 |
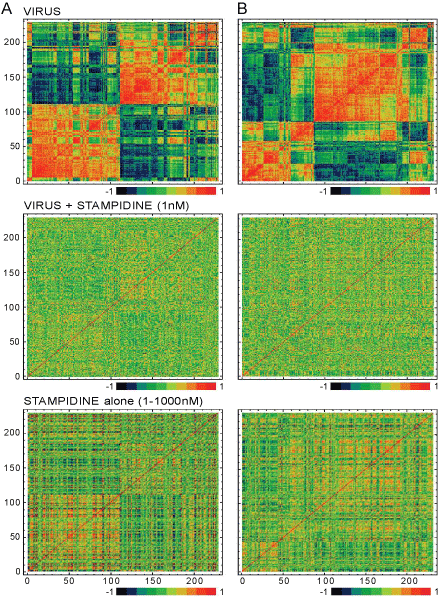
| Figure 2: Modular structures of co-regulated genes in HIV-infected cells are disrupted by Stampidine. Analysis of variance was performed to identify gene transcripts in MT-2 cells that were dynamically regulated across three time points (24 hr, 48 hr and 7 days) computed as change in signal value relative to zero-time point for each of the 3 time points in the presence of HIV alone (upper panels) (N=11) or HIV plus 1nM Stampidine (middle panels; Stampidine was added to cell cultures at the same time as HIV)) (N=17). We also examined the effects of Stampidine alone without HIV infection. MT-2 cells were treated with 0 nM (N=8) or 1-1000 nM (N=17) Stampidine for 24hrs, 48 hrs, or 7 days (lower panels) (N = 17) for two functional groups of RNA transcripts. [A]. Transcription factors (229 gene transcripts). [B]. Receptor/ Signal Transduction (232 gene transcripts). Correlation coefficients (r) were determined for each gene pair (shown on x and y axis, which refers to the gene number) and hierarchical cluster analysis (Complete Linkage, Pearson correlation as Distance metric) was used to characterize the network properties of the functional classes of genes treated with virus in the absence (upper panels) or presence (middle panels) of Stampidine. The orders of the genes are the same on both axis and a correlation value of 1 is shown on the diagonal. The correlation matrix between genes shows a network in which few sets of genes are strongly connected within modules and sparsely connected between modules. We examined interconnectedness between 3 subsets of genes in Module 1 (Gene numbers 7-37), Module 2 (Gene numbers 102- 106) and Module 3 (217-226) that contained 4 genes (SMARCD2, CALR, SSA2 and TAF6) with the most significant changes in gene expression at 2 or more time points and exhibited the largest number of connections with other transcription factor genes. The correlation coefficients are color coded blue for negative correlation coefficient values and red for positive values in the graphical representation of the clustering. |