Immunogenic and Protective Properties of Recombinant Proteins Based on Meningococcal Iga1 Protease
Received: 07-Oct-2015 / Accepted Date: 08-Dec-2015 / Published Date: 15-Dec-2015 DOI: 10.4172/2572-2050.1000102
Abstract
Recombinant proteins (M1K2–N963-LEH6, MA28–N963-LEH6 and ME135–H328-LEH6) have been created on the basis of the genome sequence of IgA1 protease of N. meningitidis serogroup B strain H44/76. It is revealed that, similarly to the native enzyme isolated earlier from N. meningitidis serogroup A strain A208, these proteins induce formation of animal protection against the infection with the virulent strain of meningococcus serogroup B. It is shown that these compounds are promising as a basis for a polyvalent anti-meningococcal vaccine.
Keywords: IgA1 protease; Recombinant proteins; Meningococcus; Immunogenicity; Protectivity
5459Introduction
At present, meningitis is a frequent neuroinfection, and the most widely distributed bacteria-caused meningitis (up to 56%) is a socially dangerous disease. Due to existence of vaccines against meningitis caused by the meningococcus serogroups A, C, Y, W135 which are based on their capsular polysaccharides [1,2] and to creation of the protein-based polycomponent serogroup B monovaccine [3], there is a possibility of preventing the meningococcal infection caused by the epidemically most dangerous serogroups. However, all currently used vaccines are, as a rule, targeted rather narrowly against a specific infectious agent. Besides, such vaccination creates for a human organism an enormous antigenic load, and therefore, it is necessary to search for variants of a polyvalent vaccine which would be capable of protecting against the variety of circulating and permanently mutating meningococcus strains. Thus, the search for new approaches for preventing the meningococcal infection is still urgent.
The bacterial IgA1 protease (IgA1pr) which is the major virulence factor of many pathogens was earlier proposed as a basis for such polyvaccine [4-6]. It is known [7] that IgA1 proteases promote the colonization of mucous membranes by bacteria and their penetration into the organism’s internal medium because of cleavage by these enzymes of the human secretory immunoglobulins A1 (sIgA1). Thus, the destruction of IgA1 proteases on this stage of the invasion can be an obstacle for development of infection making difficult an adhesion of the bacteria on the mucous membrane surface. Comparison of the primary structures of IgA1 proteases of various meningococcus serogroups and of some other pathogenous microbes (H. influenzae, N. gonorrheoeae, etc.) revealed a high homology (up to 70%) of the amino acid sequences of this enzyme [8]. Therefore, it was reasonably to suppose that vaccination with IgA1pr-based preparations would promote an appearance of a steady immunity to all meningococci and, possibly, also to other pathogens with the virulence associated with this enzyme.
Data on the protectivity of our previous preparations, the native IgA1pr of the meningococcus serogroup A strain A208 and of two variants of the recombinant IgA1pr of the meningococcus serogroup B strain H44/76 (the enzymatically active and mutant by the Ser267 residue of the enzyme catalytic center) confirmed our expectations. These three preparations were shown to have identical immunogenic and protective properties and protected up to 80% of experimental animals infected directly with meningococcus serogroups A, B, and C [9-12], These results allowed us to consider IgA1pr to be a promising basis for a polyvalent vaccine. However, the high molecular weight of the enzyme and, as a consequence, a high antigenic load on the organism associated with the immunization are serious difficulties for creation of vaccines. This made us to search for truncated immunogenic fragments of IgA1pr which would possess the protective properties and could be used as an antigenic basis of a polyvalent vaccine.
The purpose of this work was to design and prepare on the basis of the IgA1pr gene new truncated recombinant proteins with the different primary structure and to study effects of structural changes on the immunogenic and protective properties of these preparations against the meningococcal infection.
Methods
In this paper, we examined the three recombinant proteins, which are fragments of IgA1 protease precursor from N. meningitidis of serogroup B strain H44/76 [https://www.ncbi.nlm.nih.gov/protein/289063650] with different primary structures, and contain His tags at the C-terminal parts of the molecules: M1K2–N963-LEH6 (protein I), MA28–N963-LEH6 (protein II), and ME135–H328-LEH6 (protein III). Using the recombinant DNA technology, the three producer strains were obtained which provided an efficient expression of these recombinant proteins.
The earlier created producer strain E. coli BL21 (DE3)/pBIGAPS1 [10] was used for production and isolation of protein I.
