Research Article Open Access
Polymorphisms in DNA Repair Gene XRCC1 (Arg194Trp) and (Arg399Gln) and their Role in the susceptibility of Bacterial Meningitis
Daniele Maria Lopes Pinheiro1, Fabrícia Lima Fontes1, Ana Helena Sales de Oliveira1, Leonam Gomes Coutinho1, Thayse Azevedo da Silva1, Stephen L. Leib2 and Agnez-Lima LF1*1Departamento de Biologia Celular e Genética, Universidade Federal do Rio Grande do Norte, UFRN, Natal, Brazil
2Institute for Infectious Diseases, University of Bern, Friedbuehlstrasse 51, CH-3010 Bern, Switzerland
- *Corresponding Author:
- Agnez-Lima LF
Laboratório de Biologia Molecular e Genômica
-LBMG, Departamento de Biologia Celular e Genética
Centro de Biociências-UFRN Campus
Universitário, Lagoa Nova Natal - RN, Brazil
Tel: 55 84 32119209
E-mail: lfagnez@gmail.com
Received Date: December 18 , 2015 Accepted Date: January 08, 2016 Published Date: January 28, 2016
Citation: Pinheiro DML,Oliveira HS, Coutinho LG, Silva TA, Leib SL, et al. (2016) Polymorphisms in DNA Repair Gene XRCC1 (Arg194Trp) and (Arg399Gln) and their Role in the susceptibility of Bacterial Meningitis. J Meningitis 1:105. doi: 10.4172/2572-2050.1000105
Copyright: © 2016 Pinheiro DML, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Journal of Meningitis
Abstract
Meningitis is a contagious infectious disease with high rates of mortality. Most pathogenic microbes in humans have the ability to cause bacterial meningitis. However, the most common pathogens are Streptococcus pneumoniae, Neisseria meningitidis and Haemophilus influenzae type b (Hib). It was found that the susceptibility to this infectious disease may be related to genetic characteristics of the host, such as the occurrence of single nucleotide polymorphisms (SNPs). In our previous work, association of SNPs in DNA repair genes with bacterial meningitis (BM) was demonstrated. In this study we evaluated two non-synonymous SNPs of the repair gene XRCC1 Arg194Trp (rs 1799782) and Arg399Gln (rs 25487) in patients with BM and health volunteers. The patient genotypes were investigated by PCR-RFLP. DNA damages were quantified using the genomic DNA with formamidopyrimidine DNA-glycosylase (FPG). Cytokines and chemokines were quantified from cerebrospinal fluid samples from BM patients. Concerning the SNP XRCC1 Arg194Trp, none association was found relation to BM. However, a higher frequency of heterozygous genotype for XRCC1 Arg399Gln was observed in the control group compared to the BM group (P=0.043; OR=0.426). DNA damage and cytokine/chemokines levels were not positively correlated with polymorphic genotypes. In conclusion, there is an indication that the SNP XRCC1 Arg399Gln could have a possible protective effect against BM.
Keywords
Bacterial meningitis; Streptococcus Pneumoniae ; Neisseria Meningitides; Base excision repair; XRCC1; SNPs; Inflammation
Introduction
Bacterial meningitis (BM) is characterized as a severe infection in the central nervous system (CNS), which compromises the meninges. Despite the effectiveness of the vaccination and treatment with antibiotics, negative outcomes are still associated with permanent neurological dysfunctions [1-3].
Most pathogenic microbes in humans are able to cause meningitis. However, the most common pathogens that cause BM are Streptococcus pneumoniae , Neisseria meningitidis and Haemophilus influenzae type b (Hib) [4]. Although different pathogens involved in the BM evolution are recognized by different toll-like receptors (TLRs), an overlap in the activation of the same inflammatory response which starts with the induction of MyD88 – a dependent pathway of nuclear factor kappa B (NF-κB) has been reported [5]. This finding corroborates the recent studies from our group showing a similar profile of inflammatory modulators during BM caused by S. pneumoniae and N. meningitidis [6,7].
BM starts with mucosa colonization in nasopharynx by commensal microorganisms. In this step, the immune defense system is important to combat the pathogens, avoiding the disease progression [8,9]. Thus, deficiency in the innate or acquired immune response may cause predisposition to disease [10,11]. Furthermore, strategies have been developed along the evolutionary process to enable some of these pathogens to overcome this natural barrier. The pathogens may invade the bloodstream, and after blood-brain barrier disruption they gained access to subarachnoid space causing the infection.
Several events involving cytokines, chemokines and oxidative stress have been observed during the inflammatory response contributing to brain dysfunction and this is mainly caused by the host immune response rather than by BM pathogen per se [12,13]. Genetic factors, such as single-nucleotide polymorphisms (SNPs) in DNA repair genes and in immune response genes have been shown to be associated with BM occurrence [11,14].
