Research Article Open Access
Monitoring Glycosylation Profile and Protein Titer in Cell Culture Samples Using ZipChip CE-MS
Yan Wang#, Peng Feng#, Zoran Sosic* and Li Zang
Analytical Development, Biogen Inc., Cambridge, MA, USA
- *Corresponding Author:
- Zoran Sosic
Analytical Development, Biogen Inc.
Cambridge, MA, USA
Tel: 7814642000
E-mail: zoran.sosic@biogen.com
Received Date: March 30, 2017; Accepted Date: April 04, 2017; Published Date: April 08, 2017
Citation: Wang Y, Feng P, Sosic Z, Zang L (2017) Monitoring Glycosylation Profile and Protein Titer in Cell Culture Samples Using Zipchip CE-MS. J Anal Bioanal Tech 8:359. doi: 10.4172/2155-9872.1000359
Copyright: © 2017 Wang Y, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Journal of Analytical & Bioanalytical Techniques
Abstract
Rapid and sensitive product quality analysis is important for real-time monitoring during biopharmaceutical development and manufacturing. However, low level of protein concentration and complex cell culture matrix pose challenges for product quality characterization at early stages of cell line selection and process development. Here, we describe a fast and simple microfluidic ZipChip CE-MS method to measure quality attributes of monoclonal antibody protein directly from cell culture supernatant. Cell culture supernatant samples were characterized with charge-based separation using microfluidic capillary electrophoresis coupled to a high-resolution mass spectrometer. Under sample reducing conditions, multiple protein glycosylation attributes were determined on the heavy chain, whereas titer information was obtained from comparison of light chain signal intensity following sample spiking-in with heavy labeled mAb. Therefore, the protein expression and product quality can be monitored using the same method with a single microfluidic device. A total volume of ten to fifty microliter of cell culture supernatant is needed, whereas analysis time is within three minutes per sample. In addition, comparison of new method with traditional RP-LC-MS method using a set of time-course bioreactor cell culture samples has been performed. A good correlation of the levels of N-glycosylation attributes between ZipChip CE-MS of crude samples and RPLCMS analysis following Protein A (ProA) purification step has been demonstrated.
Keywords
ZipChip CE-MS; RP-LC-MS; Protein quality attribute; Titer; Recombinant monoclonal antibody; N-glycosylation; Cell culture supernatant
Abbreviations
mAb: Recombinant Monoclonal Antibody; CE: Capillary Electrophoresis; MS: Mass Spectrometry; HC: Heavy Chain; LC: Light Chain; RP-LC: Reversed Phase Liquid Chromatography; RT, Room Temperature; E/S: Enzyme to Substrate ratio; AGE: Advanced Glycation End Product; BG: Biantennary Galactosylation.
Introduction
Recombinant monoclonal antibody (mAb) production represents the fastest growing sector of approved and in pipeline developing biopharmaceutical drugs [1]. With an emphasis on the control of manufacturing process, there is increased demand for advanced product analytics to report protein quality attributes early in order to allow for potential adjustment to the manufacturing process. However, methods for at-line or in-line monitoring of protein quality attributes are still in development and only few of them have been reported [2]. Process analytical technology (PAT) tools, such as on-line Raman spectra, refractive index, pH, temperature and various gas probes are important for process control, yet they don’t offer a direct measurement of product quality attributes. In particular, the early stages of mAb development, such as clone selection and process optimization, require analytical screening approaches for intermediate samples where protein titer can be low and sample matrix is complex. Similarly, there is also need for realtime analytical methods to enable feed-back or feed-forward controls during the manufacturing of mAb batches, as opposed to traditional process that takes weeks prior to the final drug substance product quality measurement. Hence, rapid and sensitive analytical methods that potentially enable real-time quality attribute characterization are highly desirable.
Post-translational modifications, such as N- and C-terminal modifications, deamidation, oxidation, glycation and glycosylation result in a remarkable heterogeneity of mAb products. Glycosylation is one of very important post-translational modifications because it can have significant impact on antibody-dependent cell-mediated cytotoxicity (ADCC) activity [3,4], complement-dependent cytotoxicity (CDCC) activity [5], clearance [6] and immunogenicity [7,8]. Protein glycosylation can be significantly impacted by the host cell line, clone, media composition, feeding strategy and downstream processing conditions [9-11]. Therefore, N-glycosylation analysis of monoclonal antibody is widely performed in the biopharmaceutical development, and in some cases is used as release assay when glycosylation is established as a critical quality attribute [12]. To date, only a few approaches have been developed targeting at-line or in-line control of protein glycosylation during protein production. For example, highthroughput ProA purification combined with rapid subunit LC-MS analysis allowed simultaneous quantitation of multiple product quality attributes from cell culture harvest, including antibody glycosylation and glycation [13]. Another method for profiling N-glycosylation based on release of glycans, their fluorescent labelling and ultra-performance liquid chromatography (UPLC) was developed for the analysis of glycan composition of mAbs from the cell culture supernatant, [14,15]. The disadvantage of above approaches is protein purification step required prior to analysis as well as relatively lengthy analysis time. Subunit LC-MS analysis has been reported to confirm antibody identity and determine isoform glycan content directly from cell culture supernatant samples. However, extensive sample preparation including buffer exchange and enzymatic digestion were performed. In addition, high noise levels due to remaining media components interfered with MS detection [16].
Recently, a microfluidic CE-ESI-MS method has been reported to generate multiple pieces of quality information in a single analysis step, including masses of intact mAb variants, an assessment of charge heterogeneity and glycoform information of purified monoclonal antibody protein [17].