For cloning the DNA sequence encoding the protein II in the vector pET-31(a+) the primers (IgA1-Nde-28 and IgA1-Xho-963, Table 1) were inserted with sequences of the recognition sites of the restrictases NdeI and XhoI (for IgA1-Nde-28 and IgA1-Xho-963, respectively), which were present in the vector polylinker. The nucleic acid fragment was amplified according to the standard protocol in 25 PCR cycles, with the plasmid pBIGAPS1 DNA used as a template. For cloning the DNA sequence encoding the protein III in the vector pET-28(b+), the gene iga was amplified using the oligonucleotides IgA1-E135 and IgA1-H328-Xho, and with the plasmid pBIGAPS1 DNA used as a template. The desired product of PCR was treated with the restrictase XhoI and cloned into the vector pET-28b(+) (Novagen) using the unique restriction sites NcoI (the blunted one) and XhoI. In both cases the site of the restriction site XhoI was inserted to provide the His tag presence in the C-terminal region. Oligonucleotide primers used for cloning by the PCR method of the proteins II and III and for obtaining expressing constructs based on them are shown in Table 1.
| Protein | Primers |
|---|---|
| Protein II | IgA1-Nde-28-forward 5’-GACAGCCATATGGCATTGGTCAGAGACGATGTC-3’ |
| IgA1-Xho-963-reverse 5’-GGGCTCGAGATTGTACAATCGGGTAATACCG-3’ | |
| Protein III | IgA1-E135-forward 5’-GAACCAAATAAAAATTGGCATCAC-3’ |
| IgA1-H328-Xho-reverse 5’-GCGCTCGAGATGATGTTCTCCATTACCTTTGA-3’ |
Table 1: Oligonucleotide primers used for cloning by the PCR method of the proteins II and III.
The PCR-fragments and vectors were treated with restrictases, and the competent cells were transformed using the standard procedures. The target sequence of all plasmids obtained by PCR was confirmed by DNA sequencing.
A relevant expression constructs were transformed into E. coli BL21 (DE3) cells. The transformed cells were plated on an LB plate containing kanamycin and incubated at 37C overnight. Several colonies were collected and cultured in 5 ml of LB medium containing kanamycin at 37°C overnight with shaking. A relevant overnight bacterial culture was transferred to 500 ml of kanamycin containing LB medium and shaking was continued at 37°C. When OD600 of the bacterial culture reached 0.8, IPTG was added to a final concentracion of 0.5 mM and the culture was incubated at 37°C for 2 hours with shaking. The bacterial culture was centrifuged at 5,000 rpm for 10 min and the cell pellet was suspended in 20 ml of buffer (20 mM Tris, pH 8.5) containing Triton X-100. The cell pellet was sonicated on ice and then was centrifuged at 12,000 rpm and 4°C for 1 hour to obtain inclusion bodies.
Inclusion bodies were isolated and the desired proteins were purified using the common modified method described in the work [9]. For isolation, inclusion bodies were resuspended in a buffer solution containing Triton X-100, ultrasonicated, and resuspended again. After three washings, the inclusion bodies were dissolved in a buffer solution supplemented with 8 M urea. The dissolved inclusion bodies were fractionated by chromatography on Ni-Agarose. Fractions obtained as a result of elution in Imidazole gradient and containing the purified protein were combined and subjected to a step-by-step dialysis gradually decreasing the urea concentration. Upon the renaturation, the dissolved protein was purified by chromatography on Q-Sepharose. The resulting recombinant proteins were stored at −20°C. The homogeneity and molecular weight of the resulting preparations were confirmed by SDS-PAAG-electrophoresis. Amino acid sequences of target proteins were determined on the basis of nucleotide sequence of the cloned DNA.
The immunogenic and protective activity of proteins I, II and III was studied using female BALB/c mice (16 g-28 g) obtained from the Puschino Animal Breeding Center (Moscow Region). Mice were treated as per approved protocol IACUC (IBCH 58a, 15.05.15). Animals (3 groups for each protein, 5 animals in each group) were immunized by intravenous injection into the retro-orbital sinus of protein I, II or III in dose of 10 to 40 μg/mouse, of Vi-antigene polysaccharide S. typhi strain 44/46 (Vi-AG) in dose of 1 μg/mouse or of sheep erythrocytes (SE) in dose of 0.25 × 104 cells. All antigens were administered with an interval between injections of 45 days. On the 5th day after re-immunization the specific antibody level was assayed using the ELISA method by sorption of corresponding antigens on the microtiter plate [10,12]. The optical density was determined using the reader (StatFax-2100, Awareness Technology Inc.). The antibody level to the sheep erythrocytes (SE) were determined by the reaction of passive agglutination [1].