In previous work, polymorphisms in base excision repair (BER) genes, main involved in the repair of oxidized DNA damage, such as APE1, PARP-1 and OGG1 have been associated with regulation of immune response in BM [6,15]. However, none study of association between genetic variants in the X-ray repair protein crosscomplementing group 1 (XRCC1) gene, a key protein from BER which interacts with APE1 and OGG1, and meningitis inflammation has been performed.
XRCC1 is considered a key protein to interact with several enzymes as APE1 and OGG1, stimulating their activities. Several variants of XRCC1 have been described, the most common being located at codon 194 in exon 6, resulting from the substitution of arginine (Arg) with tryptophan (Trp). Another common variant is characterized by the exchange of arginine (Arg) for glutamine (Gln) at codon 399 located in exon 10 into BRCT1 domain [16,17]. The SNPs of XRCC1, Arg194Trp and Arg399Gln, have been investigated in several inflammatory diseases and cancer [18,19].
Considering the BER, as the main pathway involved in the repair of oxidative DNA damage in neurons [15], it is important to evaluate the association of genetic polymorphisms of XRCC1 with BM. In this study, we screened the occurrence of SNPs of XRCC1 Arg194Trp and Arg399Gln in the individuals with the meningitis disease. To the SNP XRCC1 Arg194Trp we did not find relation with the disease, but to the SNP XRCC1 Arg399Gln was found a higher frequency of the heterozygote genotype in the control group, indicating a possible protective effect against BM.
Materials and Methods
Ethics statement
Ethical approval for this study was given by Medical Ethics Committee at Giselda Trigueiro Hospital and by National Committee of Ethics (CONEP) with number 0052.1.051.000-05. In addition, informed consent was obtained from each patient participating in this research or the legal guardian in cases of minors/children.
Study subjects and samples
The study was conducted with a group of patients admitted at Giselda Trigueiro Hospital (HGT), reference for infectious diseases in Natal city, Rio Grande do Norte, state, Brazil. Diagnosis of each case was performed according to clinical signs and hospital routine tests, such as: Kernig’s sign and the Brudzinski’s sign, fever, headache, nausea and vomiting; cerebrospinal fluid (CSF) bacterial culture; gram staining; latex agglutination test in the CSF or blood; neutrophilic pleocytosis (≥ 500 cells/mm3) with predominance of polymorphonuclear granulocytes (PMN), evaluation the levels of protein (> 40 mg/dl) and glucose (< 40 mg/dl) and).
The exclusion criteria for this study were patients undergoing treatment with drugs (i.e. antibiotics) or afflicted with other diseases (i.e. AIDS), which could interfere in the expression of immune and/or inflammatory markers as cytokines. The control group was formed by healthy volunteers and those patients with a negative diagnosis for infectious disease and whose leukocyte levels were within the normal scale. For healthy volunteers a questionnaire was applied in order to assess the history to infectious and inflammatory diseases. In this last group of patients, CSF samples were not collected. The Blood samples were collected to obtain white blood cell counts (WBC) and inflammatory modulators such as cytokines and chemokines were analyzed.
The study involved 160 subjects (86 men and 74 women) split in two groups, BM group (n = 53) and control group (n = 107). From all the individual studied, 31 were less than 18 years old, 124 adults between 18 and 60 years old, 5 individuals aged over 60. In relation to BM group, 53 patients had positive diagnosis: 16 were diagnosed with S. pneumoniae, 7 with N. meningitidis , 6 with other pathogens, and 24 without specified etiology (Table 1). These data are consistent with other papers published by our group [6,7].
| Features | Number of cases |
|---|---|
| BM | 53 |
| Age | |
| 0-18 | 20 |
| 19-60 | 30 |
| >60 | 3 |
| Gender | |
| Male | 32 |
| Female | 21 |
| Pathogen | |
| S. pneumoniae | 16 |
| N. meningitidis | 7 |
| Other pathogens | 6 |
| Unidentified pathogens | 24 |
| Controls | 107 |
| Age | |
| 0-18 | 11 |
| 19-60 | 94 |
| >60 | 2 |
| Gender | |
| Male | 54 |
| Female | 53 |
Table 1: Clinical features of patients.
Samples processing
Blood samples were collected and processed by centrifugation at 4000 g for 3 minutes at 4°C in order to separate plasma from cells. Genomic DNA extraction was performed by the salting out procedure following the protocol described by Miller et al. [20]. The DNA integrity was observed on agarose gel 0.7%. Samples of CSF were collected immediately following lumbar puncture and centrifuged at 720 g for 5 min. Supernatants were frozen and stored at -80°C before any further procedure. Blood samples were obtained from all patients and controls, however, CSF were only obtained from patients who were submitted to LP as diagnosis routine once lumbar puncture is an invasive procedure.