To target the current challenges with protein quality assessment during biopharmaceutical development, we have developed a novel microfluidic ZipChip CE-MS method to monitor protein quality attributes of monoclonal antibody protein. Protein isoforms and glycosylation profile are measured under reducing conditions directly from cell culture supernatant without protein purification. By spiking with heavy labeled control sample, protein titer can be measured using the same method based on relative MS signal intensities of light chains. To assess the applicability of this approach to detect change in the bioreactor, the levels of Mannose 5 glycoforms in cell culture time-course samples were monitored by ZipChip CE-MS. These findings were confirmed with traditional RPLC-MS assay that relies on an additional ProA sample purification step and both approaches correlated well. Furthermore, EndoS2 enzyme treatment was applied directly to cell culture for quantitation of the afucosylation level, as one of the important monoclonal antibody quality attribute. A good correlation for afucosylation data between ZipChip CE-MS and RPLCMS analysis on ProA purified samples has been demonstrated. In order to further compare the data obtained by the two techniques, percent of biantennary galactosylation, main neutral glycoforms (G0F, G1F and G2F) and level of aglycosylated protein have been also calculated and found to be comparable. Overall, this simple and fast approach (three minutes per sample) allows for screening of multiple product quality attributes in cell culture samples and should permit for an efficient control and understanding of manufacturing process. Due to its versatility, it is foreseeable that it could be extended for monitoring of other product attributes as well.
Methods
Materials and reagents
Optima LC/MS grade Water, Acetonitrile and Formic acid (99% purity) was obtained from Fisher Scientific. Recombinant monoclonal antibody mAb and heavy-labeled mAb were produced by Cell Culture Development at Biogen. Heavy-labeled mAb was grown in culture media contained heavy labeled 13C or 15N for Arginine and Lysine amino acids. Heavy-labeled mAb was purified through ProA purification. ZipChip HR (cat#810-00140) was from 908 Devices. Reversed phase column, Tosoh TSKgel Phenyl-5PW, (cat#18756, 2.0 × 75 mm) was used for reversed-phase intact mass analysis.
Cell culture samples
Representative cell culture supernatant samples were obtained by filtration through Steriflip-GV filter unit (EMD Millipore, Cat# SE1M179M6) to remove cells and large cell debris.
Sample desalting for CE-MS and LC/MS
Both cell culture supernatant and ProA purified eluate were desalted and buffer exchanged to 50 mM ammonium bicarbonate using Thermo Zeba 96-well spin plate at 1,000 g for 2 min before further sample reduction and/or enzymatic treatment. For RP-LC/MS analysis, cell culture supernatant was first purified by Agilent Bravo small scale ProA purification step to obtain ProA purified eluate.
Sample reduction for CE-MS and LC/MS
For reduced intact mass analysis, 10 μL (or 50 μg) of desalted cell culture supernatant or 50 μg purified proA eluate or heavy-labeled mAb spiked in at final concentration of 0.25 mg/mL were reduced with 168 mM DTT for 30 min at 37°C.
EndoS2 deglycosylation and reduction for CE-MS and LC/MS
For deglycosylated reduced intact mass analysis, either cell culture supernatant or ProA purified eluate containing 50 μg of desalted protein, was treated with GlycInator (EndoS2, Genovis, Cat # A0-GL1-020) to remove N-linked carbohydrates after first GlcNAc at E/S ratio of 4:1 (cell culture supernatant) or 1:1 (ProA purified eluate). Samples were then reduced either with 168 mM DTT for 30 min at 37°C (cell culture supernatant) or with 15 mM DTT for 45 min at 25°C (ProA purified eluate).
RP-LC-MS analysis
In RP-LC-MS analysis, the heavy chain and light chain from 5 μg of each sample were separated on a Tosoh TSKgel Phenyl column and analyzed on-line by coupling to a Thermo Exactive Plus mass spectrometer. A linear gradient was started with 10% B for 1 min, and then increased from 10% to 28% in 1 min and to 36% B in 4 min; column was then washed repeatedly by 2 cycles of increasing B to 80% in 0.5 min, and decreasing to 10% in 0.5 min. Column was equilibrated with 10% B for 3 min before next injection.
CE-ESI-MS device operation and data analysis
The ZipChip CE ion source was purchased from 908 Devices and installation followed the vendor instruction manual. Device was positioned in front of the inlet of a Thermo Exactive Plus mass spectrometer (Thermo, San Jose, CA). 29 cm long separation channel of ZipChip HR was used to separate the samples. Background electrolyte (BGE) was 50% Acetonitrile containing 1% Formic Acid. The setting of mass spectrometer was mass resolution 17, 500, in-source CID 100 eV, 1 micro scan, AGC target 1E6, maximum inject time 50, spray voltage 3 kV, capillary temperature at 380°C, S-lens RF level 80, Aux gas heater temperature 400°C, sheath gas flow rate 2, Aux gas flow rate 20 with data acquired over a mass range of m/z 800-3500. Data acquisition was accomplished through Xcalibur (Thermo Scientific) tune page, which was triggered by 908 Devices CE-ESI Instrument Control to control the ZipChip output. The settings of CE were: injection load time 5 sec at 2 psi, analysis run time 3 min, pressure assist start time 10 second, CE voltage 20 KV, ESI voltage 2 KV and shield 500 V. Deconvolution of the mass spectra was performed using the deconvolution algorithm in the BiopharmarFinder 2.0 (Thermo) software. The five most abundant charge states of heavy chain were used for glycosylation profile quantitation. The most abundant two charge-state ions in light chain were used for protein concentration measurement based on intensity ratio to two corresponding ions from spiked-in heavy labeled light chain of heavy labeled mAb.
