Determination of the Cutoff Point of the Absolute Value of MGMT mRNA for Predicting the Therapeutic Resistance to Temozolomide in Glioblastoma
Received: 01-May-2019 / Accepted Date: 20-May-2019 / Published Date: 27-May-2019
Abstract
Abstract Purpose: In our previous study we have reported the absolute value of O6-methylguanine-DNA methyltransferase (MGMT) messenger RNA (mRNA) which was acquired by using real-time reverse transcription polymerase chain reaction (RT-PCR). Probably the value may be useful for envisioning both the prognosis and the outcomes of the therapy for glioblastoma (GB) treated by Temozolomide (TMZ). Methods: MGMT mRNA was studied in 55 newly diagnosed cases of GB treated with TMZ and radiation (with less than 75 y and had a Karnofsky performance status (KPS) of at least 60), based upon the technique of realtime reverse transcription polymerase chain reaction (RT-PCR) using the TaqMan probe. A receiver operating characteristic analysis was carried out in order to determine the cutoff points for progression free survival (PFS) as well as overall survival (OS). Results: In 55 patients with GB, 1200 and 3600 for PFS (specificities of 88.9 and 66.7%, and sensitivities of 36.7 and 50.0%, respectively); 1200, 2100 and 2900 copies/μg RNA for OS (specificities of 83.6, 73.9 and 69.6%, and sensitivities of 39.1, 47.8 and 52.2%, respectively) were the candidate cutoff points. Significantly longer PFS and OS were observed in patients who did not exceed 1200 copies/μg RNA (p=0.0035 for PFS and 0.0189 for OS by logrank test). Median overall survival of the GB patients who had less than 1200 copies/μg RNA treated with TMZ and radiation was 36 months. Conclusion: One Thousand and two hundreds copies/μg RNA appeared to be the most reasonable cutoff point of MGMT mRNA in GB for deciding to use other anti-tumor drugs such as Bevacizumab together with TMZ.
Keywords: O6-methylguanine-DNA methyltransferase; Glioblastoma; Real-time polymerse chain reaction; Temozolomide; Cutoff point; Bevacizumab; Receiver operating characteristic analysis
Introduction
Temozolomide (TMZ) is the only chemotherapeutic agent which has the level 1 evidence for the glioblastoma (GB) treatment. Among all solid tumors including brain tumors, it is adjudged to elicit worst prognosis [1,2]. Indeed some other anti-tumor drugs such as 1-(4-amino-2-methyl-5-pyrimidynyl) methyl-3-(2-chloroethyl)- 3-nitrosourea hydrochloride (ACNU), Bevacizumab, Human interferon-β, Vincristine, and Carmustine wafer are approved and used for GB therapy in Japan, but there seems to be very little evidence with regard to their effectiveness for GB therapy [3]. Additionally, except for Bevacizumab, there have been no reports on the effectiveness of these drugs together with TMZ. The effect of Bevacizumab in addition to TMZ-radiation has been yet controversial. O6-Methylguanine- DNA methyltransferase (MGMT) is one of the drug-resistance genes for alkylating agents such as ACNU, Procarbazine, and TMZ [4,5]. Methylation of the DNA promoter domain leads to MGMT silencing [6] and is considered as a good prognostic factor for GB as per the previous report [7]. Whether MGMT methylation can predict the clinical results of temozolomide treatment for GB was uncertain due to lack of clear evidence.
In our previous study we have reported the absolute value of MGMT messenger RNA (mRNA) which was obtained using real-time reverse transcription polymerase chain reaction (RT-PCR). It may be useful for not ony predicting the prognosis but also the clinical results of TMZ therapy for GB [8]. In the study reported here, the cutoff points of the absolute value of MGMTmRNA in GB were determined by receiver operating characteristic (ROC) analyses for developing new individual adjuvant therapy (IAT).