XRCC1 Arg194Trp and Arg399Gln genotyping: The genotypes of the polymorphisms XRCC1 Arg194Trp and Arg399Gln were detected from the amplification of DNA by the method of polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP). Primers to each polymorphisms XRCC1 Arg194Trp (rs1799782) and Arg399Gln (rs25487) were obtained from Wu et al. [18] and Sturgis et al. [21], respectively. Table 2 shows primer sequences. The PCR-RFLP technique was validated after repeated with 20% from samples, beyond the success of genotyping above 80%.
| Genetic Polymorphism |
Exon | Primer sequence | Annealing Temp. (°C) | Restriction enzyme | Genotype | Fragments(bp) |
|---|---|---|---|---|---|---|
| XRCC1 Arg194Trp (rs1799782) |
6 | F 5’GTTCCGTGTGAAGGAGGAGGA3’ R 5’CGAGTCTAGGTCTCAACCCTACTCACT3’ (138 pb) |
58.7 0C | PvuII | Arg/Arg Arg/Trp Trp/Trp |
138 138+75+63 75+63 |
| XRCC1 Arg399Gln (rs 25487) |
10 | F 5’CAGTGGTGCTAACCTAATC3’ R 5’AGTAGTCTGCTGGCTCTGG3’ (871 pb) |
58 0C | Ncil | Arg/Arg Arg/Gln Gln/Gln |
461+278+132 593+461+278+132 593+278 |
Table 2: SNPs in DNA repair genes, primers and conditions used for genotyping.
PCR-RFLP products were generated using 100 ng of genomic DNA and 25 μl of PCR reaction containing specific buffer 1X, 10 pM of each primer, 2.0 mM MgCl2, 0.3 mM of each dNTP, and 3.6 U of Taq polymerase. The PCR program had an initial denaturation step of 5 min at 94°C followed by 30 cycles of 1 min at 95°C, annealing of 1 min and 30 s for codon 194 and of 2 min for codon 399 (temperatures given in Table 2), elongation for 1 min at 72°C and a final extension step of 10 min at 72°C. PCR products were run on a 1% agarose gels and visualized with SYBR green. The amplified fragments were digested with appropriate restriction endonucleases PvuII and Ncil for SNPs XRCC1 Arg194Trp and Arg399Gln, respectively (Table 2) and the resulting cleavage products were separated by electrophoresis on 8% polyacrylamide gel for 1 h at 100 V and detected with silver staining according to the protocol described by Sanguinetti et al. [22].
Detection of FPG sensitive sites
The genomic DNA from patients was submitted to treatment with formamidopyrimidine DNA glycosylase (FPG) (New England Biolabs) as described by da Silva et al. [6]. This enzyme is a functional analog of OGG1 in bacteria and acts as a bifunctional glycosylase that recognizes oxidized damage as 8-oxodG due to its β-lyase activity [23]. The sensitive sites to FPG were measured by densitometry of DNA, seen after electrophoresis in 1% agarose gel with ImageJ 1.42q software (Wayne Rasband National Institutes of Health, USA). The number of double strand breaks (DSBs) were calculated as described by D’Ambrosio et al. [24], using Poisson distribution. These data were associated with presence or absence of the SNPs in the studied group.
Measurement of inflammatory mediators
The cytokine and chemokine measurements were performed using the CSF of patients with BM. Inflammatory modulators levels were measured by multiplex suspension array using the Bio-Plex 200 suspension array system (Bio-Rad, Hercules, CA, USA) and using microsphere based multiplex assays. Human Cytokine/Chemokine Panel (MPXHCYTO-60k, Millipore) was used including a set of 12 cytokines and chemokines (TNF-α, IL-6, IL-1β, IL-2, INF-γ, IL-10, IL-1Ra, MIP-1α/CCL3, MIP-1β/CCL4, MCP-1/CCL2, G-CSF and IL-8/CXCL8). The samples were processed and measured in duplicates and according to the manufacturer’s instructions [6,7]. This assay allowed for the quantification of cytokines and chemokines over a broad range of 3.2–10.000 pg/ml, using Bio-Plex manager 4.01 software. Due to the low quantity of biological material, it was not possible to make the determination in all the patients.