Results
Reduced mass analysis of cell culture supernatant by ZipChip CE-MS
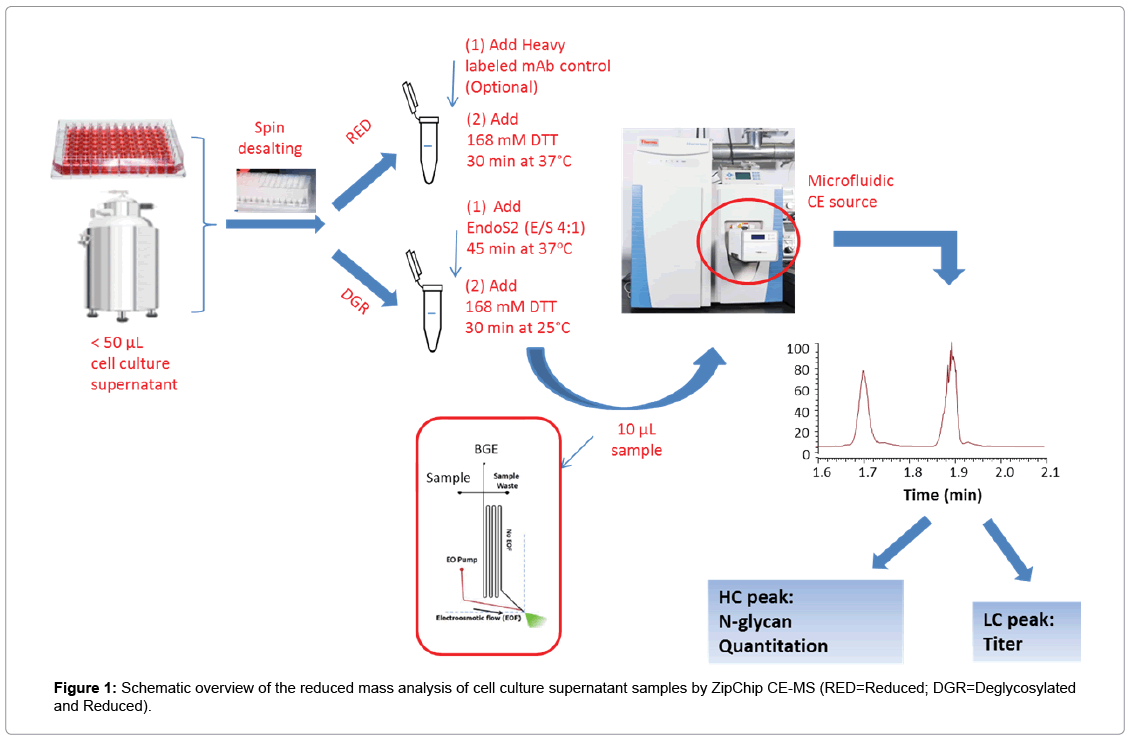
The workflow for use of ZipChip CE-MS to characterize monoclonal antibody directly from cell culture supernatant is shown in Figure 1. Reduced mass analysis was selected over intact mass analysis to increase sensitivity of identification and quantitation of HC and LC isoforms. Cell culture supernatant containing secreted monoclonal antibody was buffer exchanged to 50 mM ammonium bicarbonate during desalting step to reduce the potential ionic interference to capillary electrophoresis separation and mass spectrometry detection. Sample was then reduced with 168 mM DTT at 37°C for 30 min. For titer measurement, heavy-labeled protein was mixed into the sample prior to sample reduction step. To measure afucosylation level, sample reduction was performed after EndoS2 treatment. The HC and LC related species were assigned and quantitated based on the charge separation profile, detected masses and relative signal intensity. Since there are no N-glycosylation variants present on light chain, the protein concentration can be readily calculated using relative signal intensity of the light chain versus heavy-labeled light chain under sample reducing conditions. The ZipChip CE-MS workflow can obtain protein concentration and quality attributes information faster compared to previously reported orthogonal approaches [15].
Optimization of the sample reduction step
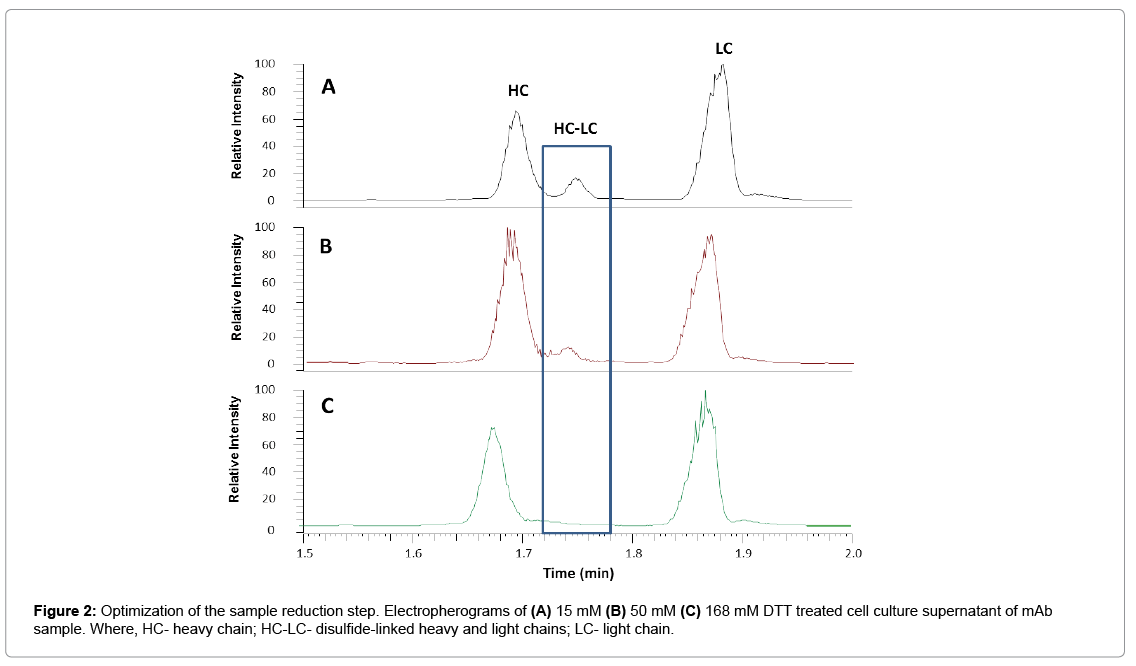
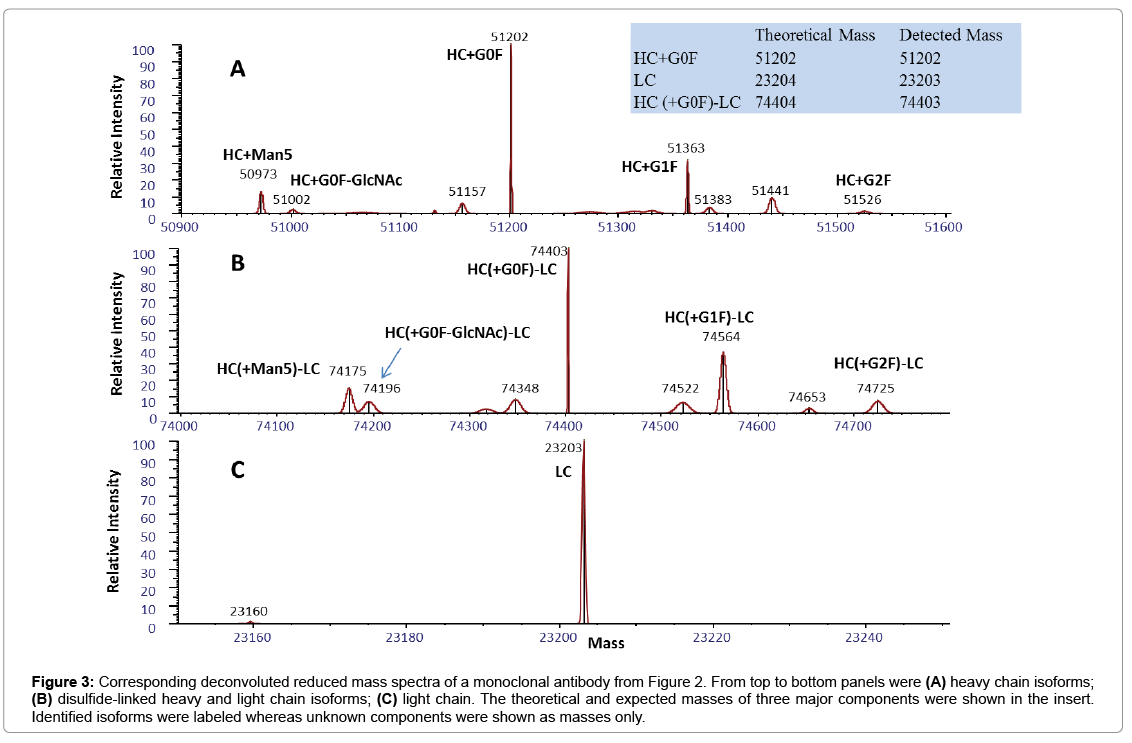
