The Clinical Value of Abnormal TBX15 Hypermethylation in HCC
Received: 05-Aug-2021 / Accepted Date: 19-Aug-2021 / Published Date: 26-Aug-2021 DOI: 10.4172/aot.s4.1000002
Abstract
Background: HCC is one of the most common and invasive malignant tumors in the world. The objective of this research was to explore the methylation of TBX15 and determine the clinical value of TBX15 Hypermethylation in Hepatocellular Carcinoma (HCC).
Methods: The methylation status of the TBX15 gene in 61 pairs of HCC patient tumor and paracancerous tissues was analyzed with bisulfite and methylation-specific PCR, and the association between TBX15 Hypermethylation and specific clinical features was determined.
Results: TBX15 Hypermethylation was associated with patient prognosis. Eighty percent of the tumor and associated paracancerous tissues were hypermethylated. The methylation levels of TBX15 in all 61 tumors were higher than those in the paired paracancerous tissues. The difference between the two tissue types was statistically significant (P<0.001). The methylation level of TBX15 was increased in tumor tissues with vascular invasion (P>0.05) and poor envelope integrity (P<0.05). Additionally, Hypermethylation of TBX15 was statistically associated with advanced cancer stage (P<0.05). The Area Under the Curve (AUC) value (0.98) indicated that TBX15 Hypermethylation had a high confidence value for diagnosing HCC. The Kaplan-Meier analysis suggested that TBX15 Hypermethylation was associated with low relapse-free survival rates. The Cox proportional hazards regression analysis indicated that TBX15 Hypermethylation was an independent factor affecting prognosis.
Conclusion: TBX15 Hypermethylation was related to malignant behavior and was identified as a potential indicator for poor prognosis. Therefore, TBX15 Hypermethylation can be used as a biomarker for the diagnosis and prognostication of HCCs.
Keywords: TBX15; DNA hypermethylation; Hepatocellular carcinoma; Biomarker; Hepatology
Introduction
Hepatocellular Carcinoma (HCC) is one of the most common and aggressive malignant tumors in the world, and it is also the third most common cause of death in the world [1,2]. The disease causes nearly 600,000 deaths worldwide every year, and its morbidity and mortality have been increasing in recent years. Although surgical treatment and medical management strategies for HCC have progressed, the overall prognosis of HCC patients is still not satisfactory because liver cancer patients undergoing treatment often suffer from terminal-stage cancer. The 5-year survival rate of HCC is approximately 17% [3,4]. Although the Alpha-Fetoprotein(AFP) tumor marker is routinely used in early screening for HCC, which has reduced the mortality of liver cancer, its sensitivity and specificity are still limited [5,6]. Therefore, discovering a valuable cellular biomarker specific for HCC and exploring new diagnosis and treatment strategies are crucial.
The alteration of DNA methylation is a key epigenetic event in cancer. Such alteration has been confirmed to play a vital role in cancer and other human diseases [7]. DNA methylation often occurs in Tumor Suppressor Genes (TSGs), and Hypermethylation can cause inappropriate transcriptional silencing [8,9], leading to the occurrence and development of cancer. Current research shows that the T-box gene family is a systematically conserved transcription factor family that regulates a variety of developmental processes [10] and the proliferation and differentiation of tissue-specific stem and progenitor cells [11,12]. The T-box gene family is divided into 5 subfamilies. The TBX15 gene belongs to the Tbx2 subfamily [13]. It is involved in a variety of biological processes, such as cell proliferation, differentiation, metastasis and apoptosis. Evidence shows that TBX15 affects cell proliferation and apoptosis via antiapoptotic effects in cancer [14]. Therefore, it is considered a TSG. For example, in ovarian cancer, immunohistochemistry analysis showed a negative correlation between the TBX15 methylation level and TBX15 protein expression [15]. TBX15 methylation is also significantly correlated with pathological stage, Gleason score and biochemical relapse in prostate cancer [16]. The studies of Gao [17], Zheng [18] and others have found that HCC tumor tissues display higher methylation of the TBX15 gene promoter than paracancerous tissues. However, there is no research on the relationship between TBX15 methylation and liver cancer clinical characteristics. Therefore, in this study, the methylation status of TBX15 in HCC and adjacent paracancerous tissues was determined to study the relationship between the methylation status and clinical characteristics of HCC patients. Finally, an evaluation of its clinical value is provided.