Statistical analysis
Statistical analysis was performed using GraphPad-Prism5 software and WINPEPI version 11.15 (2011). First, the allele and genotype frequencies to BM group and control group were determined by direct gene counting. Genotypic distributions in Hardy-Weinberg equilibrium were analyzed by the classical method of X2-test (twotailed) using Helix SVS program. Logistic regression analyses were used to calculate odds ratios (OR) with 95% confidence interval (CI) and corresponding P-values for the association between the susceptibility to infectious disease (BM) and each variant. The effects of sex and age were also evaluated in this analysis. Comparisons obtained statistical significance and differences in proportions in the genotypic frequencies between BM and control group were analyzed using the Chi square test (two-tailed) or Fisher’s exact test. STATA software (version 11.0; Stat Corporation, College Station, Texas, USA) was used to conduct this statistical analysis.
The study power was calculated using WINPEPI version 5.4 software (2008). Differences in the levels of inflammatory modulators and DNA damage level were analyzed using the non-parametric t-test (Mann-Whitney two-tailed). Values of P ≤ 0.05 were considered statistically significant.
Results
Analysis of genetic polymorphisms
Fifty-three BM patients and 107 control subjects were tested for polymorphisms XRCC1 Arg399Gln and Arg194Trp. The genotype distribution for both SNPs was in the Hardy-Weinberg equilibrium. For the SNP XRCC1 Arg399Gln, the frequency of heterozygote (Arg/ Gln) was significantly higher in controls (P = 0.043) than in BM patients. The frequency of homozygote (Gln/Gln) was more frequent in the control group although the difference was not significant. The frequency of Gln allele was higher in the control group (Table 3). Significant differences between groups were not observed, neither for the genotypes nor the alleles of SNP XRCC1 Arg194Trp (P = 0.68) (Table 4).
| Genotype | Control group n (%)=107 | BM group n (%)=51 | P | OR (95% CI) | Adjusted Pa | Adjusted OR (95% CI)a |
|---|---|---|---|---|---|---|
| XRCC1 Arg399Gln | ||||||
| Arg/Arg | 50 (47.17) | 35 (68.63) | 0.043* | 1 | 0.048* | 1 |
| Arg/Gln | 48 (44.34) | 14 (24.45) | 0.426 (0.204 – 0.889) | 0 | 0.415 (0.196 – 0.878) | |
| Gln/Gln | 9 (8.49) | 2 (3.92) | 0.317 (0.065 – 1.560) | 0.356 (0.707 – 1.795) | ||
| Gln allele frequency | 0.3 | 0.17 |
Table 3: Allelic and genotypic frequencies of BM group compared to control group for XRCC1 Arg399Gln SNP. BM: bacterial meningitis; OR: odds ratio; CI: confidence interval. Data were analyzed by multivariate logistic regression. *Statistical significance (P<0.05). The reference group in each of the analysis was the most prevalent genotype. A data adjusted for age and gender.
| Genotype | Control group n (%)=105 | BM group n (%)=53 | P | OR (95% CI) | Adjusted Pa | Adjusted OR (95% CI)a |
|---|---|---|---|---|---|---|
| XRCC1 Arg194Trp | ||||||
| Arg/Arg | 95 (90.48) | 49 (92.45) | 0.68 | 1 | 0.94 | 1 |
| Arg/Trp | 10 (9.52) | 4 (7.55) | 0.776 (0.231 – 2.600) | 0.954 (0.273 – 3.332) | ||
| Trp/Trp | 0 (0) | 0 (0) | ||||
| Trp allele frequency | 0.04 | 0.04 |
Table 4: Allelic and genotypic frequencies of BM group compared to control group for XRCC1 Arg194Trp SNP. BM: bacterial meningitis; OR: odds ratio; CI: confidence interval. Data were analyzed by multivariate logistic regression. *Statistical significance (P<0.05). The reference group in each of the analysis was the most prevalent genotype. A data adjusted for age and gender.
Detection of DNA damage
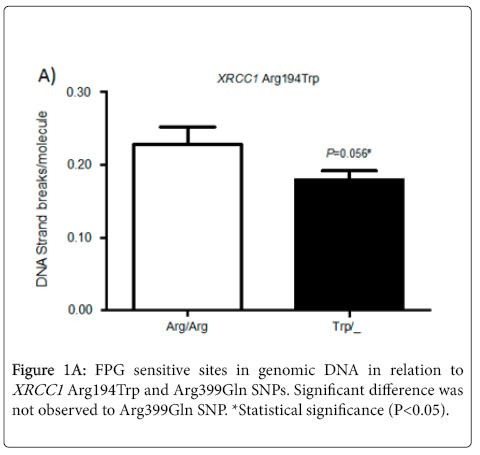
FPG sensitive sites were quantified in genomic DNA obtained from individuals included in this study. The samples were analyzed in relation to the presence or absence of the variant allele. Although there was no significant difference, but a borderline value (P = 0.056) was observed for SNP XRCC1 Arg194Trp, with a small reduction in FPG sensitive sites (Figure 1A).