To develop the reduced condition for CE-MS analysis, 15 mM DTT concentration was first used, in order to mimic the reduction condition for RP-LC-MS analysis of ProA purified mAbs. Electropherograms of representative cell culture supernatant samples after reduction with 15 mM, 50 mM or 168 mM DTT are shown in Figure 2. The separation of HC and LC was completed in less than 2.0 min, with all protein peaks detected in a migration time window between 1.6 and 2.0 min and with an average width at base of 5 s. However, under 15 mM reduction conditions (Figure 2A), there was an extra peak, labeled as HC-LC that was increasing over time upon sample being repeatedly injected (data not shown). Figure 3 shows deconvoluted mass spectra of the HC and LC peaks corresponding to the expected heavy chain with major glycosylation isoforms (mass 51202 Da for HC-G0F, 51363 Da for HC-G1F and 51526 for HC-G2F, Figure 3A) and light chain (mass 23203 Da, Figure 3C). The detected mass 74403 Da, 74564 Da, 74725 Da (Figure 3B) from HC-LC peak agree well with expected mass of heterodimers of three HC glycosylated forms with LC. It was concluded that the artifact peak was due to disulfide scrambling as result of low DTT concentration and no denaturant in the sample buffer. Therefore, different reduction reagent concentration and dilution conditions were tested. The concentration of 168 mM DTT proved to work best to minimize the HC-LC presence, as shown in analysis of sample stored at 4°C for 24 hours (Figure 2C). From these results, an addition of 168 mM DTT was selected as optimum concentration of reducing reagent.
Figure 3: Corresponding deconvoluted reduced mass spectra of a monoclonal antibody from Figure 2. From top to bottom panels were (A) heavy chain isoforms; (B) disulfide-linked heavy and light chain isoforms; (C) light chain. The theoretical and expected masses of three major components were shown in the insert. Identified isoforms were labeled whereas unknown components were shown as masses only.
