Indexed In
- Open J Gate
- Genamics JournalSeek
- Academic Keys
- JournalTOCs
- China National Knowledge Infrastructure (CNKI)
- Ulrich's Periodicals Directory
- RefSeek
- Hamdard University
- EBSCO A-Z
- Directory of Abstract Indexing for Journals
- OCLC- WorldCat
- Publons
- Geneva Foundation for Medical Education and Research
- Euro Pub
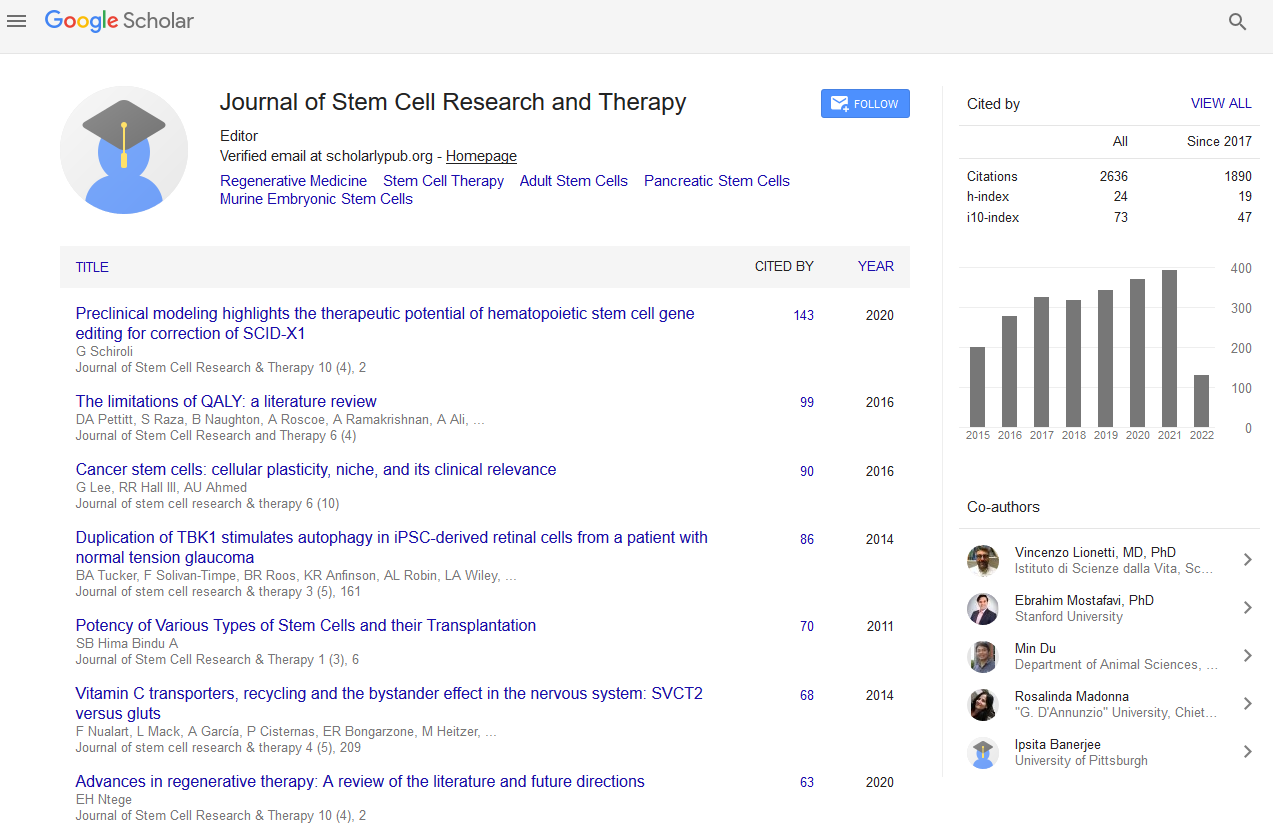
- Google Scholar
Useful Links
Share This Page
Journal Flyer

Open Access Journals
- Agri and Aquaculture
- Biochemistry
- Bioinformatics & Systems Biology
- Business & Management
- Chemistry
- Clinical Sciences
- Engineering
- Food & Nutrition
- General Science
- Genetics & Molecular Biology
- Immunology & Microbiology
- Medical Sciences
- Neuroscience & Psychology
- Nursing & Health Care
- Pharmaceutical Sciences
Abstract
Comparative Gene Expression Profiling of Stromal Cell Matrices that Support Expansion of Hematopoietic Stem/Progenitor Cells
Abhilasha Tiwari, Christophe Lefevre, Mark A Kirkland, Kevin Nicholas and Gopal Pande
The bone marrow microenvironment maintains a stable balance between self-renewal and differentiation of hematopoietic stem/progenitor cells (HSPCs). This microenvironment, also termed the “hematopoietic niche”, is primarily composed of stromal cells and their extracellular matrices (ECM) that jointly regulate HSPC functions. Previously, we have demonstrated that umbilical cord blood derived HSPCs can be maintained and expanded on stromal cell derived cellular matrices that mimic the complexity of the hematopoietic niche. The results indicated that matrices prepared at 20% O2 with osteogenic medium (OGM) were best suited for expanding committed HSPCs, whereas, matrices prepared at 5% O2 without OGM were better for primitive progenitors. Based upon these results we proposed that individual constituents of these matrices could be responsible for regulation of specific HSPC functions. To explore this hypothesis, we have performed comparative transcriptome profiling of these matrix producing cells, which identified differential expression of both known niche regulators, such as Wnt4, Angpt2, Vcam and Cxcl12, as well as genes not previously associated with HSPC regulation, such as Depp. MetaCore analysis of the differentially expressed genes suggests the down-regulation of several ECM related pathways and up-regulation of Ang-Tie2 and Wnt signaling pathways in OGM under high O2 (20%). Our findings provide an overview of several known and unique genes and pathways that play potential key roles in the support of HSPCs by stromal cells, both ex vivo and in vivo, and could be helpful in understanding the complex network of signaling and communication in hematopoietic niches.