Patients and Methods
Patients
A total of fifty-five patients who were newly diagnosed for GB meeting the criteria of less than 75 y and with a Karnofsky performance status (KPS) of a minimum of 60 (Eastern Cooperative Oncology Group performance status: 0-2) were included in the study. These patients were treated from 2006 to 2016 at the five different Institutes which were participating constituents of Tokyo Consortium for Brain Tumor Treatment. Forty patients were treated at Tokyo Medical University Hospital, Tokyo, 10 patients at National Cancer Center Hospital, Tokyo; 1 patient at Shioya Hospital, International University of Health and Welfare, Yaita; 3 patients at Kawasaki Hospital, Hitachiohta; and remaining 1 patient at Tokyo-Nishi Tokushukai Hospital, Tokyo, Japan. All of the 55 GB patients received temozolomide and radiation after surgery in accordance with the Stupp protocol [2]. Written informed consent for the quantitation of MGMT mRNA in tumor samples was provided by all patients. RT-PCR based quantitation of MGMT mRNA (Cutoff Point of MGMT mRNA in Glioblastoma) was approved by the Ethics Committee at Tokyo Medical University in the year 2005, at Kitasato University in the year 2002, at the International University of Health and Welfare in the year 2012, and at Tokushukai Hospitals in the year 2015.
Real-time polymerase chain reaction based quantitation of MGMT mRNA absolute value
Collection of tumor samples and quantitation of MGMT mRNA was performed by Special Reference Laboratory Co. Ltd., Hino, Japan. The method used to quantify the absolute value of MGMT mRNA by RT-PCR was described previously [9]. Briefly, the guanidinium thiocyanate-phenol-chloroform mediated extraction was performed using Isogen (WAKO Junyaku) for extracting total RNA from either ~10 mg of freshly obtained tumor samples stored at 4°C in QIAGEN RNAlater Tissue Protect Tubes (AMBION Inc) or tissues frozen at -70°C [10]. From 1 μg of the extracted total RNA, the complementary DNA (cDNA) was synthesized and was subsequently incubated at 37°C for 60 minutes. The real-time polymerase chain reaction was carried out using a TaqMan Universal Master Mix (Applied Biosystems) comprising of 120 nM of each primer [11], 200 nM of probe (5- CGA GCA GTG GGA GGA GCA ATG AGA-3), and 2.5 μL of each cDNA sample, with denaturation at 95°C for 10 minutes and 50 cycles (at 95°C for 30 seconds, 60°C for 40 seconds, and 72°C for 30 seconds) in a real-time PCR system. The levels of glyceraldehyde-3- phosphate dehydrogenase (GAPDH) mRNA expression were used as a quantitative internal control. Using a standard curve, the expression level of each mRNA was calculated. In order to obtain an even more accurate quantification, the MGMT mRNA expression level of each sample was normalized by the expression of the GAPDH gene.
Statistical analysis
All the statistical analyses were carried in Microsoft® Excel Tokei Software. The progression-free survival periods and overall survival of the 55 GB patients were analyzed, and the cases with less than 8.1 months progression-free survival and 15 months overall survival were judged to be TMZ resistant according to the results of Brain Tumor Registry Japan [12]. The cutoff points were determined in GB treated with TMZ less than 75 y and KPS of at least 60 by ROC analysis. The specificity and sensitivity of each cutoff point were calculated. Kaplan- Meier analysis was performed to evaluate each survival time, and the logrank (Mantel-Cox) test was considered for analysing the binary variables (less than and at least the cutoff points). 2-tailed p values are reported. Determination of the statistical significance of the data analysis was set at a probability level of 5% (p=0.05).
Results
ROC analysis for selecting candidate cutoff points for MGMTmRNA in GB
The candidate cutoff points in each GB group were calculated by ROC analysis. In the 55 GB patients treated with TMZ and radiation who were less than 75 y and had a KPS of at least 60, 1200 and 3600 copies/μg RNA were the candidate cutoff points for predicting PFS (specificities of 88.9 and 66.7%, and sensitivities of 36.7 and 50.0%, respectively), and 1200, 2100 and 2900 copies/μg RNA were the candidate cutoff points for predicting OS (specificities of 83.6, 73.9 and 69.6%, and sensitivities of 39.1, 47.8 and 52.2%, respectively) [6]. Kaplan-Meier analyses of GB patients with less than and at least the candidate cutoff points calculated by ROC analyses were used to estimate each survival time, and the logrank (Mantel-Cox) tests were used to evaluate binary variables. In the 55 GB patients treated with TMZ and radiation who were less than 75 y and had a KPS of at least 60, progression free survival (PFS) and overall survival (OS) were found to be significantly longer in those who had less than 1200 copies/μg RNA (p=0.0035 for PFS and 0.0189 for OS by logrank test) as shown in Figures 1-5. Median overall survival of the GB patients who had less than 1200 copies/μg RNA treated with TMZ and radiation was 36 months. The summary of all the analytical results has been presented in Table 1.