The improved detection capability of low abundance charge variants by ZipChip CE-MS
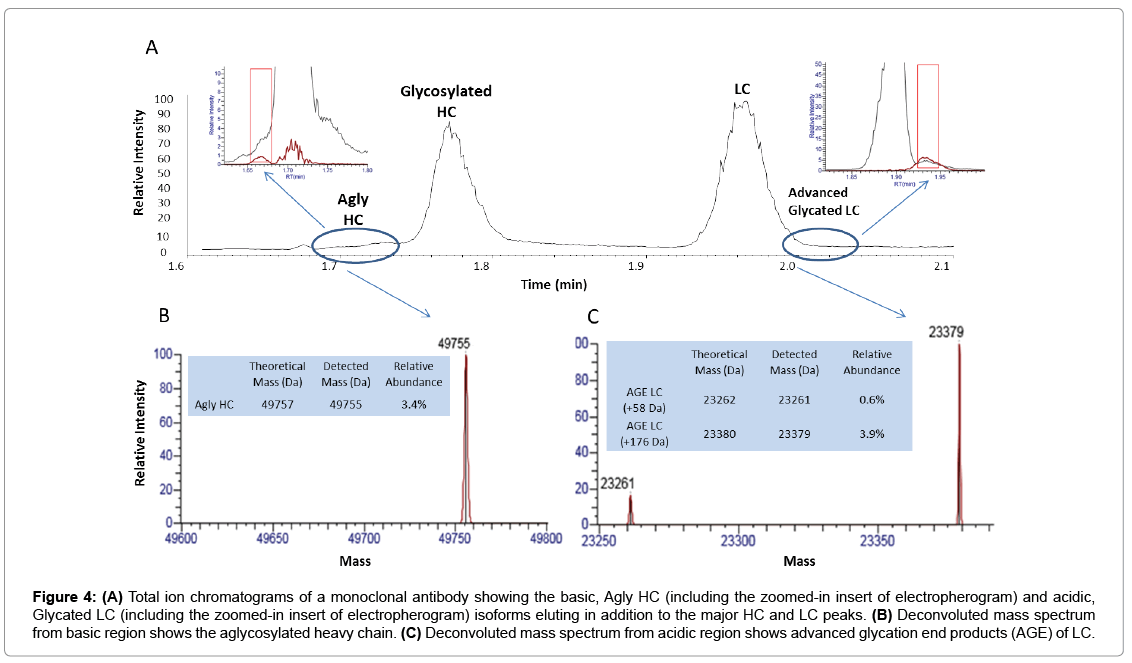
In addition to the major HC and LC peaks in the cell culture sample under optimized reducing conditions, there are also minor basic and acidic variants detected to the left and right of the HC and LC major peaks (Figure 4A). These minor peaks were deconvoluted to MWs of 49756 Da (matching aglycosylated HC, Figure 4B) and 23365 Da (matching glycated LC), respectively. The relative abundance was 3.4% for aglycosylated HC, 0.6% and 3.9%, respectively, for advanced glycated LC components (Figure 4C). As these minor species may coeluate with their 1 net charge counterparts (i.e., HC and LC isoforms) in traditional RP-LC-MS, a charge-based separation observed by Zipchip CE-MS could help against ion suppression effects and potentially allow for improved detection of similar, low abundant species.
Figure 4: (A) Total ion chromatograms of a monoclonal antibody showing the basic, Agly HC (including the zoomed-in insert of electropherogram) and acidic, Glycated LC (including the zoomed-in insert of electropherogram) isoforms eluting in addition to the major HC and LC peaks. (B) Deconvoluted mass spectrum from basic region shows the aglycosylated heavy chain. (C) Deconvoluted mass spectrum from acidic region shows advanced glycation end products (AGE) of LC.
Titer determination by ZipChip CE-MS
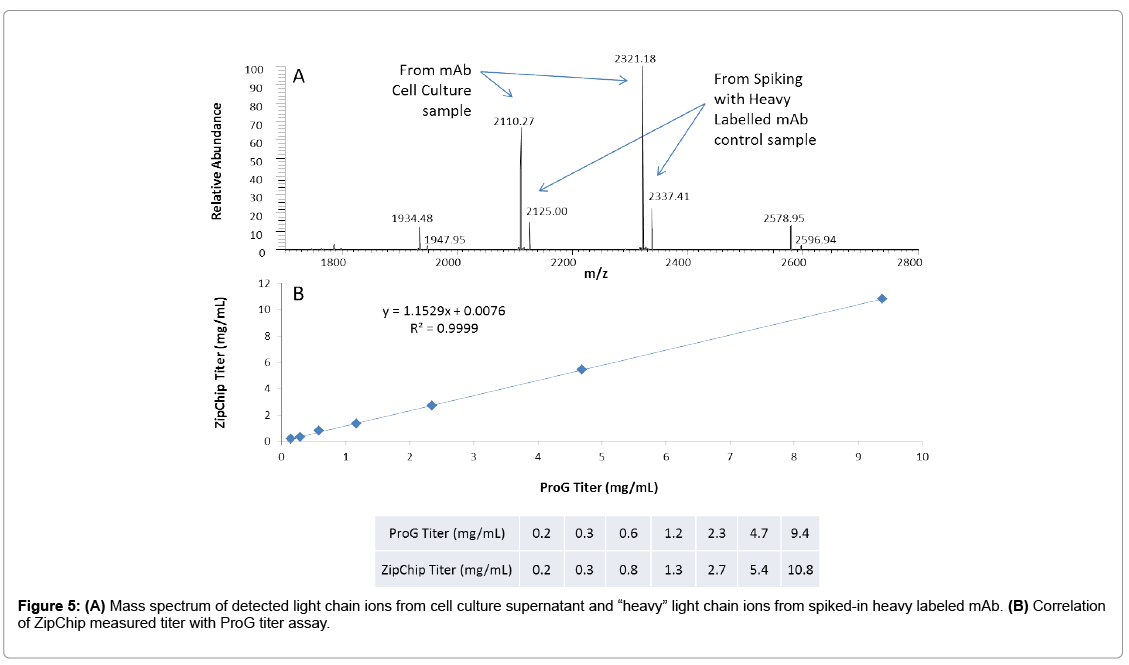
Here, we have also demonstrated ZipChip CE-MS method that can be applied to determine the protein concentration by utilizing the heavy isotope-labeled mAb as internal calibrant. This material was mixed into the cell culture supernatant sample before reduction step. In the heavylabeled mAb, all lysine and arginine residues were substituted by their 13C and 15N containing counterparts (purity >99%). The heavy labeled mAb behaves same as the control, original mAb in terms of ionization efficiency. This internal calibrant provides a way to determine “light” mAb concentration by calculating the ratio of “light” ions (from cell culture supernatant) and “heavy” ions (from the spiked-in internal calibrant). As the modifications on light chain are at very low level, this approach provides more accurate measurement than using complex heavy chain and its isoforms. Figure 5A shows the mass spectrum of “light” light chain ions and “heavy” light chain ions. The concentration of spiked heavy-labeled mAb is 0.25 mg/mL. Therefore, the titer of cell culture supernatant can be calculated by 0.25* [intensity of extracted ion chromatogram (XIC) of light chain ions 2110 and 2321y/intensity of XIC of heavy chain ions 2125 and 2337].