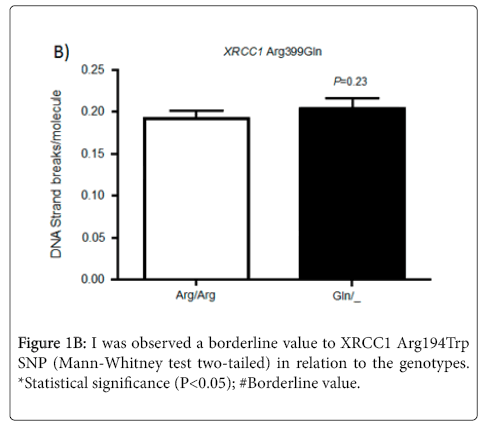
No statistical difference in the levels of DNA damage was observed for individuals with the variant or wild-type alleles for SNP XRCC1 Arg399Gln.
Considering no significant difference was observed between cases and control (data not shown), all individuals was grouped and the analysis was done take into consideration the presence or absence of the variant allele.
Discussion
Bacterial meningitis has been known to be an important cause of mortality and morbidity. The disease evolution is mainly influenced by the host immune response determined by several genes that regulate the intensity of the inflammatory response to infection [8-11].
Measurement of cytokines and chemokines in patients with BM
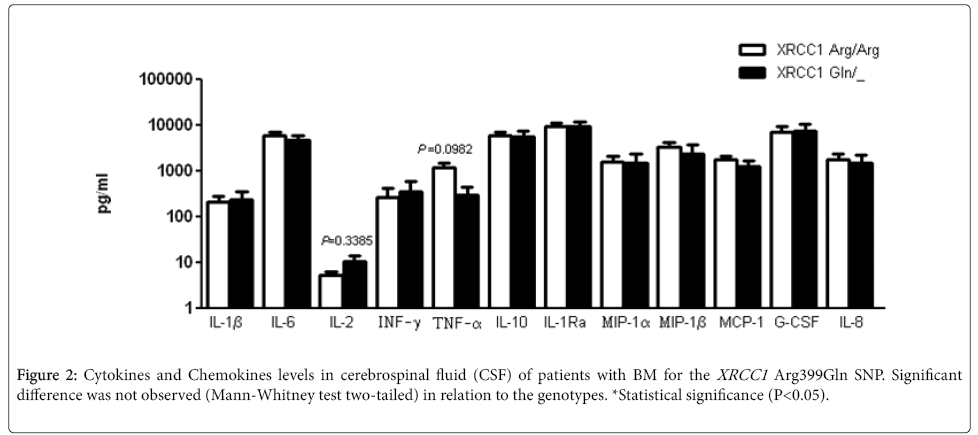
The main cytokines and chemokines involved in the BM inflammatory response were examined in the CSF samples from patients. BM patients carrying XRCC1 Gln allele did not showed statistical difference in the cytokines and chemokines levels (Figure 2). For SNP XRCC1 Arg194Trp, no statistical difference was also observed in the concentration of cytokines and chemokines (data not shown).
In previous work, we obtained data showing association of the SNPs APE1 Asn148Glu, OGG1 Ser326Cys, and PARP-1 Val762Ala with BM. Furthermore, the SNP in APE1 also showed effect on expression of some cytokines as IL-6 and IL-8 [6]. Since XRCC1 is a key enzyme in BER pathway, being a scaffold protein that is responsible to recruit other proteins as APE1 and OGG1 [16,17], similar data could be expected. However, our data showed that SNP XRCC1 Arg194Trp is not associated with BM. In addition, for the SNP XRCC1 Arg399Gln, we found a possible protective effect against BM once the Gln variant allele was more present in the control group, consequently significant difference in allelic and genotypic frequencies was observed. Although, we used a low number of patients in our work we obtained allele and genotypic frequencies similar to the ones reported in the literature [25-27].
Other authors also found no association of SNP XRCC1 Arg194Trp with rheumatoid arthritis and Alzheimer’s disease [19,28]. The function of this SNP has not been established yet, but studies have suggested a protective role against cancer. Some experimental data indicate that the Arg194Trp variant is associated with increased genomic stability in response to genotoxic agents [29,30]. In agreement with these data, in our work we observed a trend in the reduction of FPG sensitive sites in the genomic DNA from BM patients, which may be associated to the role of XRCC1 in stimulating the OGG1 activity [17].