| Survival | Cutoff point (copy/µg RNA) | Sensitivity (%) | Specificity (%) | Median Less than | Survival (mo) At least | Logrank (p=) |
|---|---|---|---|---|---|---|
| Progression free survival | 1200 | 36.7 | 88.9 | 36 | 10 | 0.0035 |
| 3600 | 50 | 66.7 | 15 | 10 | 0.1183 | |
| Overall survival | 1200 | 39.1 | 82.6 | 36 | 14 | 0.0189 |
| 2100 | 47.8 | 73.9 | 30 | 14 | 0.06 | |
| 2900 | 52.2 | 69.6 | 30 | 14 | 0.0528 |
Table 1: Summary of Receiver Operatic Characteristic Analyses and Kaplan-Meier Analyses for Determination of the Cutoff Point of the Absolute Value of MGMTmRNA for Predicting the Therapeutic Resistance to Temozolomide in Glioblastoma.
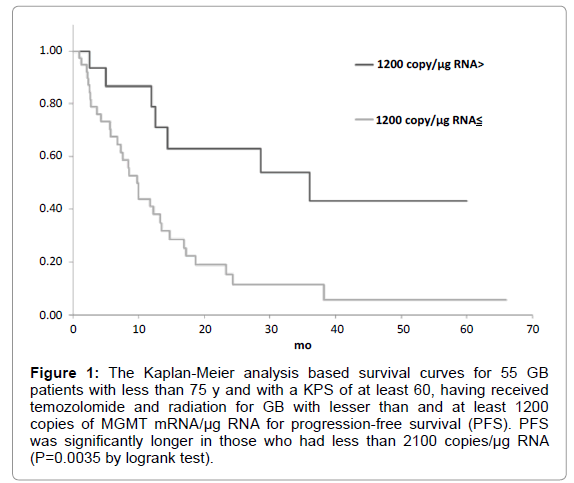
Figure 1: The Kaplan-Meier analysis based survival curves for 55 GB patients with less than 75 y and with a KPS of at least 60, having received temozolomide and radiation for GB with lesser than and at least 1200 copies of MGMT mRNA/µg RNA for progression-free survival (PFS). PFS was significantly longer in those who had less than 2100 copies/µg RNA (P=0.0035 by logrank test).
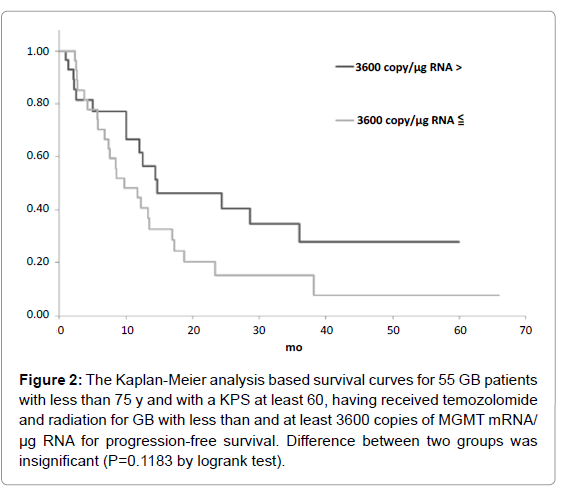
Figure 2: The Kaplan-Meier analysis based survival curves for 55 GB patients with less than 75 y and with a KPS at least 60, having received temozolomide and radiation for GB with less than and at least 3600 copies of MGMT mRNA/ µg RNA for progression-free survival. Difference between two groups was insignificant (P=0.1183 by logrank test).