In Figure 5B, representative cell culture supernatant with protein concentration of ~10 mg/mL as measured by ProG titer assay was serially diluted to 0. 2 mg/mL with null cell culture media. Each dilution point was further diluted by 10 folds with water before loading onto ZipChip. The calculated and expected protein concentration showed very good correlation of R2=0.9999. This result indicates that ZipChip CE-MS could be applied to measure protein concentration accurately encompassing concentration range from 0.2 to 10 mg/mL with good linearity.
N-glycosylation detection and correlation results between ZipChip CE-MS and RP-LC-MS
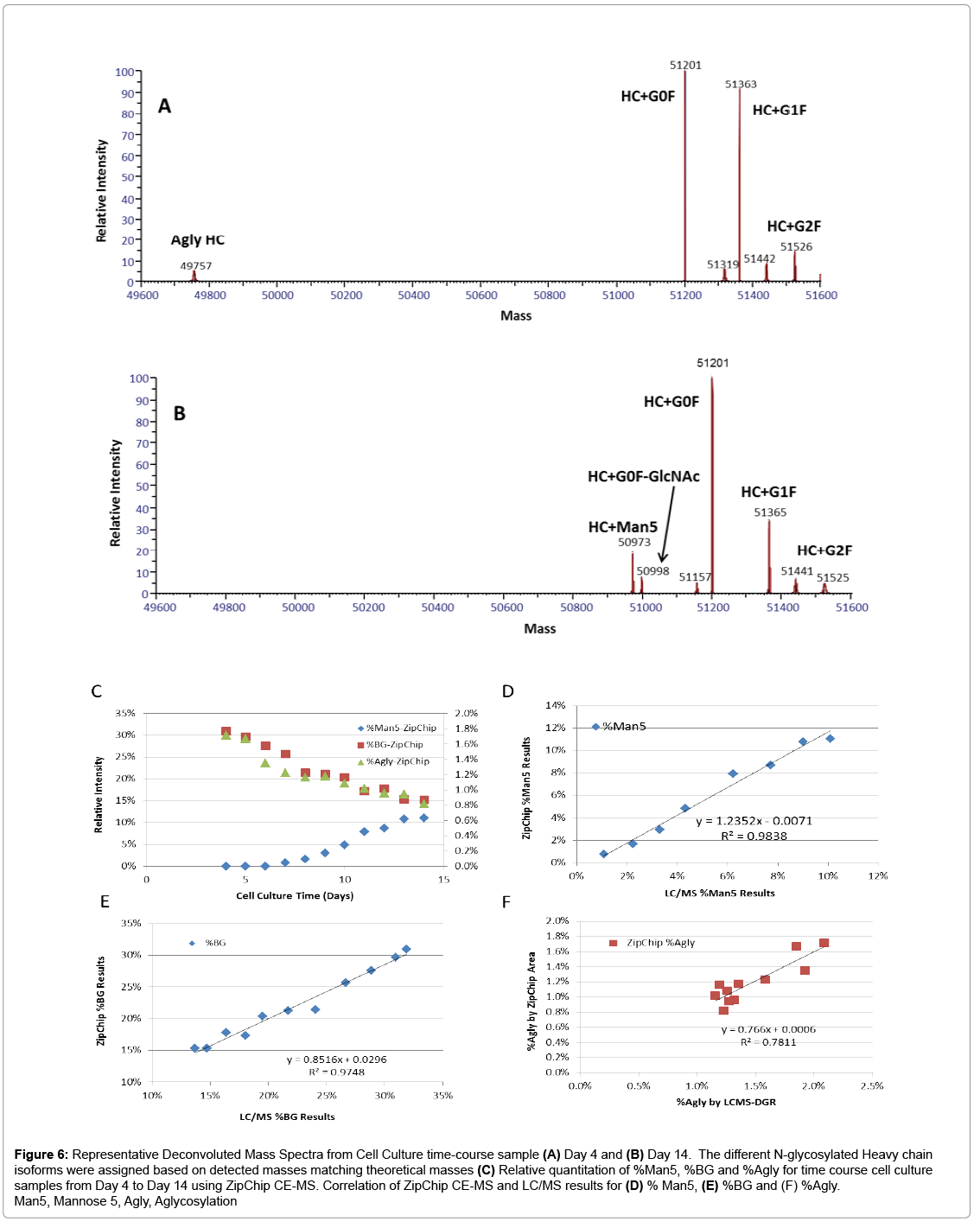
To demonstrate the capability of the ZipChip CE-MS for screening of cell culture quality attributes, a set of bioreactor time-course samples was analyzed by both ZipChip CE-MS and LC-MS using cell culture supernatant or ProA purified protein, respectively. Figure 6 shows the example of deconvoluted HC spectra from ZipChip CE-MS data of Day 4 and Day 14 cell culture supernatant. At Day 4, aglycosylated HC was detected and its level decreased over the course of culture duration (Figure 6A), while Man5 and G0F-GlcNAc were new species that appeared in Day 14 sample (Figure 6B). These trends were also confirmed by traditional RPLC-MS assay. The levels of five glycoforms (Man5, G0F-GlcNAc, G0F, G1F, and G2F) as well as aglycosylated species were quantitated in the Day 4 to Day 14 samples. Minor and major glycoforms species were detected at levels ranging from 0.7% for Man5 to 45.4% for G0F. Figure 6C shows changes in %Man5, %Biantennary Galactosylation (%BG) and %AGly levels for these attributes measured on different days of bioreactor cell culture process. Figure 6D and 6E shows good correlation of quantitation results between two methods for %Man5 (R2=0.98), and %BG (R2=0.97). As the %agly level in these samples is in the range of 0.8-1.7%, which is at the lower end of accurate quantitation for both CE-MS and LC-MS methods, resulting correlation factor is relatively low (R2=0.78) (Figure 6F).