For the mixed immunization with Vi-AG or SE the samples were administered as the mixture using the same scheme. In this case the level of specific antibodies against each antigen was evaluated in comparison with this value obtained after immunization by each antigen separately. The antibody titer in blood of the immunized animals in comparison with the intact animals pointed to the immunogenicity of these preparations.
The protective activity of IgA1pr was studied using mice immunized as described above. Mice were infected intraperitoneally by live virulent cultures of meningococcus serogroup B strain H44/76 in dose of 0.25 × 106 microbial cells on the 12th day after re-immunization. The protective activity was evaluated by bacteriemia (the number of colony formed cells (CFU)) in the blood samples of infected mice 4 hours after infection by meningococcal culture by seeding the blood samples of these animals on plates with solid nutrient medium or by the number of mice survived 5 days after infection in comparison with those in the control group of unimmunized mice [10].
Statistical analysis was performed using the software MS Office Excel, the program [Probit Analysis] and t Student's test. Analysis of the distribution of potential B-cell and T-cell epitopes in the molecule structure of IgA1pr was performed using traditional methods of predicting the antigenic structure of the protein by the service https://www.iedb.org. The molecular weights of proteins were calculated using the server http://web.expasy.org/protparam.
Results And Discussion
Preparation of recombinant proteins
Earlier the producer strain E. coli BL21 (DE3)/pBIGAPS1 was created capable of providing and accumulating as inclusion bodies of the full-size IgA1pr from the serogroup B strain H44/76 which was a protein with the M1K2–N963-LEH6 sequence (protein I) [10]. A method was developed for preparing from these inclusion bodies of an active enzyme which was characterized by the specific activity toward immunoglobulin A1 and by presence, in addition to the proteolytic enzyme, the N-terminal signaling peptide (K2–A27) and the His tag in the C-terminal part of the molecule. In animal experiments this protein was shown to have a high immunogenic activity and the ability to protect the mice against infection with a living virulent culture of three main meningococcus serogroups: A, B and C. The preparation was studied in detail and taken as the basis for development of the meningococcal polyvaccine [12].
The new truncated variants of IgA1pr were obtained to study the role of structural changes on the immunogenic and protective properties of these preparations against the meningococcal infection. For this purpose the recombinant plasmid DNAs including the corresponding nucleotide sequences and two producer strains providing the effective expression of IgA1pr variants (proteins II and III) as inclusion bodies in the host cell were designed.
The protein II with the primary sequence MA28–N963-LEH6, similarly to the protein I is an active enzyme with the specific activity, but unlike the protein I it lacks the signaling peptide (K2–A27). The protein III with the sequence ME135–H328-LEH6 is significantly shorter than the protein II and contains a sequence located in the region of IgA1pr which is specified by a high density of potential T- and spatial B-cellular epitopes.
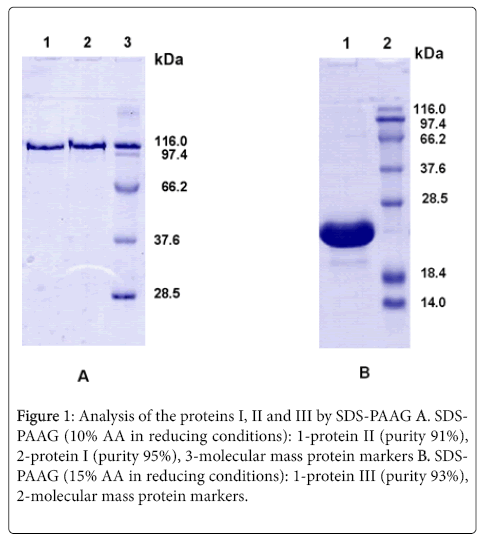
The proteins I, II and III were isolated from inclusion bodies and obtained in high purified form (see Methods) using the following stages: washing and dissolving of the inclusion bodies in denaturing conditions, renaturation of the proteins and chromatographic purification on the Ni-Agarose and Q-Sepharose FF. The homogeneity and molecular weights of the obtained proteins were confirmed by SDS-PAAG-electrophoresis. The purity of the preparations was about 91-95% (Figure 1). Amino acid sequences of target proteins were determined on the basis of nucleotide sequence of the cloned DNA.
