 |
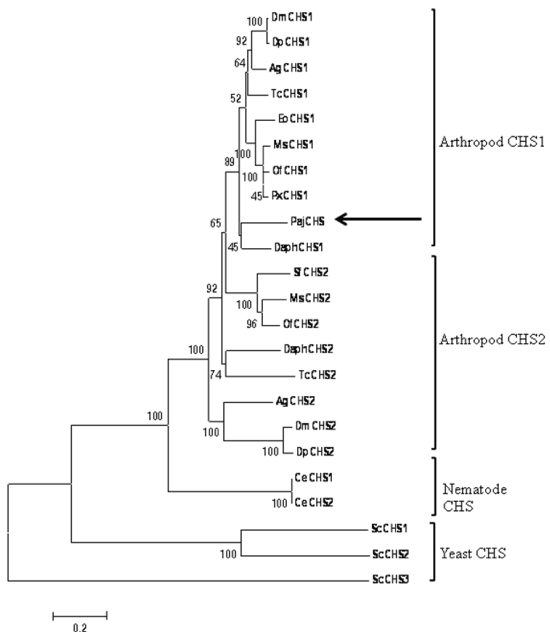
| Figure 2: Phylogenetic tree of CHSs from various species. The Phylogenetic tree was constructed by minimum evolution algorithm using Molecular Evolutionary Genetics Analysis (MEGA4) software (see Materials and methods). Bootstrapping test was performed with 1000 replicates. The scale bar represents 0.2 amino acid substitution per site. Species and GenBank accession numbers: P. japonica, PajCHS (KC131026), D. melanogaster, DmCHS1 (NM_079509), DmCHS2 (NM_079485); D. pseudoobscura, DpCHS1 (XP_001359390), DpCHS2 (XP_001352881); A. gambiae, AgCHS1 (XM_321337), AgCHS2 (AY056833); T. castaneum, TcCHS1AAQ55059), TcCHS2 (AAQ55061); E. obliqua, EoCHS1 (ACA50098), P. xylostella, PxCHS1 (BAF47974), M. sexta, MsCHS1 (AY062175), MsCHS2 (AY821560); S. frugiperda, SfCHS2 (AAS12599); Ostriniafurnacalis OfCHS1 (ACB13821), OfCHS2 (ABB97082), Caenorhabditiselegans, CeCHS1 (Np_492113), CeCHS2 (Np_493682);Saccharomyces cerevisiae,ScCHS1 (PO8004); ScCHS2 (AAA34493) ScCHS3 (P29465); D. pulex, DaphCHS1( EFX76951), DaphCHS2 (EFX80669). |