Indexed In
- Open J Gate
- Genamics JournalSeek
- CiteFactor
- Cosmos IF
- Scimago
- Ulrich's Periodicals Directory
- Electronic Journals Library
- RefSeek
- Hamdard University
- EBSCO A-Z
- Directory of Abstract Indexing for Journals
- OCLC- WorldCat
- Proquest Summons
- Scholarsteer
- ROAD
- Virtual Library of Biology (vifabio)
- Publons
- Geneva Foundation for Medical Education and Research
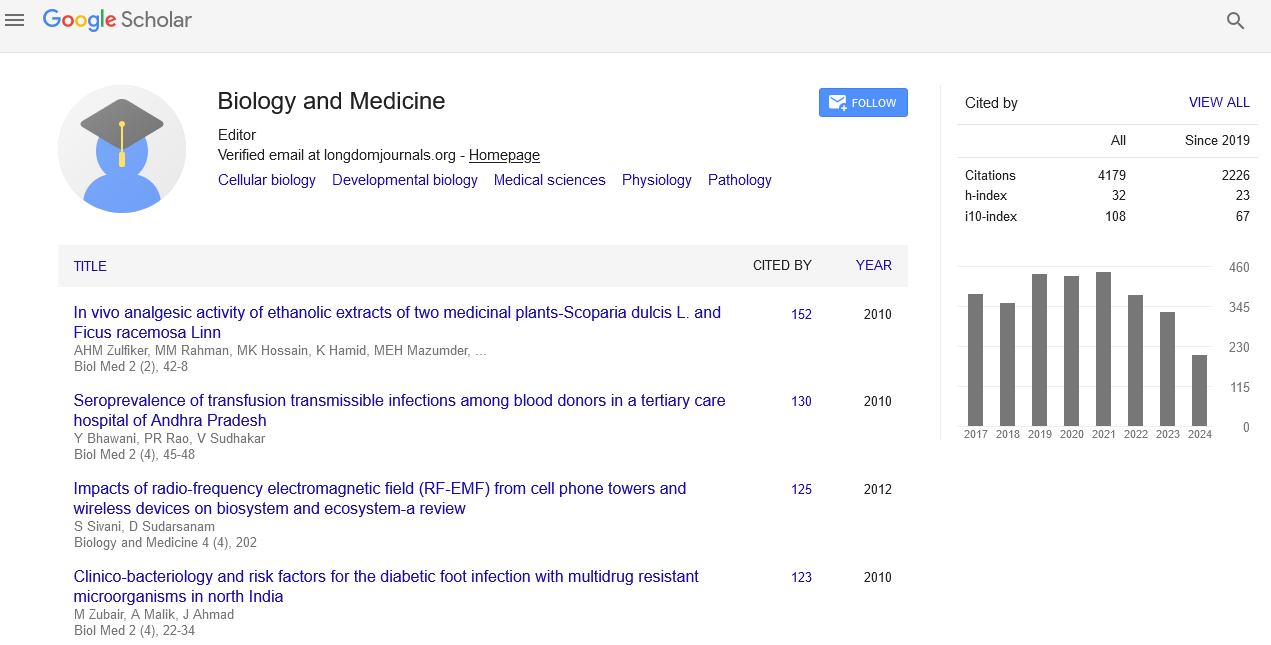
- Google Scholar
Useful Links
Share This Page
Journal Flyer

Open Access Journals
- Agri and Aquaculture
- Biochemistry
- Bioinformatics & Systems Biology
- Business & Management
- Chemistry
- Clinical Sciences
- Engineering
- Food & Nutrition
- General Science
- Genetics & Molecular Biology
- Immunology & Microbiology
- Medical Sciences
- Neuroscience & Psychology
- Nursing & Health Care
- Pharmaceutical Sciences
Abstract
Role of MicroRNA Molecules in Colon Cancer Etiology
Farid E Ahmed
The biogenesis of micro (mi) RNA suggests that nearly 3% of human genes are encoded for micro (mi) RNAs, and computer predictions indicate that more than 30% of protein coding genes in humans are coded by miRNAs through an imperfect binding to the 3’ untranslated region (UTR) of target messenger (m) RNA affecting gene silencing and leading to either transcription repression, or induction of messenger (m) RNA degradation. The expression of several miRNAs in noninvasive body fluids/excrements has been linked to development of colorectal cancer (CRC) and its progression. The majority of miRNA genes are oriented antisense to neighboring genes, and miRNA genes are transcribed by polymerases II and III. The rate of evolution of miRNAs has been very slow, which has permitted morphological innovation by making gene expression specific, a process that permitted the genesis of complex organisms. Bacteria lack true miRNAs. Bioinformatics approaches to predict putative miRNA target genes has been facilitated by finding that miRNA target recognition is partly based on simple sequence complementarity, and exact base pairing between miRNAs and their targets is required only in the first six to eight bases from the 5′ end of the miRNA.