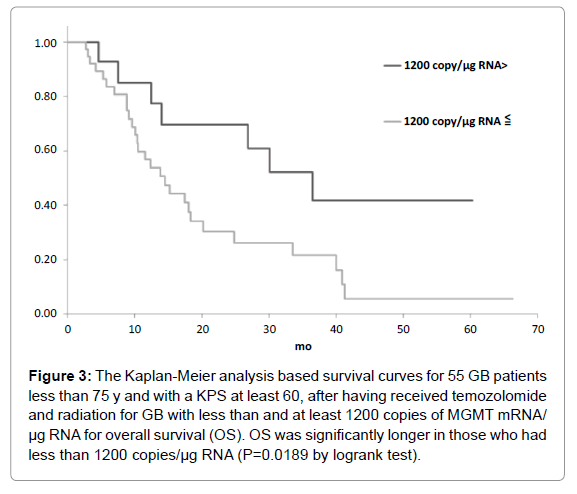
Figure 3: The Kaplan-Meier analysis based survival curves for 55 GB patients less than 75 y and with a KPS at least 60, after having received temozolomide and radiation for GB with less than and at least 1200 copies of MGMT mRNA/ µg RNA for overall survival (OS). OS was significantly longer in those who had less than 1200 copies/µg RNA (P=0.0189 by logrank test).
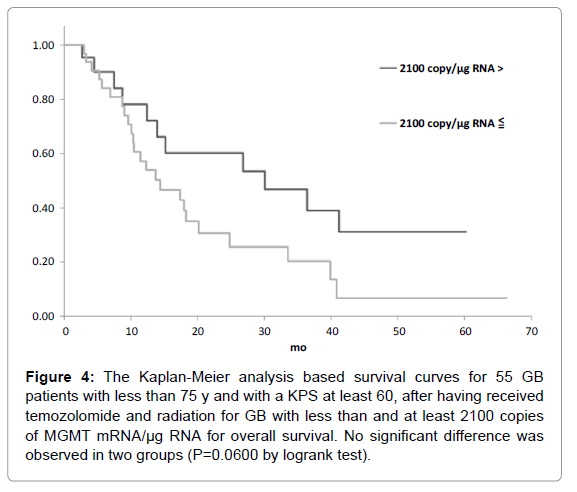
Figure 4: The Kaplan-Meier analysis based survival curves for 55 GB patients with less than 75 y and with a KPS at least 60, after having received temozolomide and radiation for GB with less than and at least 2100 copies of MGMT mRNA/µg RNA for overall survival. No significant difference was observed in two groups (P=0.0600 by logrank test).
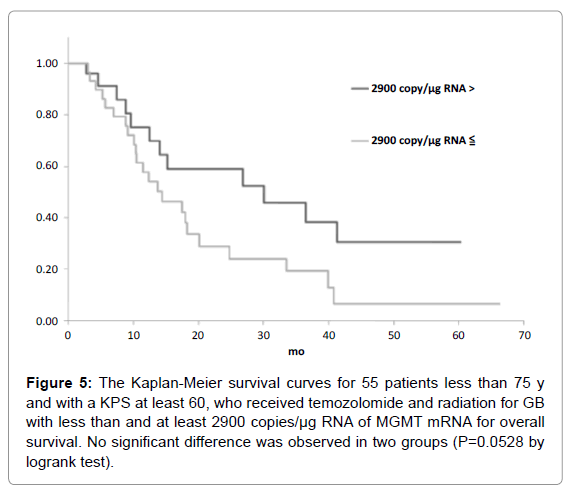
Figure 5: The Kaplan-Meier survival curves for 55 patients less than 75 y and with a KPS at least 60, who received temozolomide and radiation for GB with less than and at least 2900 copies/µg RNA of MGMT mRNA for overall survival. No significant difference was observed in two groups (P=0.0528 by logrank test).