Materials and Methods
Tissue samples
Subjects were chosen from patients who underwent radical liver resection at the First Associated Hospital of Zhejiang University. The patients met the following criteria:
• The subject was undergoing radical resection for liver cancer.
• The subject was suffering from only HCC without any derived or secondary carcinomas.
• No radiotherapy or chemotherapy was administered before surgery.
All patients signed an informed consent form. The resected tumor tissue from the liver and its adjacent tissues were collected post-surgery. The paracancerous tissues were obtained from tissues surrounding the tumor within a radius of 1 cm. All tissues collected were stored in liquid nitrogen tanks at -80°C. The TNM classification method was used for clinical staging. Among the enrolled patients, 11 patients had stage I disease, 4 patients had stage II disease, 43 patients had stage III disease, and 3 patients had stage IV disease. In this study, the average age of the patients was 58.3 years with a standard deviation of 11.04. The follow-up time was from the end of the surgery to September 2020. The average follow-up time was 383.19 days with a standard deviation of 86.29. All but 3 patients kept in contact during the follow-up period. This study was approved by the Ethics Committee of the First Affiliated Hospital of Zhejiang University.
Experiments
DNA extraction: DNA was extracted from tissue samples, and a Nano drop 2000 (Thermo Fisher Scientific Co. Ltd) was used to test the quality of genomic DNA. The concentration of all samples was ≥ 20 ng/μL and the total amount was ≥ 1 μg. All DNA absorbance ratios at 260/280 nm were between 1.8 and 2.0. Agarose gel electrophoresis was used to detect the integrity of genomic DNA. The electrophoresis bands of all the samples were clearly visible without RNA contamination.
Bisulfite conversion and multiplex amplification: The DNA methylation level was analyzed with MethylTargetTM (Genesky Biotechnologies Inc., Shanghai, China), an NGS-based multiple- target CpG methylation analysis method. The CpG island between the 2K region upstream of Transcription Start Site (TSS) and the 1K region downstream of the first exon was analyzed with Gene CpG v1.0 software. The evaluation standards were as follows: observed/ expected ratio>0.60, percent C+ percent G>50.00% and length>200 bp. The search ultimately yielded two CpG island locations. Four targeted regions before the promoter region (ranging from TSS1500 to TSS200) were chosen and transformed into bisulfite-converted sequences. The design of primers for methylation experiments (Table 1) was performed using Methylation Fast Target V4.1 software. The primer standard characteristics were as follows: Tm: 58-62°C, primer length: 18-36 bp, product size range: 150-270, and minimum CpG site number: 10. Genomic DNA (400 ng) was subjected to sodium bisulfite treatment using the EZ DNA Methylation™-GOLD Kit (Zymo Research) according to the manufacturer’s protocols. Multiplex PCR was performed with optimized primer combination sets. A 20 μL PCR mixture was prepared for each reaction and included 1x reaction buffer (Takara), 3 mM Mg2+, 0.2 mM dNTPs, each primer (0.1 μM), 1U HotStarTaq polymerase (Takara) and 2 μL template DNA. The cycling program ran with the following conditions: 95°C for 2 min; 11 cycles of 94°C for 20s, 63°C for 40s with a 0.5°C decrease per cycle, and 72°C for 1 min; 24 cycles of 94°C for 20s, 65°C for 30s, and 72°C for 1 min and 72°C for 2 min.
| Target | Length | Distance to TSS |
Primer F | Primer R |
|---|---|---|---|---|
| TBX15_46 | 197 | -523 | AATTTTGGGGAGTTGGAAGTAGG | TCTACCTTATCTCCAACTTCTCTACTAAAACC |
| TBX15_47 | 211 | -406 | GATGTTAGTTTTTTGATATTTTGGTTGGA | AACCTCTAAAACRCCRAATTCTCTACC |
| TBX15_48 | 210 | -882 | GGATGAGAAAAGAGTTGGGAAAG | CATTCACTACTACCRAACTTCAACAA |
| TBX15_49 | 209 | 93 | GATTTYGYGGATAGYGGTTTAGATATAGTT | CAAACTATAAACCCCCTAACCTAAACT |
Table 1: Primers used in this study.
