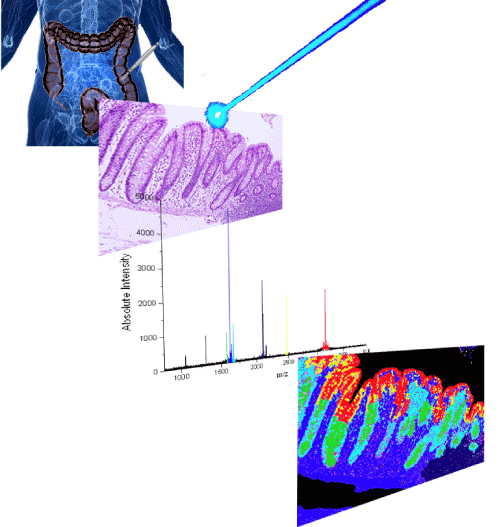
| Figure 4: Mass Spectrometric Imaging. The traditional application follows the use of tissue samples, formalin fixed, and paraffin embedded, or frozen. These samples are then sliced (3-10mm thick) and stained with haematoxylin and eosin. These tissue slices then have Matrix applied to them to help ionisation of proteins (can also be carried out without this step). The sample is shot with a laser and the ionised proteins measured using the traditional Matrix Assisted Laser Desorption Ionisation (MALDI) method. The ion intensity information is then converted into colour information and this is computationally overlaid on the image of the original sample, therefore generating a visual representation of protein localisation. |