Discussion
As mentioned in the introduction section, MGMT is silenced by hypermethylation of DNA promoter domain [6]. Many tests of MGMT, such as immunohistochemistry, methylation-specific PCR, pyrosequencing, Southern blotting, and the absolute values of MGMT mRNA, have been reported [13]. Among these methods, immunohistochemistry and methylation-specific PCR are not fundamentally quantitative, and therefore they are inappropriate for ascertaining the indications and procedures for TMZ therapy, although Hegi et al. [7], substantiates the predictive potential of a quantitative methylation-specific PCR for the evaluation of MGMT promoter methylation. Immunohistochemistry yields low specificity while it is not quantitative as per the analysis of MGMT at the protein level [14]. The activity of MGMT appears to have high correlation with clinical resistance to alkylating agents though mRNA levels as quantitated by RT-PCR might not correlate with enzymatic activity always [15]. mRNA expression has been preferentially investigated because quantitation of MGMT activity is quite complex requiring longer time for clinical utility [11,16]. Real-time polymerase chain reaction technique is relatively simple, quick and clinically relevant for the estimating the extent of resistance to the alkylating agents as described previously. In a previous study, the results pertaining to the methylation specific PCR and sequencing were compared to the MGMT mRNA expression level based on real-time quantitative RT-PCR and have reported that the extent of MGMT mRNA expression was strongly correlated with the methylation status of MGMT promoter [17]. Hence, the level of MGMT mRNA expression appears to be relatively more precise prognostic factor than the result from methylation-specific PCR.
In fact, recently pyrosequencing was effectively used for the purpose of quantitative detection of methylation. The extent of MGMT methylation based on pyrosequencing was previously reported as factor of prognosis in case of GB therapy by temozolomide as well as radiation, but clear cutoff points for applicability of pyrosequencing to estimate MGMT methylation was not ascertained [18]. Several reports on cutoff points of methylation of MGMT promoter based on pyrosequencing and the precise selection of hot spots of promoter methylation impacts the results [19,20]. On the contrary, based on SYBR Green method, Everhard et al. reported a methylation status concordance level of as high as 85% between pyrosequencing and RNA expression level by RT-PCR and further recommended that if repression of transcription is the key mechanism for higher chemosensitivity of tumors with MGMT methylation, then a substantial rate of discordance warrants caution while deciding therapeutic strategy solely based on MGMT methylation status [21].
On the other hand, almost all previous clinical studies have excluded elderly patients and those with a low KPS. Age and KPS have been reported to be the most significant prognostic factors in GB [19]. An important objective of the present study was to identify the cutoff point for MGMT mRNA absolute value for adding other anti-tumor agents to TMZ and radiation therapy. The present study showed that, in GB patients treated with TMZ and radiation who were less than 75 y and who had a KPS of at least 60, 1200, 2100, 2900 and 3600 copies/μg RNA of MGMTmRNA were the candidate cutoff points for predicting both PFS and OS. Among these, significantly longer PFS and OS were observed in patients who did not exceed 1200 copies/μg RNA. One Thousand and two hundreds copies/μg RNA appeared to be the most reasonable cutoff point of MGMTmRNA in GB for deciding to use other anti-tumor drugs such as Bevacizumab together with TMZ. GB at least 1200 copies/μg RNA seemed to be able to tolerate TMZ, other anti-tumor drug may be used without TMZ. The fact that MGMT is a suicide enzyme and TMZ itself consume the MGMT molecule is one of the reasons why the Stupp protocol of continuous administration of low-dose temozolomide at the first adjuvant therapy is significantly effective [22]. TMZ should be use in GB with at least 1200 copies/μg RNA of MGMTmRNA in order to overcome resistance to TMZ by MGMT.
Bevacizumab and human interferon-β (hIFN-β) are available for glioma therapy in Japan. Bevacizumab was approved for use in Japan in 2013 for both initially diagnosed and recurrent malignant gliomas. Bevacizumab is reported to be more effective for treating MGMTmethylated tumors than unmethylated tumors [23]. Although phase III studies have reported that a course of bevacizumab with TMZ and radiation conferred prolonged PFS for patients with GB, OS was not prolonged. If we consider cross-over cases in which Bevacizumab was used after recurrence in placebo groups, Bevacizumab seems to be effective in selected glioma cases. Human interferon (hIFN)-β has been used for glioma therapy since 1985, especially in Japan. Although there is no evidence on the effectiveness hIFN-β addition to TMZ for prolonging PFS and OS, hIFN- β was found to downregulate MGMT expression and appears to be useful for treating GB with unmethylated MGMT, i.e., in the presence of high levels of MGMT mRNA [24,25]. A recent report showed that interferon-α/β enhances TMZ activity against MGMT-positive glioma stem-like cells [26].