Index PCR: PCR amplicons were diluted and amplified using indexed primers. Specifically, a 20 μL mixture was prepared for each reaction and included 1x reaction buffer (NEB Q5TM), 0.3 mM dNTPs, 0.3 μM F primer, 0.3 μM index primers, 1 U Q5TM DNA polymerase (NEB) and 1 μL diluted template. The cycling program ran with the following conditions: 98°C for 30s; 11 cycles of 98°C for 10s, 65°C for 30s, 72°C for 30s and 72°C for 5 min. PCR amplicons (170 bp-270 bp) are separated by Agarose electrophoresis and purified using the QIAquick Gel Extraction kit (QIAGEN).
Sequencing: Libraries from different samples were quantified and pooled together, followed by sequencing on the Illumina MiSeq platform according to the manufacturer’s protocols. Sequencing was performed with the 2 × 300 bp paired-end mode.
Statistical analysis: Using R language (version 4.0.2) for statistical analysis, Student’s t-test was used to analyze the methylation level differences between tumor tissues and paired paracancerous tissues. The associations between cancer tissue methylation status and clinicopathological characteristics were analyzed by Wilcoxon exact rank tests. A Receiver Operating Characteristic (ROC) curve was generated to determine the TBX15 gene methylation threshold, and the area under the ROC curve (AUC) was determined to evaluate the diagnostic value in HCC. The determination of the ability of TBX15 methylation in tumor tissues to predict relapse-free survival was performed via Kaplan-Meier survival analysis and log-rank tests. Cox proportional hazards regression analysis was performed to evaluate the prognostic utility of TBX15 methylation in HCC. Results were determined significant with a standard of P<0.05.
Results
Differences in TBX15 methylation levels between tumor tissues and paracancerous tissues
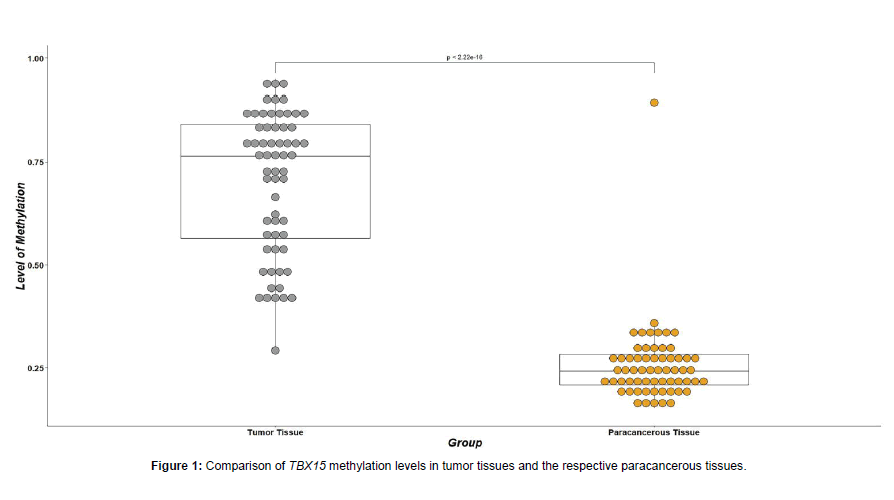
MethylTarget™ technology was used to analyze the methylation levels of tumor tissues and paired paracancerous tissues. Data are expressed in a “mean ± Standard Deviation (SD)” format. The methylation level was calculated as the number of methylated reads at the site divided by the total number of reads at the site. The methylation levels of four targeted regions within the TBX15 promoter region were measured. The methylation levels of each region were higher in tumor tissues than in the respective paracancerous tissues (P<0.01), (Table 2). Additionally, 80% of tumor tissues displayed Hypermethylation, while the adjacent paracancerous tissues showed hypomethylation. In addition, the methylation levels of TBX15 in all 61 tumor tissues were higher than those in the paired adjacent paracancerous tissues. The difference was statistically significant (P<0.001, (Figure 1).
| Target | Tumor tissues (mean ± SD) | Paracancerous tissues (mean ± SD) | P-value` |
|---|---|---|---|
| TBX15_46 | 74.05 ± 15.89 | 30.41 ± 10.92 | P<0.01 |
| TBX15_47 | 68.94 ± 19.33 | 20.55 ± 9.99 | P<0.01 |
| TBX15_48 | 70.87 ± 17.91 | 29.73 ± 11.83 | P<0.01 |
| TBX15_49 | 65.32 ± 18.91 | 16.60 ± 9.40 | P<0.01 |
| TBX15 | 70.12 ± 16.86 | 25.52 ± 9.68 | P<0.01 |
Table 2: Methylation analysis of four target region in tumor tissues.