In relation to the SNP XRCC1 Arg399Gln, the literature data show evidences of its association with rheumatoid arthritis, Parkinson’s disease, increased in breast cancer risk, among others [19,31,32]. Some groups have tried to explain their results by ethnic data because populations with different ancestry may cause differences in observed frequencies (Figure 1B) However, the ancestral influence on the current Brazilian population is the result of effects such as, migration and miscegenation, mainly European, African and Amerindian populations and then resulting in a pattern of linkage disequilibrium. So, it is not possible a detailed analysis of the data found in ethnic closed groups for most of the studies conducted in Brazil, which does not have a specific ethnic pattern [33].
Some authors have obtained evidence that the SNP XRCC1 Arg399Gln may affect the XRCC1 functions. The residue 399 is located in BRCT1 domain of the XRCC1 protein, which is important for protein-protein interaction. Experimental data indicated that the Gln variant affect the capability of BRCT1 domain to interact with other proteins, reducing the ability of XRCC1 to coordinate BER [34]. In cells expressing the XRCC1 Gln variant, differences in the intranuclear localization or in the ability to recruit repair enzymes to the damage sites were not observed in relation to XRCC1 wild type protein. However, a reduced stability of the repair foci was observed in cells expressing Gln variant. Furthermore, differences in repair profile were observed after treatment of the cells with methyl methanesulfonate (MMS) or hydrogen peroxide, suggesting that this SNP may affect the repair under stress conditions [35]. Corroborating these data, in non-small cell lung cancer patients with polymorphism to Arg399Gln a lower 8-oxoG incision activity was observed in lung tissues, but not in leukocytes. This suggests that this SNP may influence the OGG1 activity in tissues exposed to stress [36].
Despite XRCC1 interacting with OGG1 which stimulates its activity 2 to 3-fold and being important in the repair of 8-oxoG [17], we did not observe differences in the occurrence of DNA damage in relation to XRCC1 genotypes in our work. Since BM is a localized infection in CNS, and the genomic DNA was obtained from blood samples, our data corroborate with previous researches who propose that the SNP Arg399Gln may affect the repair function mainly in cells under stress [35,36]. In contrast to the commitment of the DNA repair functions, the SNP Arg399Gln has also been associated with a reduced risk of some types of cancer, showing a protective role [37]. To explain this contradictory data, Stern et al. proposed the hypothesis that the reduced repair activity of Gln variant could predispose cells to increased apoptosis or senescence, thus preventing the tumor progression.
In the context of BM, it is not clear how the Gln variant could be associated to the protection of the disease. It was demonstrated that monocytes are highly sensitive to oxidative stress due to a reduced level of XRCC1, PARP-1 and LigIII proteins. After the differentiation in macrophages and dendritic cells, the repair proteins levels are restored. The authors proposed that the deficiency in repair of monocytes may be important to the regulation of inflammatory response, since the selective killing of monocytes during oxidative stress induced by macrophage may act as a negative regulatory feedback, reducing the macrophage differentiation and avoiding the excess of oxidative stress during inflammation [38]. In this scenario, it could be important to investigate the effect of the XRCC1 Gln variant in monocytes/ macrophage differentiation process.
Cytokines have an important role in controlling infection. Their deficiency may cause predisposition to diseases but in excess or in the wrong tissues can cause serious complications as observed in BM [8-11]. Some BER enzymes as APE1, OGG1 and PARP-1 were implicated in the regulation of expression of some cytokines [6]. However, in this work we did not find any association between SNPs in XRCC1 with cytokines and chemokines expression. Despite this, the hypothesis that XRCC1 plays an important role in inflammation should not be ruled out, since a possible protective role of Gln allele was observed in our study. Considering that BM is a complex disease, the interaction between XRCC1 with other important proteins in modulating inflammatory response should also be evaluated, since XRCC1 is a scaffold protein important not only for BER, but also in nucleotide excision repair (NER) and non-homologous end joining (NHEJ) [39].
Conclusion
In conclusion, we found no association of the SNP XRCC1 Arg194Trp with the disease and an indication that the Gln XRCC1 allele of the SNP XRCC1 Arg399Gln may play a protective role for BM. However, the low frequency of cases of meningitis and the poor conditions of the public health system in Natal (Brazil) did not allow for a more precise stratification of patients. With this, our results are preliminary and which will be necessary to extend this study to a larger cohort in the future to confirm this finding to understand the mechanisms involved in this protection.
References
- Miranda J and Tunkel AR (2009) Strategies and new developments in the management of bacterial meningitis. Infect Dis Clin North Am 23:925-43.
- Thigpen MC, et al. (2011) Bacterial meningitis in the United States, 1998-2007. N Engl J Med 364:216-25.