A new IAT based on the results of MGMTmRNA quantitation by real-time RT-PCR is recommended in initially diagnosed GB. GB with less than 1200 copies/μg RNA seems to have a good prognosis and no treatment other than TMZ and radiation is used, at least in the initial adjuvant therapy. For GB with at least 1200 copies/μg RNA, the co-use of Bevacizumab and TMZ accompanied by radiation may be an effective modality. Although presented our study has been retrospective and on limited patients, MGMTmRNA quantitation is seemed to be available to improve the adjuvant therapy for GB.
Conclusions
One Thousand and two hundreds copies/μg RNA appeared to be the most reasonable cutoff point of MGMTmRNA in GB for deciding to use other anti-tumor drugs such as Bevacizumab together with TMZ. A multicenter randomized clinical trial will be needed to obtain evidence regarding this new IAT based on MGMTmRNA quantitation.
Acknowledgment
We thank the Center for Molecular Biology and Cytogenetics Gene Analysis Section, Special References Laboratory, Inc., Hino, Japan, for sample collection and quantitation of MGMT mRNA.
References
- Anderson E, Grant R, Lewis SC, Whittle IR (2008) Randomized Phase III controlled trials of therapy in malignant glioma: where are we after 40 years? Br J Neurosurg 22: 339-349.
- Stupp R, Mason WP, van den Bent MJ, Weller M, Fisher B, et al. (2005) Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N Engl J Med 352: 987-996
- Yoshida J, Kajita Y, Wakabayashi T, Sugita K (1994) Long-term follow-up results of 175 patients with malignant glioma: importance of radical tumour resection and postoperative adjuvant therapy with interferon, ACNU and radiation. Acta Neurochir (Wien) 127: 55-59.
- Bocangel DB, Finkelstein S, Schold SC, Bhakat KK, Mitra S, et al. (2002) Multifaceted resistance of gliomas to temozolomide. Clin Cancer Res 8: 2725-2734.
- Tano K, Shiota S, Collier J, Foote RS, Mitra S. (1990) Isolation and structural characterization of a cDNA clone encoding the human DNA repair protein for O6-alkylguanine. Proc Natl Acad Sci USA 87: 686-690.
- Esteller M, Hamilton SR, Burger PC, Baylin SB, Herman JG (1999) Inactivation of the DNA repair gene O6-methylguanine-DNA methyltransferase by promoter hypermethylation is a common event in primary human neoplasia. Cancer Res 59: 793-797.
- Hegi ME, Diserens AC, Gorlia T, Hamou MF, de Tribolet N, et al. (2005) MGMT gene silencing and benefit from temozolomide in glioblastoma. N Engl J Med 352: 997-1003.
- Tanaka S, Akimoto J, Narita Y, Oka H, Tashiro T (2014) Is the absolute value of O6 – methylguanine - DNA methyltransferase gene messenger RNA a prognostic factor or does it predict the results of treatment of glioblastoma with Temozolomide? J Neurosurg 121: 818-826.
- Tanaka S, Oka H, Fujii K, Watanabe K, Nagao K, et al. (2005) Quantitation of O6 – Methylguanine - DNA Methyltransferase Gene Messenger RNA in Gliomas by Means of Real-Time RT-PCR and Clinical Response to Nitrosoureas. Cell Mol Neurobiol 25: 1067-1071.
- Chomczynski P, Sacchi N (1987) Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal Biochem 162: 156-159.
- Tanaka S, Kobayashi I, Utsuki S, Oka H, Fujii K, et al. (2003) O6-methylguanine-DNA methyltranspherase gene expression in gliomas by means of real-time quantitative RT-PCR and clinical response to nitrosoureas. Int J Cancer103: 67-72.