Figure 6: Representative Deconvoluted Mass Spectra from Cell Culture time-course sample (A) Day 4 and (B) Day 14. The different N-glycosylated Heavy chain isoforms were assigned based on detected masses matching theoretical masses (C) Relative quantitation of %Man5, %BG and %Agly for time course cell culture samples from Day 4 to Day 14 using ZipChip CE-MS. Correlation of ZipChip CE-MS and LC/MS results for (D) % Man5, (E) %BG and (F) %Agly. Man5, Mannose 5, Agly, Aglycosylation
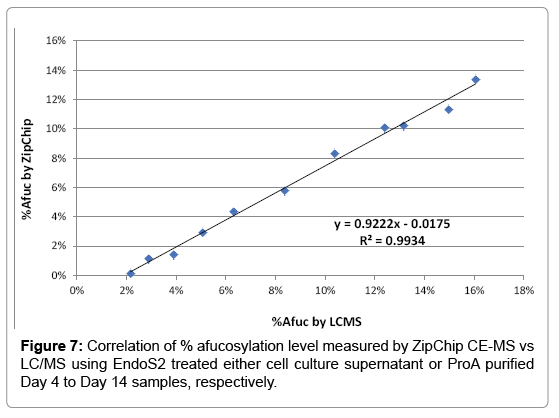
Afucosylation level can affect the effector function of mAb, which is a critical quality attributes to monitor during mAb development [4]. GlycINATOR enzyme (EndoS2) converts the N-glycosylated heavy chain species into two forms, one heavy chain attached with a GlcNAc and another one with a core-fucosylated GlcNAc, in addition to presence of the aglycosylated heavy chain. The RP-LC/MS based workflow to quantitate afucosylation using ProA purified mAb has been reported [18]. We herein tested the strategy of EndoS2 direct treatment of cell culture samples followed by ZipChip CE-MS analysis. Afucosylation levels determined by ZipChip CE-MS method from Day 4 to Day 14 were compared with results obtained from RP-LCMS method. Both methods showed very good correlation with R2=0.99 (Figure 7). This result demonstrated the capability of CE-MS method for rapid afucosylation monitoring from cell culture supernatant samples as well.
To make an initial method’s assessment between ZipChip CE-MS and established RPLC-MS assay, cell culture sample and corresponding ProA sample were analyzed multiple times. As shown in Table 1, comparable data for percent of Mannose 5, G0F, G1F and G2F glycoforms and %BG index were obtained between two methods using slightly different starting concentrations of cell culture supernatant and ProA purified sample, respectively. Two-fold dilutions of cell culture sample with null cell culture media down to 0.75 mg/mL revealed comparable and consistent data measurement of %G0F, %G1F, %G2F and %BG quantitation, but higher variation for the least abundant glycoform i.e., Mannose 5 (data not shown). Considering that titer concentration above 1.5 mg/mL are typically achievable at early days of mAb bioreactor process, ZipChip CE-MS can be used to monitor changes of minor glycoforms at approximately 1% relative abundance, which is comparable to the detection sensitivity of RPLC-MS using purified protein. Thus, this technique can be applied as a fast screening tool for monitoring glycosylation changes in cell culture samples.
| Sample Description | Average | Std Dev | RSD | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| LC/MS (N= 6) | %Man5 | %BG | G0F | G1F | G2F | %Man5 | %BG | G0F | G1F | G2F | %Man5 | G2F | %BG |
| Harvest after Pro A step1mg/mL | 1.7% | 16.3% | 67.3% | 23.1% | 3.8% | 0.2% | 0.2% | 0.4% | 0.2% | 0.1% | 9.9% | 3.8% | 1.1% |
| Zip Chip (N=4) | |||||||||||||
| Harvest1.5mgml | 1.1% | 16.0% | 68.6% | 23.4% | 3.6% | 0.2% | 0.5% | 1.1% | 0.9% | 0.2% | 14.0% | 4.6% | 3.1% |
Table 1: Comparison of the ZipChip CE-MS and LC-MS for mAb N-glycosylation quantitation. The ZipChip CE-MS data were obtained from four replicates of cell culture sample at 1.5 mg/mL. The RP-LC/MS results were obtained from six replicates using ProA purifed sample at 1 mg/mL. BG,biantennary galactosylation, % BG=[2*%G2F+%G1F]/[2*(G0F+G1F+G2F)].