- Van De Beek D, Brouwer MC, Thwaites GE, Tunkel AR (2012) Advances in treatment of bacterial meningitis. Lancet 380:1693-702.
- Kim KS (2010) Acute bacterial meningitis in infants and children. Lancet Infect Dis 10:32-42.
- Mogensen TH, Paludan SR, Kilian M, Ostergaard L (2006) Live Streptococcus pneumoniae, Haemophilusinfluenzae, and Neisseria meningitidis activate the inflammatory response through Toll-like receptors 2, 4, and 9 in species-specific patterns. J LeukocBiol 80:267-77.
- Da Silva TA, Fontes FL, Coutinho LG, De Souza FR, De Melo JT, et al. (2011) SNPs in DNA repair genes associated to meningitis and host immune response. Mutat Res 713:39-47.
- Coutinho LG, Grandgirard D, Leib SL and Agnez-Lima LF (2013) Cerebrospinal-fluid cytokine and chemokine profile in patients with pneumococcal and meningococcal meningitis. BMC Infect Dis 13:326.
- Carbonnelle E, Hill DJ, Morand P, Griffiths NJ, Bourdoulous S, et al. (2009) Meningococcal interactions with the host. Vaccine 27:B78-B89.
- Mook-Kanamori BB, Geldhoff M, Van Der Poll T, Van De Beek D (2011) Pathogenesis and pathophysiology of pneumococcal meningitis. ClinMicrobiol Rev 24: 557-91.
- Brouwer MC, De Gans J, Heckenberg SG, Zwinderman AH, Van Der Poll T, Van De Beek D (2009) Host genetic susceptibility to pneumococcal and meningococcal disease: a systematic review and meta-analysis. Lancet Infect Dis 9: 31-44.
- Sanders MS, Van Well GT, Ouburg S, Morre SA, Van Furth AM (2011) Genetic variation of innate immune response genes in invasive pneumococcal and meningococcal disease applied to the pathogenesis of meningitis. Genes Immun 12: 321-334.
- Leib SL, Kim YS, Chow LL, Sheldon RA, Tauber MG (1996) Reactive oxygen intermediates contribute to necrotic and apoptotic neuronal injury in an infant rat model of bacterial meningitis due to group B streptococci. J Clin Invest 98: 2632-2639.
- Prasad K, Rai NK, Kumar A (2012) Use of corticosteroids and other adjunct therapies for acute bacterial meningitis in adults. Curr Infect Dis Rep 14: 445-453.
- Netea MG and Van Der Meer JW (2011) Immunodeficiency and genetic defects of pattern-recognition receptors. N Engl J Med 364: 60-70
- Parsons JL, Dianov GL (2013) Co-ordination of base excision repair and genome stability. DNA Repair (Amst) 12: 326-333.
- Vidal AE, Boiteux S, Hickson ID, Radicella JP (2001) XRCC1 coordinates the initial and late stages of DNA abasic site repair through protein-protein interactions. EMBO J 20: 6530-6539.
- Marsin S, Vidal AE, Sossou M, Menissier-De Murcia J, Le Page F, et al. (2003) Role of XRCC1 in the coordination and stimulation of oxidative DNA damage repair initiated by the DNA glycosylase hOGG1. J BiolChem 278: 44068-44074.
- Wu MT, Chen SY, Wu TN, Hwang HY, Ho CK, et al. (2003) No association between polymorphisms of the DNA repair geneXRCC1 and cervical neoplasm risk. Environ Health Prev Med 8: 100-103.
- Yosunkaya E, Karakurt F, Cetin E, Ozgonenel L, Onaran I, et al. (2010) Rheumatoid arthritis risk associates with DNA repair gene XRCC1 Arg399Gln polymorphism in Turkish patients. RheumatolInt 32: 1265-1269.
- Miller SA, Dykes DD, Polesky HF (1988) A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res 16: 1215.
- Sturgis EM, Castillo EJ, Li L, Zheng R, Eicher SA, et al. (1999) Polymorphisms of DNA repair gene XRCC1 in squamous cell carcinoma of the head and neck. Carcinogenesis 20: 2125-2129.
- Sanguinetti CJ, Dias Neto E, Simpson AJ (1994) Rapid silver staining and recovery of PCR products separated on polyacrylamide gels. Biotechniques 17: 914-921.
- Frosina G (2006) Prophylaxis of oxidative DNA damage by formamidopyrimidine-DNA glycosylase. Int J Cancer 119: 1-7.
- D'ambrosio SM, Gibson-D'ambrosio RE, Brady T, Oberyszyn AS, Robertson FM (2001) Mechanisms of nitric oxide-induced cytotoxicity in normal human hepatocytes. Environ Mol Mutagen 37: 46-54.