Determination of the immunogenic and protective activity
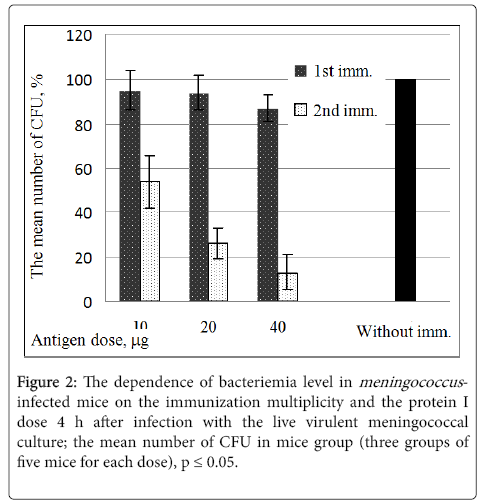
The protective activity of protein I in dependence on the immunization multiplicity and the protein I dose was evaluated in the mice experiments after infection with the live virulent meningococcal culture of serogroup B. The protection effect was evaluated by number of CFU [12].
Figure 2 shows that the double immunization of mice with the dose of 40 μg the protein I provided the optimal protection: the number of CFU in this case was 12% as compared with the number of CFU (100%) for the control (non-immunized) animals.
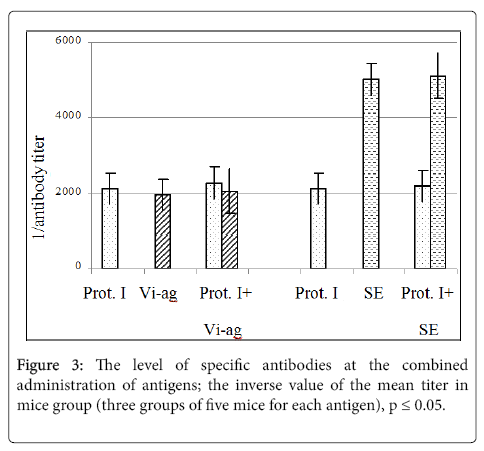
The effect of foreign antigens to the level of antibodies specific to the protein I after the combined immunization of mice with the recombinant protein I and the polysaccharide Vi-antigen from S. typhi strain 44/46, or with the recombinant protein I and sheep erythrocytes (SE). Specific antibodies were determined in sera of the mice immunized with bacterial antigens using the solid phase ELISA with the appropriate antigens sorbed onto the plate [12]. The antibodies to sheep erythrocytes (SE) were determined by the passive agglutination reaction [13]. The results are shown in Figure 3.
Thus it was shown that at the immunization of mice with the protein I combined with heterologic antigens (the Vi-antigen from S. typhi or SE) the level of specific antibodies to each of them did not change (Figure 3) that indicated the absence of the competition between these antigens during formation of the immunity.
The immunogenic and protective properties of the proteins I, II and III were then studied.
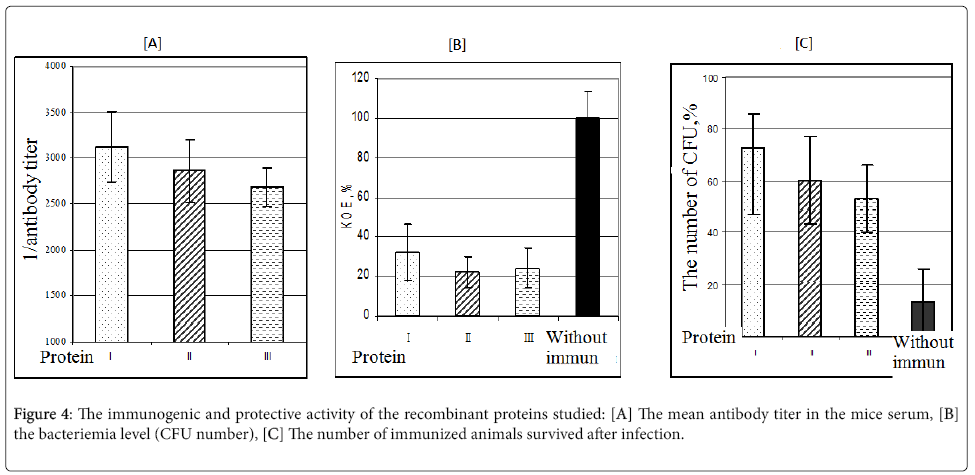
The immunological comparability of these proteins was confirmed by data of cross-titration of the antibodies to the protein I with the proteins I, II and III sorbed on the microtiter plate (Figure 4A). The level of antibodies to the protein I was statistically not different when titrated with these antigens, i.e., 1:3122 ± 381, 1:2860 ± 446, and 1:2680 ± 215, respectively.