- Narita Y, Shibui S (2015) Committee of Brain Tumor Registry of Japan Supported by the Japan Neurosurgical Society. Trends and outcomes in the treatment of gliomas based on data during 2001-2004 from the Brain Tumor Registry of Japan. Neurol Med Chir (Tokyo) 45: 286-295.
- Dullea A, Marignol L (2016) MGMT testing allows for personalised therapy in the temozolomide era. Tumour Biol 37: 87-96.
- Nagane M, Kobayashi K, Ohnishi A, Shimizu S, Shiokawa Y (2007) Prognostic significance of O6-methylguanine-DNA methyltransferase protein expression in patients with recurrent glioblastoma treated with temozolomide. Jpn J Clin Oncol 37: 897-906.
- Silber JR, Mueller BA, Ewers TG, Berger MS (1993) Comparison of O6-methylguanine-DNA methyltransferase activity in brain tumors and adjacent normal brain. Cancer Res 53: 3416-3420.
- Tanaka S, Kamitani H, Amin MR, Watanabe T, Oka H, et al. (2000) Preliminary individual adjuvant therapy for gliomas based on the results of molecular biological analyses for drug-resistance genes. J Neurooncol 46: 157-171.
- Kreth S, Thon N, Eigenbrod S, Lutz J, Ledderose C, et al. (2011) O-methylguanine-DNA methyltransferase (MGMT) mRNA expression predicts outcome in malignant glioma independent of MGMT promoter methylation. PLoS One 6:e17-156.
- Dunn J, Baborie A, Alam F, Joyce K, Moxham M, et al. (2009) Extent of MGMT promoter methylation correlates with outcome in glioblastomas given temozolomide and radiotherapy. Br J Cancer 101: 124-131.
- Villani V, Casini B, Pace A, Prosperini L, Carapella CM, et al. (2015) The prognostic value of pyrosequencing-detected MGMT promoter hypermethylation in newly diagnosed patients with glioblastoma. Dis Markers 604-719.
- Xie H, Tubbs R, Yang B (2015) Detection of MGMT promoter methylation in glioblastoma using pyrosequencing. Int J Clin Exp Pathol 8: 1790-1796.
- Everhard S, Tost J, El Abdalaoui H, Crinière E, Busato F, et al. (2009) Identification of regions correlating MGMT promoter methylation and gene expression in glioblastomas. Neuro Oncol 11: 348-356.
- Tolcher AW, Gerson SL, Denis L, Geyer C, Hammond LA, et al. (2003) Marked inactivation of O6-alkylguanineDNA alkyltransferase activity with protracted temozolomide schedules. Br J Cancer 88: 1004-1011.
- Lai A, Tran A, Nghiemphu PL, Pope WB, Solis OE, et al. (2011) Phase II study of bevacizumab plus temozolomide during and after radiation therapy for patients with newly diagnosed glioblastoma multiforme. J Clin Oncol 29: 142-148.
- Motomura K, Natsume A, Kishida Y, Higashi H, Kondo Y, et al. (2011) Benefits of interferon-ß and temozolomide combination therapy for newly diagnosed primary glioblastoma with the unmethylated MGMT promoter: a multicenter study. Cancer 117: 1721-1730.
- Natsume A, Ishii D, Wakabayashi T, Tsuno T, Hatano H, et al. (2005) IFN-beta down-regulates the expression of DNA repair gene MGMT and sensitizes resistant glioma cells to temozolomide. Cancer Res 65: 7573-7579.
- Shen D, Guo CC, Wang J, Qiu ZK, Sai K, et al. (2015) Interferon-a/ß enhances temozolomide activity against MGMT-positive glioma stem-like cells. Oncol Rep 34: 2715-2721.
Citation: Tanaka S, Akimoto J, Narita Y (2019) Determination of the Cutoff Point of the Absolute Value of MGMT mRNA for Predicting the Therapeutic Resistance to Temozolomide in Glioblastoma. J Oncol Res Treat 4: 139.
Copyright: © 2019 Tanaka S, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Select your language of interest to view the total content in your interested language
Share This Article
Open Access Journals
Article Usage
- Total views: 3435
- [From(publication date): 0-2019 - Dec 09, 2025]
- Breakdown by view type
- HTML page views: 2558
- PDF downloads: 877