Mutations in the Insect Transcriptome.
Received: 05-May-2016 / Accepted Date: 06-May-2016 / Published Date: 09-May-2016 DOI: 10.4172/2161-0681.1000e122
Keywords: Moth; Chemosensory protein; RNA; Proteome; Pheromone
313653Introduction
It is just the way imposed by the genetic code in the deep inner core of tissue cells in all organisms. A series of defined triplets in the DNA encodes for a precise combination of amino acids at the protein level. ATGGATTGCGTG (DNA or gene) changes to AUGGAUUGCGUG (RNA, single gene product) to encode Methionine, Aspartate, Cysteine and Valine amino acid residue, building thereby a crucial motif for the protein function. The motif cannot be changed without disrupting the protein function, which is directly related to the structure of that protein. So functioning of the “first place” life molecular element, i.e. “protein”, resides in the synthesis of a faithful copy of the product encoded in the genetic material, i.e. “DNA”. One piece of DNA (gene) codes one single piece of protein motif. From DNA-to-RNA-to protein, there is no mistake allowed. Any frameshift mutation or single base substitution that is disruptive to the genetic code, thus altering the number of bases and their position in the original sequence is not allowed by nature. There is no mistake possible and if it does, the result is a pathological disease [1-2].
To prevent any error in the protein translation process from gene and RNA, eukaryotic (and prokaryotic) cells have developed sophisticated proofreading mechanisms, i.e. specific enzymes called RNA polymerases that are capable of backtracking thereby minimizing the number of single nucleotide base mutations. This ensures accuracy in the transition from gene, RNA to protein [3]. However, this does not exclude the possibility that the cell can eventually switch the protein function thanks to single point mutations in key structural features of that protein.
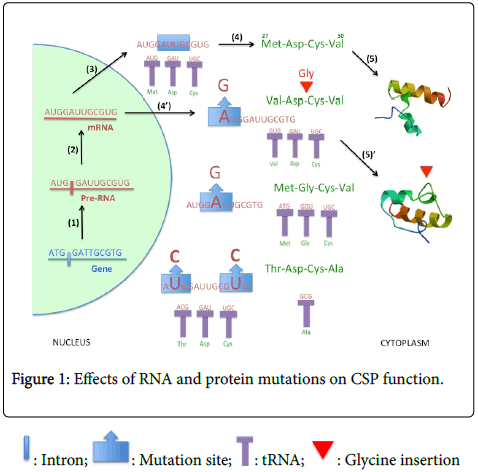
Other specific enzymes, namely adenosine deaminase acting on RNA (ADAR) and adenosine deaminases acting on tRNA (ADAT), are known to mediate A-to-G and U-to-C conversion, respectively, not only in human, insects and plants, but also in yeasts and bacteria [4-5]. Therefore, apparently in all various organisms, RNA can change and eventually codes for a new protein bearing novel function. In the brain of insects such as the fruitfly Drosophila melanogaster and the German cockroach Blatella germanica, it has been shown that edited RNA transcripts encode for voltage-dependent sodium channels with different gating properties [6]. Interestingly, however, editing of RNA transcripts is not limited to brain ion channels, but apparently expands also on some many more other gene families [7]. The recent discovery of RNA (and protein) mutations in the chemosensory protein (CSP) gene family in various tissues of the silkworm moth Bombyx mori, particularly in the female pheromone gland, strongly challenges the dogmatic concept that mutation is lethal and does not serve any possibility for a change in protein function [8-9]. The mutations highlighted in B. mori CSPs (Glycine insertion, Cysteine substitutions, etc) are pinpointed in the key features of the protein structure, strongly arguing for CSP-RNA nucleotide substitutions through or mediated via RNA editing to produce new proteins with novel functions in pheromone synthesis [10] (Figure 1).