The protective activity of the obtained proteins was determined after infection with the live virulent meningococcal culture of serogroup B. The results are shown in Figures 4B and 4C. The indicators of protective activity were the CFU number and the number of immunized animals survived after infection.
It was shown that after meningococcal infection of immunized mice the protective activity of these proteins was similar. According bacteriemia data (4 h after infection) the bacteria number in the blood samples of immunized animals was decreased by 60% to 70% (Figure 4B), and the number of animals survived after infection was 50% to 70% (Figure 4C). The difference in these values for all the proteins studied was not statistically significant.
Thus, it has been shown that the recombinant proteins II (MA28– N963-LEH6) and III (ME135–H328-LEH6) created on the basis of IgA1 protease and significantly different in the primary structure and the molecular weight possess the comparable immunogenic and protective properties and can be used as a basis for a polyvalent antimeningococcal vaccine.
References
- Frasch CE (1995) Meningococcal vaccines: Past, present and future, in Meningococcal Disease. K Cartwright (ed.),Chichester, John Wiley and sons, Chichester, UK
- Sierra GV, Campa HC, Varcacel NM, et al. (1991) Immunogenicity of meningococcal PorA antigens in OMV vaccines. Nat. Inst. Public Health Ann: 195-210.
- Gossger N, Snape MD, Yu LM, Finn A, Bona G, et al. (2012)Immunogenicity and tolerability of recombinant serogroup B meningococcal vaccine administered with or without routine infant vaccinations according to different immunization schedules: a randomized controlled trial. JAMA 307: 573-582.
- Henderson IR, Nataro JP (2001) Virulence functions of autotransporter proteins. Infect. Immun 69: 1231-1243.
- Mistry D, Stockley RA (2006) IgA1 protease.Int J. Biochem Cell Biol 38: 1244-1248.
- Plaut AG, Bachovchin WW (1994) IgA-specific prolyl endopeptidases: serine type. Methods Enzymo l244: 137-151.
- Kazeeva TN, Shevelev AA (2007)Â Unknown functions of immunoglobulins A. Biochemistry (Mosc)72:485-494.
- Lomholt H, Poulsen K, Kilian M (1995)Â Comparative characterization of the iga gene encoding IgA1 protease in Neisseria meningitidis, Neisseria gonorrhoeae and Haemophilusin fluenzae. MolMicrobiol 15: 495-506.
- Serova OV, Me006Cnikov EE, Zinchenko AA, et al. (2011)Recombinant iga1 protease from neisseria meningitidis. obtaining and properties. Biofarmatsevt Zh 3: 42-47.
- Rumsh LD, Serova OV, Zinchenko AA, (2011) Polynucleotide encoding mutant recombinant IgA1 protease from Neisseria meningitidis of serogroup b. recombinant plasmid DNA comprising said polynucleotide. host cell containing said plasmid DNA. recombinant IgA1 protease of Neisseria meningitidis of serogroup b. method of producing mature form of IgA1 protease.
- YagudaevaEYu, Zhigis LS, Razgulyaeva OA (2010)Isolation and determination of the activity of IgA1 protease from Neisseria meningitides. Russian Journal of Bioorganic Chemistry36: 81-89.
- Kotel’nikova OV, Alliluev AP, Drozhzhina EYu (2013) Biochemistry (Moscow) Supplement Series B Biomedical Chemistry 7.
Citation: Zinchenko AA, Alliluev AP, Serova OP, Gordeeva EA, Zhigis LS, et al. (2015) Immunogenic and Protective Properties of Recombinant Proteins Based on Meningococcal Iga1 Protease. J Meningitis 1:102. DOI: 10.4172/2572-2050.1000102
Copyright: ©2015 Zinchenko AA, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Select your language of interest to view the total content in your interested language
Share This Article
Open Access Journals
Article Tools
Article Usage
- Total views: 12398
- [From(publication date): 7-2016 - Jul 16, 2025]
- Breakdown by view type
- HTML page views: 11416
- PDF downloads: 982