Discussion
In this work, we report a fast and simple ZipChip CE-MS method to monitor protein N-glycosylation quality attributes and titer by directly analyzing cell culture supernatant without sample purification required. This approach allows multiple product quality information in a single analysis, such as determination of mAb HC and LC isoform masses, and titer concentration, as described here. Analogous information is generally carried out by several methods, for example, RP-LC-MS for mass determination, ELISA-based Octet or UV UPLC for protein concentration. Additionally, sample turnaround by ZipChip CE-MS can be achieved within 1.5 h or less for described workflows, including sample preparation, CE-MS analysis and data processing time. The reduced mass analysis using ZipChip CE-MS for nonpurified cell culture supernatant samples allows speed and simplicity required at the early stage of cell line screening, process development and batch manufacturing. This approach can be used for detection of undesired shifts in product quality that may lead to adjustment of related manufacturing parameters to ensure process being within the expected ranges. With the future implementation of automated sample loading, its application could be expanded as a new analytical tool for at-line and in-line cell culture performance monitoring.
Acknowledgements
We thank Dr. Yinyin Li for kindly providing heavy labeled monoclonal antibody for titer determination. We thank Dr. Seth Madren and Dr. Linda Yi for helpful discussions.
References
- Ecker DM, Jones SD, Levine HL (2015) The therapeutic monoclonal antibody market. MAbs 7: 9-14.
- Pais DA, Carrondo MJ, Alves PM, Teixeira AP (2014)Towards real-time monitoring of therapeutic protein quality in mammalian cell processes. Curr Opin Biotechnol 30: 161-167.
- Boyd PN, Lines AC, Patel AK (1995)The effect of the removal of sialic acid, galactose and total carbohydrate on the functional activity of Campath-1H. Mol Immunol 32: 1311-1318.
- Shinkawa T, Nakamura K, Yamane N, Shoji-Hosaka E, Kanda Y, et al. (2003)The absence of fucose but not the presence of galactose or bisecting N-acetylglucosamine of human IgG1 complex-type oligosaccharides shows the critical role of enhancing antibody-dependent cellular cytotoxicity. J Biol Chem 278: 3466-3473.
- Chung S, Quarmby V, Gao X, Ying Y, Lin L, et al. (2012)Quantitative evaluation of fucose reducing effects in a humanized antibody on Fcgamma receptor binding and antibody-dependent cell-mediated cytotoxicity activities. MAbs 4: 326-340.
- Goetze AM, Liu YD, Zhang Z, Shah B, Lee E, et al. (2011)High-mannose glycans on the Fc region of therapeutic IgG antibodies increase serum clearance in humans. Glycobiology 21: 949-959.
- Padler-Karavani V, Yu H, Cao H, Chokhawala H, Karp F, et al. (2008)Diversity in specificity, abundance, and composition of anti-Neu5Gc antibodies in normal humans: potential implications for disease. Glycobiology 18: 818-830.
- Bosques CJ, Collins BE, Meador JW (2010)Chinese hamster ovary cells can produce galactose-alpha-1,3-galactose antigens on proteins. Nat Biotechnol 28: 1153-1156.
- Pacis E, Yu M, Autsen J, Bayer R, Li F (2011)Effects of cell culture conditions on antibody N-linked glycosylation--what affects high mannose 5 glycoform. Biotechnol Bioeng 108: 2348-2358.
- Schiestl M, Stangler T, Torella C, Cepeljnik T, Toll H, et al. (2011) Acceptable changes in quality attributes of glycosylated biopharmaceuticals. Nat Biotechnol 29: 310-312.
- Konno Y, Kobayashi Y, Takahashi K, Takahashi E, Sakae S, et al. (2012) Fucose content of monoclonal antibodies can be controlled by culture medium osmolality for high antibody-dependent cellular cytotoxicity. Cytotechnology 64: 249-265.
- Higgins E (2010)Carbohydrate analysis throughout the development of a protein therapeutic. Glycoconj J 27: 211-225.
- Dong J, Migliore N, Mehrman SJ, Cunningham J, Lewis MJ, et al. (2016) High-Throughput, Automated Protein A Purification Platform with Multiattribute LC-MS Analysis for Advanced Cell Culture Process Monitoring. Anal Chem.
- Burnina I, Hoyt E, Lynaugh H, Li H, Gong B (2013) A cost-effective plate-based sample preparation for antibody N-glycan analysis. J Chromatogr A 1307: 201-206.
- Doherty M, Bones J, McLoughlin N, Telford JE, Harmon B, et al. (2013)An automated robotic platform for rapid profiling oligosaccharide analysis of monoclonal antibodies directly from cell culture. Anal Biochem 442: 10-18.
- Henninot A, Terrier A, Charton J, Urbain R, Fontayne A, et al. (2015) Characterization of monoclonal antibodies by a fast and easy liquid chromatography-mass spectrometry time-of-flight analysis on culture supernatant. Anal Biochem 491: 52-54
- Redman EA, Batz NG, Mellors JS, Ramsey JM (2015) Integrated microfluidic capillary electrophoresis-electrospray ionization devices with online MS detection for the separation and characterization of intact monoclonal antibody variants. Anal Chem 87: 2264-2272.
- Liu S, Zang L (2016) Rapid quantitation of monoclonal antibody N-glyco-occupancy and afucosylation using mass spectrometry. Anal Biochem 509: 142-145.
Relevant Topics
Recommended Journals
Article Tools
Article Usage
- Total views: 6636
- [From(publication date):
April-2017 - Aug 18, 2025] - Breakdown by view type
- HTML page views : 5338
- PDF downloads : 1298