Relationships between the TBX15 methylation level and clinicopathological characteristics of patients with HCC
This study explored the association between the TBX15 methylation level and clinicopathological characteristics of patients with HCC. Several factors including age, sex, AFP level, HBV status, TNM stage and tumor size, presence of thrombi and/or vascular invasion and envelope integrity were included in this study. As indicated in (Table 3), male patient TBX15 methylation levels were significantly higher than those of female patients (P<0.05). TBX15 methylation levels were increased in tumor tissues with vascular invasion (P>0.05) and tumor tissues with poor envelope integrity (P<0.05), which may indicate a poor prognosis outcome. In addition, Hypermethylation of TBX15 was related to stage III and IV disease (P<0.05). There was no significant correlation between methylation and other clinicopathological characteristics (P>0.05).
| Feature | n | Methylation level | P-value |
|---|---|---|---|
| Age | |||
| >50 | 16 | 68.28 ± 19.21 | 0.64 |
| ≤ 50 | 45 | 70.78 ± 16.32 | |
| Sex | |||
| Male | 48 | 79.43 ± 17.56 | 0.0046 |
| Female | 13 | 67.60 ± 16.11 | |
| AFP | |||
| >20 | 32 | 72.49 ± 16.96 | 0.27 |
| ≤ 20 | 29 | 67.51 ± 16.95 | |
| HBV status | |||
| HBV (+) | 49 | 70.72 ± 17.28 | 0.64 |
| HBV (-) | 12 | 68.60 ± 17.05 | |
| Vascular invasion | |||
| Yes | 29 | 70.95 ± 18.60 | 0.63 |
| No | 32 | 69.37 ± 15.67 | |
| Envelope integrity | |||
| Intact | 11 | 63.20 ± 16.03 | 0.02 |
| Lost | 50 | 73.67 ± 16.68 | |
| Tumor size | |||
| >5 cm | 35 | 66.85 ± 17.27 | 0.12 |
| ≤ 5 cm | 26 | 72.90 ± 16.52 | |
| Thrombus | |||
| Yes | 12 | 71.33 ± 18.67 | 0.8 |
| No | 49 | 69.83 ± 16.76 | |
| TNM stage | |||
| I+II | 15 | 60.83 ± 17.76 | 0.02 |
| III+IV | 46 | 73.15 ± 15.77 | |
Table 3: Relationship between the TBX15 methylation level and clinicopathological characteristics of patients with HCC.
Diagnostic value of TBX15 methylation in HCC
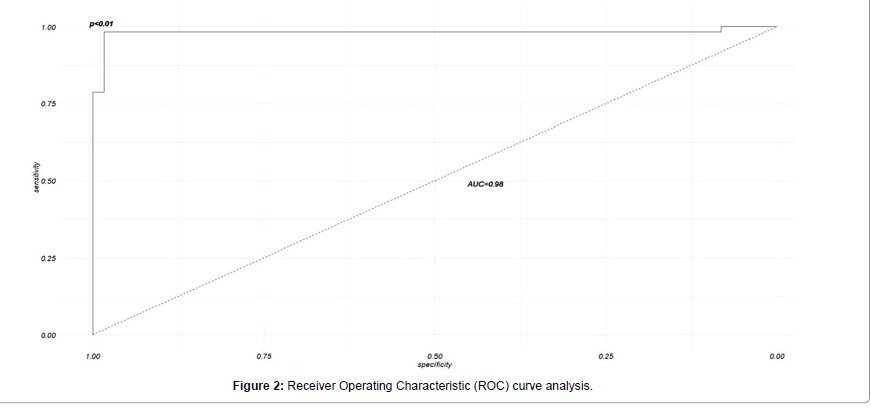
ROC curve analysis was performed to determine the diagnostic potential of TBX15 methylation. When the threshold was 38% for the methylation level, the Youden index was the largest, with a sensitivity of 96%, specificity of 84% and AUC of 0.98 (CI=0.952-0.998, P<0.01), (Figure 2). The diagnostic concordance rate was 90%.
