 |
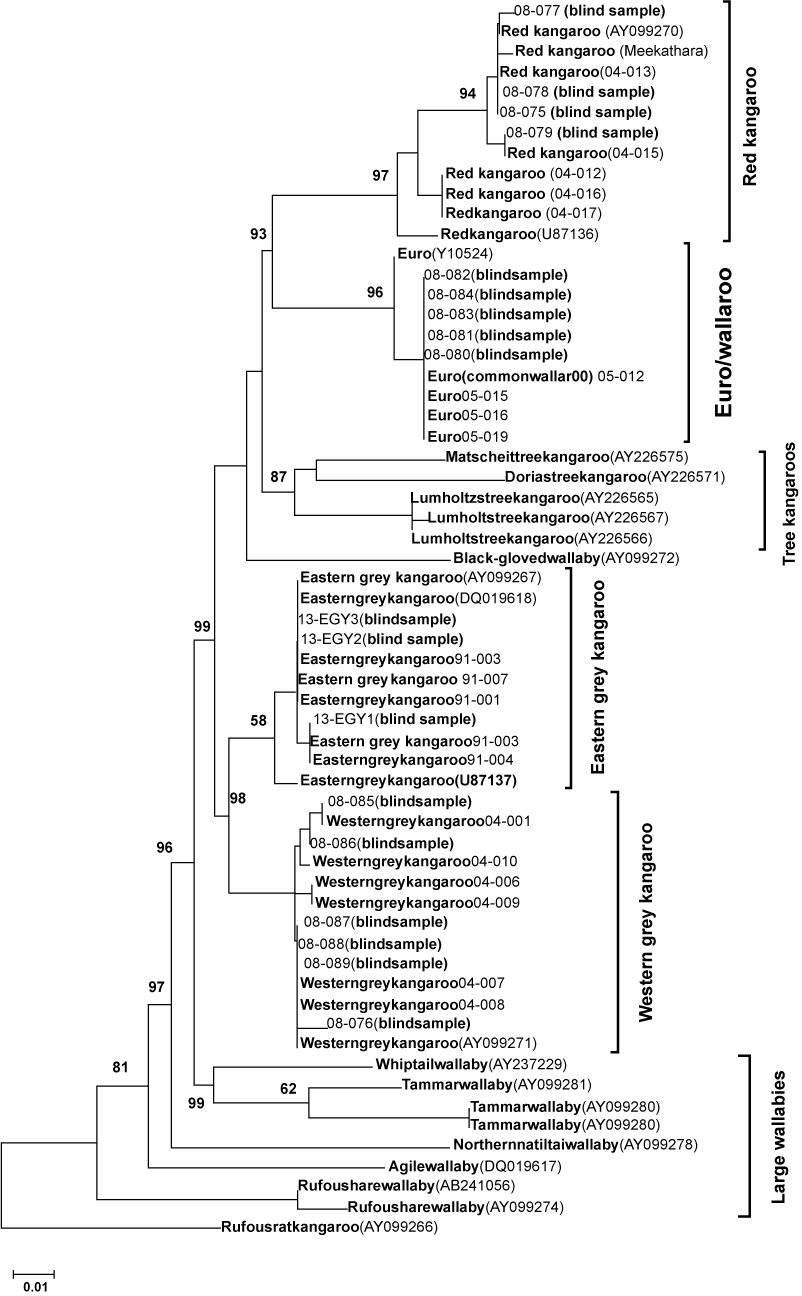
| Figure 1: Relationships of 61specimensfrom kangaroos and wallabies that would be considered suitably large to take for the commercial industry. The phylogeny was inferred using the Neighbor-Joining method [10]. The optimal tree with the sum of branch length = 0.7499 is shown. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (1000 replicates) are shown next to the branches [21]. The tree is drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. The evolutionary distances were computed using the Maximum Composite Likelihood method [10] and are in the units of the number of base substitutions per site. All positions containing gaps and missing data were eliminated from the dataset (complete deletion option). There were a total of 249 positions in the final data. Numbers in brackets are from GenBank and all other numbers are from our laboratory. |