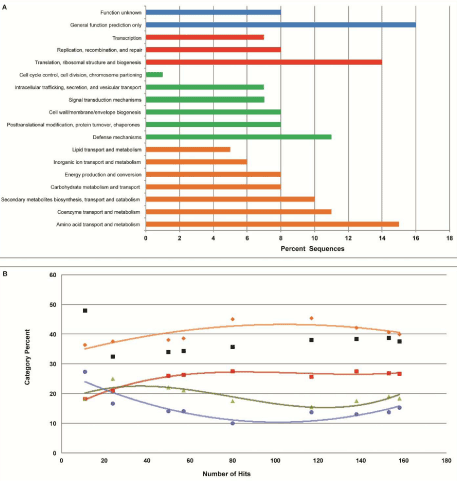
(A) Major COG categories consisted of color-coded sub categories: (Blue) Poorly Characterized, (Red) Information Storage and Processing, (Green) Cellular Processes and Signaling, and (Orange) Metabolism. Parameters for sequence comparison were set using the recommended limits of 1e-5 e-Value cutoff, a minimum identity of 60 percent, and a minimum alignment length of 15 base pairs.
(B) Change in percent of each major COG category as fosmid sequence hits increased. At each point, the sequences were classified using the same COG categories in figure 2A. As an increasing number of clones were sequenced, each category approaches saturation. The black data points represent the percentage of hits relative to the number of sequences for each respective x coordinate.