Association between TBX15 methylation and the prognosis of HCC patients
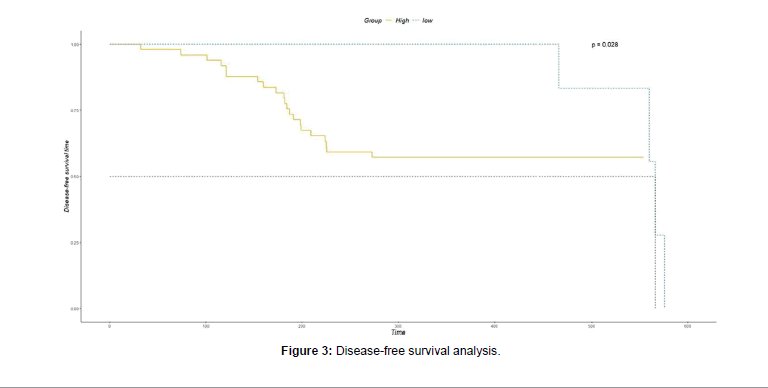
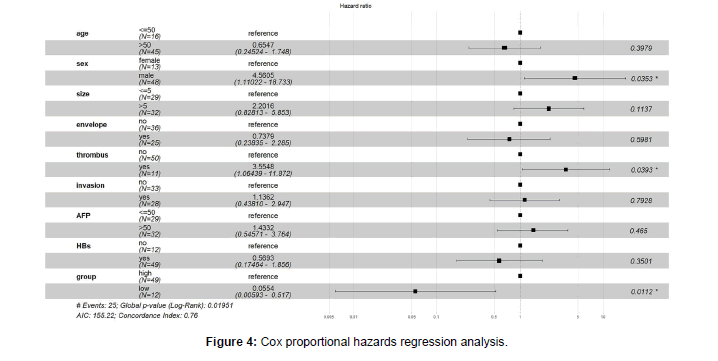
Kaplan-Meier survival analysis according to the relapse-free survival rate was performed to evaluate the prognostic potential of TBX15 methylation. Based on the methylation level threshold obtained by the ROC curve analysis, HCC patients were divided into a Hypermethylation group (n=49) and a hypomethylation group (n=12). During the follow-up, 21 people experienced relapse in the Hypermethylation group, with two dying from cancer. Four people had a relapse in the hypomethylation group, with two dying from cancer. During the follow-up period, 3 patients in the Hypermethylation group lost contact with the research team. The relapse-free survival time of the Hypermethylation group was significantly lower than that of the hypomethylation group (P<0.01), (Figure 3). Cox proportional hazards regression model analysis was used to further study the prognostic value of TBX15 methylation in HCC patients. The results showed that the risk of relapse in the hypomethylation group was 5.6% lower than that in the Hypermethylation group, (P<0.01) (Figure 4 and Table 4).
| Variables | Multivariate model | |||
|---|---|---|---|---|
| Number of patients | HR | 95% CI of HR | P value | |
| Age | ||||
| ≤ 50 | 16 | 1 | 0.25-1.75 | 0.4 |
| >50 | 45 | 0.65 | ||
| Sex | ||||
| Female | 13 | 1 | 1.11-18.73 | 0.035 |
| Male | 48 | 4.56 | ||
| Size | ||||
| ≤ 50 | 29 | 1 | 0.83-5.96 | 0.11 |
| >50 | 32 | 2.2 | ||
| Envelope integrity | ||||
| Lost | 36 | 1 | 0.24-2.29 | |
| Intact | 25 | 0.74 | 0.6 | |
| Thrombus | ||||
| No | 50 | 1 | 1.06-11.9 | 0.04 |
| Yes | 11 | 3.55 | ||
| Invasion | ||||
| No | 33 | 1 | 0.44-2.95 | 0.8 |
| Yes | 28 | 1.14 | ||
| AFP | ||||
| ≤ 50 | 29 | 1 | 0.55-3.77 | 0.47 |
| >50 | 32 | 1.43 | ||
| HBs | ||||
| No | 12 | 1 | 0.17-1.86 | 0.35 |
| Yes | 49 | 0.57 | ||
| Methylation | ||||
| High | 49 | 1 | 0.0059-0.52 | 0.01 |
| Low | 12 | 0.05 | ||
Table 4: Multivariate cox regression analysis data.
Discussion
Research shows that the annual incidence of HCC increases by approximately 3.1% per year [19]. Seventy- five percent of patients in this study diagnosed with HCC had stage III or IV disease. The overall prognosis of HCC patients after treatment is poor. Clinically, the most important thing is to identify HCC patients and provide active treatment measures. In addition, patients must also be monitored after treatment for relapse to reduce mortality. Therefore, it is imperative to find biomarkers that can be used to diagnose HCC and predict the prognosis of HCC patients. DNA methylation is the most common and widely studied epigenetic modification and is an important mechanism related to the occurrence and development of cancers; thus, it is a promising factor to consider when attempting identifying biomarkers.