The gene is initially transcribed into precursor messenger RNA (pre-mRNA), which is further processed (splicing of introns) to lead to mRNA molecule (steps 1 & 2). The mature mRNA is exported out the nucleus and translated into protein following the genetic code (step 3). This leads for instance in a motif such as Methionine-Aspartate- Cysteine-Valine (step 4) as found in silkworm moth Bombyx mori chemosensory protein-1 (BmorCSP1, AY766376, AF509239, JQ085850-JQ085858, JQ253592-JQ253630; BmorCSP1, AAM34276, AAV34688, 2JNT_A). The composition of the mature protein is unchanged and forms an alpha-helical profile typical for CSP proteins (5). However, eventually mRNA is edited thanks to ADAR and/or ADAT adenosine deaminase enzymes, converting A-to-G and/or U-to- C on specific key base locations. This leads to conversion of protein motif from Met-Asp-Cys-Val to Val-Asp-Cys-Val, Met-Gly-Cys-Val and/or Thr-Asp-Cys-Ala mutant peptides (4’). Using modeling (SWISS-MODEL) shows that such pinpointed amino acid mutations have no effects on the protein structure (2JNT_A), but insertion of Gly near Cys in the protein significantly affects the alpha-helical profile on crucial binding sites (5’). The succession of the two mutation events (first RNA, then protein) might therefore lead to a CSP with different binding affinities and thereby new functional role.
References
- Crick FHC, Barnett L, Brenner S, Watts-Tobin RJ (1961) General nature of the genetic code for proteins. Nature 192: 1227-1232.
- Veltman JA, Brunner HG (2012) De novo mutations in human genetic disease. Nature Rev Genet 13: 565-575.
- Wang D, Bushnell DA, Huang X, Westover KD, Levitt M, et al. (2009) Structural basis of transcription: backtracked RNA polymerase II at 3.4 angstrom resolution. Science 324: 1203-1206.
- Maas S, Gerber AP, Rich A (1999) Identification and characterization of a human tRNA-specific adenosine deaminase related to the ADAR family of pre-mRNA editing enzymes. ProcNatlAcadSci USA 96: 8895-8900.
- Bass BL (2002) RNA editing by Adenosine Deaminases that Act on RNA. Annu Rev Biochem 71: 817-846.
- Song W, Liu Z, Tan J, Nomura Y, Dong K (2004) RNA editing generates tissue- specific sodium channels with distinct gating properties. J BiolChem 279: 32554- 32561.
- St Laurent G, Tackett MR, Nechkin S, Shtokalo D, Antonets D, et al. (2013) Genome-wide analysis of A-to-I RNA editing by single molecule sequencing in Drosophila. Nat StructMolBiol 20: 1333-1339.
- Picimbon JF(2003) Biochemistry and evolution of CSP and OBP proteins. In: Blomquist GJ, Vogt RG (Eds) Insect Pheromone Biochemistry and Molecular Biology, The Biosynthesis and Detection of Pheromones and Plant Volatiles. Elsevier Academic Press, San Diego, London, pp. 539-566.
- Picimbon JF (2014) RNA mutations in the moth pheromone gland. RNA Dis 1: e240.
- Xuan N, Bu X, Liu YY, Yang X, Liu G X, et al. (2014) Molecular evidence ofRNA editing in Bombyx chemosensory protein family. PLoS ONE 9: e86932.
Citation: Picimbon JF (2016) Mutations in the Insect Transcriptome. J Clin Exp Pathol 6:e121. DOI: 10.4172/2161-0681.1000e122
Copyright: © 2016 Picimbon JF. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Select your language of interest to view the total content in your interested language
Share This Article
Recommended Journals
Open Access Journals
Article Tools
Article Usage
- Total views: 13410
- [From(publication date): 6-2016 - Aug 20, 2025]
- Breakdown by view type
- HTML page views: 12343
- PDF downloads: 1067

 : Intron;
: Intron;  : Mutation site;
: Mutation site;  : tRNA;
: tRNA;  : Glycine insertion
: Glycine insertion