 |
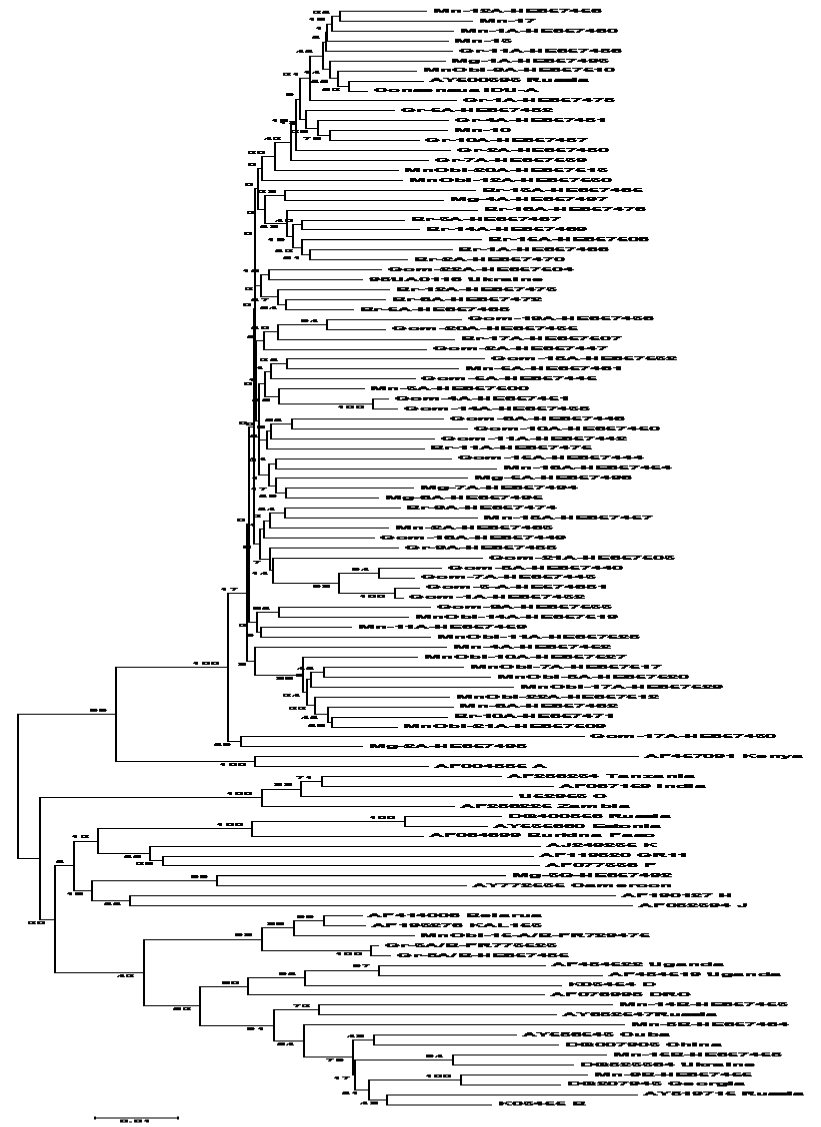
| Figure 4: Phylogenetic analysis of HIV-1 gene pol sequences from a newly diagnosed HIV-infected antiretroviral naïve patients isolates. Reference sequences of HIV-1 and viruses specific for the epidemic in the former Soviet Union and Belarus as well as A-K subtypes are included. Reference sequences of HIV-1 subtypes are labeled by their GenBank accession numbers. For IDU-A viruses, the consensus of viruses from Svetlogorsk1 is used as a reference. The neighbor-joining tree were built by using the Kimura two-parameter distance estimation method and pairwise gap deletion. Bootstrap analysis was done with 100 replications, values above 70 are shown. |