TBX15 is an important transcription factor that can regulate the proliferation and differentiation of tissue-specific stem cells and progenitor cells [11,12]. The expression of TBX15 can inhibit tumor cell growth and invasion. TBX15 can also change the ratio of pro-apoptotic proteins to antiapoptotic proteins. TBX15 can negatively affect tumor regulation [20]. DNA Hypermethylation can cause TBX15 expression to be down regulated. TBX15 is hypermethylated and down regulated in ovarian cancer [21], pancreatic cancer [22] and esophageal cancer [23]. In this study, we confirmed that the methylation levels of TBX15 in HCC tissues were higher than those in paired paracancerous tissues. The methylation of TBX15 may lead to the down regulation of its expression and the development of tumors. In addition, the association between TBX15 Hypermethylation and the clinicopathological characteristics of HCC patients was also explored. The results showed that TBX15 methylation levels were increased in incomplete capsules (P<0.05). The TBX15 methylation level was significantly increased in patients with advanced-stage cancer, suggesting that TBX15 methylation is associated with malignant behavior.
To determine the TBX15 methylation cutoff value, ROC curve analysis was used, and the AUC value of TBX15 methylation was calculated to be 0.98 (P<0.01). The ROC curve analysis indicated that TBX15 methylation can be used as a diagnostic biomarker for HCC. The Cox proportional hazards regression model showed that TBX15 methylation was an independent factor affecting the prognosis of HCC. To verify the impact of TBX15 methylation on the prognosis of HCC patients, the disease-free survival rate of HCC patients with different TBX15 methylation statuses was studied. Patients with hypermethylated TBX15 had significantly worse prognosis than patients with hypomethylated TBX15. Patients with hypomethylated TBX15 were more likely to experience relapse and had a shorter relapse-free survival time than those with hypermethylated TBX15. Therefore, patients with hypermethylated TBX15 can receive earlier intervention to achieve better curative effects. Due to the limited follow-up time, the relationship between TBX15 methylation and overall survival was not evaluated. In the future, more patients who complete follow-up can be studied for a longer duration than was used in this study to obtain more meaningful results regarding the correlation between TBX15 and the overall survival rate.
Conclusion
The results of research on DNA methylation have been applied clinically. Some reports have shown that gene methylation can be detected in body fluids such as urine, plasma, and serum and tissues. In addition, gene silencing caused by TSG methylation is reversible. Demethylating reagents such as 5-azacytidine can re-express previously silenced genes in cancer cells. Clinically, Demethylating reagents such as decitabine or azacytidine have been used to treat solid carcinomas. Therefore, using gene methylation as a biomarker for tumor diagnosis and treatment is a very promising research strategy. However, more clinical trials are needed to further determine the clinical value of TBX15 methylation.
Acknowledgement
AJE provided English editing assistance.
Funding
This research was supported by the San-Ming Program from the Shenzhen Healthcare Research Project (SZSM201412006).
Availability of Data and Materials
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Author Contributions
Rui-kun Zhang and Jia-lin Liu conceived of the study design and determined appropriate surveys and survey items for the study. Zhu Hai collected the data. Rui-kun Zhang and Zhu Hai performed the data analysis. Hai-yang Xie and Jia-lin Liu interpreted analysis results and participated in drafts and revisions of this manuscript. All authors reviewed and provided approval for the final version of the manuscript.
Ethics Approval and Consent to Participate
This research has been approved by the Ethics Committee of the First Associated Hospital of Zhejiang University. All subjects signed an informed consent and all methods are done according to the Declaration of Helsinki together with relevant rules and standards.
Patient Consent for Publication
All patients who participated also consented for the publication of this manuscript.
Competing Interests
Not applicable..
References
- Bosch FX, Ribes J, Borras J (1999) Epidemiology of primary liver cancer. Semin Liver Dis 19: 271-285.
- Dufour JF, Johnson P (2010) Liver cancer: From molecular pathogenesis to new therapies: Summary of the EASL single topic conference. J Hepatol 52: 296-304.
- EL Serag HB (2011) Hepatocellular carcinoma. New England Journal of Medicine 365: 1118-1127.
- Njei B, Rotman Y, Ditah I, Lim K (2015) Emerging trends in hepatocellular carcinoma incidence and mortality. Hepatology 61: 191-199.
