Indexed In
- Open J Gate
- Genamics JournalSeek
- CiteFactor
- Cosmos IF
- Scimago
- Ulrich's Periodicals Directory
- Electronic Journals Library
- RefSeek
- Hamdard University
- EBSCO A-Z
- Directory of Abstract Indexing for Journals
- OCLC- WorldCat
- Proquest Summons
- Scholarsteer
- ROAD
- Virtual Library of Biology (vifabio)
- Publons
- Geneva Foundation for Medical Education and Research
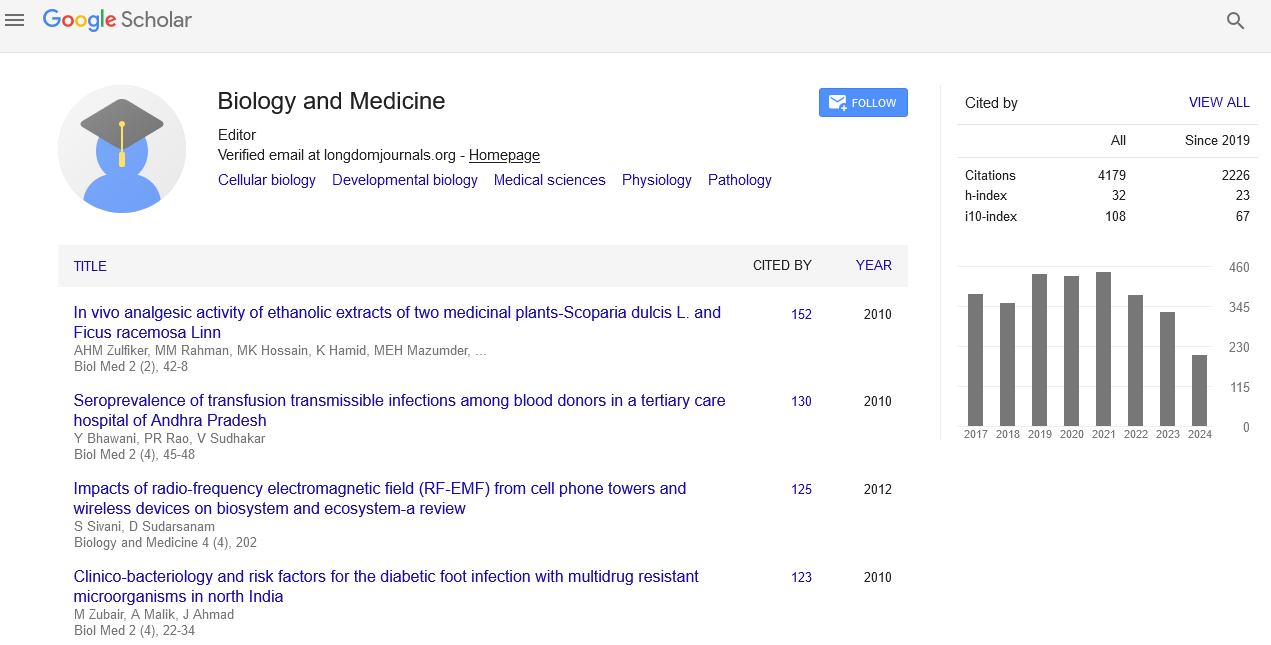
- Google Scholar
Useful Links
Share This Page
Journal Flyer

Open Access Journals
- Agri and Aquaculture
- Biochemistry
- Bioinformatics & Systems Biology
- Business & Management
- Chemistry
- Clinical Sciences
- Engineering
- Food & Nutrition
- General Science
- Genetics & Molecular Biology
- Immunology & Microbiology
- Medical Sciences
- Neuroscience & Psychology
- Nursing & Health Care
- Pharmaceutical Sciences
Abstract
Phylogeny reconstrution of ubiquitin conjugating (E2) enzymes
Muhummadh Khan, Kaiser Jamil
Phylogeny reconstruction has attracted much attention from biologists and computer scientists from gene-order dataover the last few years. Ubiquitin is widespread in eukaryotes and plays a pivotal role in selective ATP dependent protein degradation in cells. It is also highly expressed in several types of cancers. Hence, in order to understand its diversification in eukarya we undertook this study to determine its phylogeny using various computational methods. All sequences of Ubiquitin conjugating enzyme were extracted from protein data bank (PDB), and using MaximumLikelihood method as implemented in PHYML and the unrooted phylogenetic tree was constructed. This tree had four major clusters and one mini cluster. We calculated the divergence for the clusters of the gene from the site-specific change of functional importance in protein sequence evolution. The divergence was then mapped on the 3-D structure of the ube2c protein obtained from the PDB Data bank: (IL7K) using RASMOL. From this study, we conclude that we have been able to point out the exact sites where active evolution of E2s is taking place and it is apparent that these sites are subject to a strong purifying selection. We anticipate that this information would have some useful implications in neoplasia as there are reports that mutations in this protein are likely to promote tumour progression.