- Duarte MC, Colombo J, Rossit AR, Caetano A, Borim AA, et al. (2005) Polymorphisms of DNA repair genes XRCC1 and XRCC3, interaction with environmental exposure and risk of chronic gastritis and gastric cancer. World J Gastroenterol 11: 6593-6600.
- Falagan-Lotsch P, Rodrigues MS, Esteves V, Vieira R, Amendola LC, et al. (2009) XRCC1 gene polymorphisms in a population sample and in women with a family history of breast cancer from Rio de Janeiro. Genet MolBiol 32:255-259.
- Yang SF, Xu YJ, Xie JG, Zhang ZX (2009) hOGG1 Ser326Cys and XRCC1 Arg399Gln polymorphisms associated with chronic obstructive pulmonary disease. Chin Med J (Engl) 122: 960-966.
- Dogru-Abbasoglu S, Aykac-Toker G, Hanagasi HA, Gurvit H, Emre M, Uysal M (2007) The Arg194Trp polymorphism in DNA repair gene XRCC1 and the risk for sporadic late-onset Alzheimer's disease. NeurolSci 28: 31-34.
- Jiang J, Liang X, Zhou X, Huang R, Chu Z, et al. (2010) DNA repair gene X-ray repair cross complementing group 1 Arg194Trp polymorphism on the risk of lung cancer: a meta-analysis on 22 studies. J ThoracOncol 5: 1741-1747.
- Ginsberg G, Angle K, Guyton K, Sonawane B (2011) Polymorphism in the DNA repair enzyme XRCC1: utility of current database and implications for human health risk assessment. Mutat Res 727: 1-15.
- Wu K, Su D, Lin K, Luo J, Au WW (2011) XRCC1 Arg399Gln gene polymorphism and breast cancer risk: a meta-analysis based on case-control studies. Asian Pac J Cancer Prev 12: 2237-2243.
- Gencer M, Dasdemir S, Cakmakoglu B, Cetinkaya Y, Varlibas F, et al. (2012) DNA repair genes in Parkinson's disease. Genet Test Mol Biomarkers 16: 504-507.
- Pena SD, Di Pietro G, Fuchshuber-Moraes M, Genro JP, Hutz MH, et al. (2011) The genomic ancestry of individuals from different geographical regions of Brazil is more uniform than expected. PLoS One 6: 17063.
- Monaco R, Rosal R, Dolan MA, Pincus MR, Brandt-Rauf PW (2007) Conformational effects of a common codon 399 polymorphism on the BRCT1 domain of the XRCC1 protein. Protein J 26: 541-546.
- Hanssen-Bauer A, Solvang-Garten K, Gilljam KM, Torseth K, Wilson DM, et al. (2012) The region of XRCC1 which harbours the three most common nonsynonymous polymorphic variants, is essential for the scaffolding function of XRCC1. DNA Repair (Amst) 11: 357-366.
- Janik J, Swoboda M, Janowska B, Ciesla JM, Gackowski D, et al. (2011) 8-Oxoguanine incision activity is impaired in lung tissues of NSCLC patients with the polymorphism of OGG1 and XRCC1 genes. Mutat Res 710: 21-31.
- Stern MC, Umbach DM, Van Gils CH, Lunn RM, Taylor JA (2001) DNA repair gene XRCC1 polymorphisms, smoking, and bladder cancer risk. Cancer Epidemiol Biomarkers Prev 10: 125-131.
- Bauer M, Goldstein M, Christmann M, Becker H, Heylmann D, Kaina B (2011) Human monocytes are severely impaired in base and DNA double-strand break repair that renders them vulnerable to oxidative stress. ProcNatlAcadSci U S A 108: 21105-21110.
- Hanssen-Bauer A, Solvang-Garten K, Akbari M, Otterlei M (2012) X-ray repair cross complementing protein 1 in base excision repair. Int J MolSci 13: 17210-1722.
Relevant Topics
- Acute Meningititis
- Advances in Meningitis Diagnosis
- Advances in Meningitis Treatment
- Contageous Meningitis
- Etiology of Meningitis
- Fungal meningitis
- Meningitis
- Meningitis Septicaemia
- Meningitis Statistics
- Meningococcal vaccine
- Meningoencephalitis
- Pneumococcal meningitis
- Streptococcus meningitis
- Tuberculosis Meningitis
- Viral Meningitis
Recommended Journals
Article Tools
Article Usage
- Total views: 11149
- [From(publication date):
June-2016 - Aug 19, 2025] - Breakdown by view type
- HTML page views : 10193
- PDF downloads : 956