- Bruix J, Sherman M (2011) Management of hepatocellular carcinoma: An Update. Hepatology 53: 1020-1022.
- Sherman M (2010) Serological surveillance for hepatocellular carcinoma: Time to quit. J Hepatol 52: 614-615.
- Zhu J (2006) DNA methylation and hepatocellular carcinoma. J Hepatobiliary Pancreat Surg 13: 265-273.
- Baylin SB, Herman JG (2000) DNA hypermethylation in tumorigenesis: Epigenetics joins genetics. Trends Genet 16: 168-174.
- Jones PA, Laird PW (1999) Cancer epigenetics comes of age. Nat Genet 21: 163-167.
- Lee KY, Singh MK, Ussar S, Wetzel P, Hirshman MF, et al. (2015) TBX15 controls skeletal muscle fibre-type determination and muscle metabolism. Nat Commun 6: 8054.
- Showell C, Binder O, Conlon FL (2004) T-box genes in early embryogenesis. Developmental dynamics: An Official publication of the American Association of Anatomists 229: 201-218.
- Takashima Y, Suzuki A (2013) Regulation of organogenesis and stem cell properties by T-box transcription factors. Cell Mol Life Sci 70: 3929-3945
- Sheeba CJ, Logan MP (2017) The roles of t-box genes in vertebrate limb development. Curr Top Dev Biol 122: 355-381.
- Arribas J, Gimenez E, Marcos R, Velazquez A (2015) Novel antiapoptotic effect of TBX15: Overexpression of TBX15 reduces apoptosis in cancer cells. Apoptosis 20: 1338-1346.
- Gozzi G, Chelbi ST, Manni P, Alberti L, Fonda S, et al (2016) Promoter methylation and downregulated expression of the TBX15 gene in ovarian carcinoma. Oncol Lett 12: 2811-2819.
- Kron K, Liu L, Trudel D, Pethe V, Trachtenberg J, et al. (2012) Correlation of ERG expression and DNA methylation biomarkers with adverse clinicopathologic features of prostate cancer. Clin Cancer Res 18: 2896-2904.
- Gao F, Liang H, Lu H, Wang J, Xia M, et al. (2015) Global analysis of DNA methylation in hepatocellular carcinoma by a liquid hybridization capture-based bisulfite sequencing approach. Clin Epigenetics 7: 86.
- Zheng Y, Huang Q, Ding Z, Liu T, Xue C, et al. (2018) Genome-wide DNA methylation analysis identifies candidate epigenetic markers and drivers of hepatocellular carcinoma. Brief Bioinform 19: 101-108.
- Ryerson AB, Eheman CR, Altekruse SF, Altekruse SF, Ward JW, et al. (2016) Annual Report to the Nation on the Status of Cancer, 1975-2012, featuring the increasing incidence of liver cancer. Cancer 122: 1312-1337.
- He ML, Chen Y, Peng Y, Jin D, Du D, et al. (2002) Induction of apoptosis and inhibition of cell growth by developmental regulator hTBX5. Biochem Biophys Res Commun 297: 185-192.
- Wu TI, Huang RL, Su PH, Mao SP, Wu CH, et al. (2019) Ovarian cancer detection by DNA methylation in cervical scrapings. Clin Epigenetics 11: 166.
- Majumder S, Taylor WR, Yab TC, Berger CK, Dukek BA, et al. (2019) Novel methylated DNA markers discriminate advanced neoplasia in pancreatic cysts: Marker discovery, tissue validation and cyst fluid testing. Am J Gastroenterol 114: 1539-1549.
- Qin Y, Wu CW, Taylor WR, Sawas T, Burger KN, et al. (2019) Discovery, validation, and application of novel methylated DNA markers for detection of esophageal cancer in plasma. Clin Cancer Res 25: 7396-7404.
Citation: Zhang RK, Zhu H, Xie HY, Liu JL (2021) The Clinical Value of Abnormal TBX15 Hypermethylation in HCC. J Oncol Res Treat S4: 002. DOI: 10.4172/aot.s4.1000002
Copyright: © 2021 Zhang RK, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Select your language of interest to view the total content in your interested language
Share This Article
Open Access Journals
Article Tools
Article Usage
- Total views: 2129
- [From(publication date): 0-2021 - Dec 09, 2025]
- Breakdown by view type
- HTML page views: 1393
- PDF downloads: 736