Research Article Open Access
Expression Profile of Angiogenesis Related Genes in Head and Neck Squamous Cell Carcinoma
| S. Jung1*, S. Sielker1,2, N. Purcz3, C. Sproll4, A. Raem2 and J. Kleinheinz1 | ||
| 1Research Unit Vascular Biology of Oral Structures (VABOS), Department of Cranio-Maxillofacial Surgery, University Hospital Muenster, Waldeyerstraße 30, D-48149 Muenster, Germany | ||
| 2Arrows biomedical Deutschland GmbH, Muenster, Germany | ||
| 3Department of Cranio-Maxillofacial Surgery, University Hospital Kiel, Germany | ||
| 4Department of Cranio-Maxillofacial Surgery, University Hospital Duesseldorf, Germany | ||
| Corresponding Author : | Susanne Jung MD, DMD Research Unit Vascular Biology of Oral Structures (VABOS) Department of Cranio-Maxillofacial Surgery University Hospital Muenster, Waldeyerstraße 30 D-48149 Muenster, Germany Tel: +49-251-834-7004 Fax: +49-251-834-7184 E-mail: Susanne.Jung@ukmuenster.de |
|
| Received January 04, 2013; Accepted January 23, 2013; Published February 01, 2013 | ||
| Citation: Jung S, Sielker S, Purcz N, Sproll C, Raem A, et al. (2013) Expression Profile of Angiogenesis Related Genes in Head and Neck Squamous Cell Carcinoma. J Biochip Tissue Chip 3:104. doi:10.4172/2153-0777.1000104 | ||
| Copyright: © 2013 Jung S, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. | ||
Related article at Pubmed Pubmed  Scholar Google Scholar Google |
||
Visit for more related articles at Journal of Bioengineering and Bioelectronics
Abstract
Objectives: The molecular patterns of tumorigenesis of head and neck squamous cell carcinoma are mostly unknown to date. The concept of this investigation was to expose a genetic fingerprint of angiogenesis in HNSCC with special regard to VEGF as the outstanding mediator of neo-angiogenesis.
Material and Methods: A panel of significant transcriptional alterations of VEGF related genes in 83 cancer samples compared to healthy mucosa with microarray technique was established. RT-PCR and immunohistochemistry were performed to confirm the microarray results.
Results: In four selected marker genes (VEGFA, RAC2, IL8, PLA2G10) we detected a highly significant
transcriptional alteration; three of the genes (VEGFA, RAC2, IL8) showed up-regulation in 88% to 100% of the samples, PLA2G10 down-regulation in 99% of the analyzed tissue samples. Additionally a significant transcription alteration in angio-miRNAs was detected: miR-21 and miR-31 were over-expressed in upto 87% and upto 89% respectively in the analyzed samples. MiR-20a, miR-126, miR-378, miR-17, miR-19a, miR-27b and miR let-7f were down-regulated in 83% to 95% of the specimens. A specific pattern subjected to the different tumor types could not be verified.
Conclusion: An expression profile of VEGF related genes was established and selective angio-miRNAs were identified as potential biomarkers on the basis of 83 HNSCC samples. Identification of new relevant oncogenes and insight into the process of tumor vasculogenesis with a highly sensitive method opens new vistas in cancer biology.
| Keywords | |
| Oral cancer; Head and neck squamous cell carcinoma; VEGF; Angio-miRNA; Expression profiling; Microarray analysis | |
| Introduction | |
| Head and Neck Squamous Cell Carcinoma (HNSCC) represents an important pathology of the upper digestive tract, being the sixth most common cancer diagnosed around the world with an incidence of over 500,000 newly diagnosed cases worldwide each year [1]. | |
| Therapeutical procedures have been developed and refined during the last years but still locoregional relapse occurs in 60% of the patients and metastatic disease develops in 15–25%. The prognosis for advanced or recurrent disease is poor [2]. Additionally, about 5% of the patients come down with second primary tumors each year. Although improvements have been achieved in surgical techniques, radiation therapy protocols and chemotherapeutic regimes, the overall fiveyear survival rate for this disease remains at about 50% and has not significantly improved during the last three decades [3]. | |
| For the development of new effective therapeutical strategies and focused intervention it is of vital importance to further elucidate the biomolecular determinants of the head and neck carcinogenesis process. One crucial mechanism of tumor growth and therefore strongly related to the prognosis is neo-angiogenesis. Blood supply is essential for tumor growth and metastasis, since the blood not only supplies nutrients and oxygen but also provides a vascular route for hematogenous spread to distant sites. | |
| Increased microvessel density is related to patients’ prognosis [4]; in an immunohistochemical analysis of over 50 specimens of oral squamous cell carcinoma micro vessel density was clearly higher in malignant tissue compared to normal mucosa and pre-malignant lesions. In addition, a clear correlation of VEGF-A expression and poor prognosis was shown already. In a multivariate logic regression analysis of Seki et al. [6] patients with high expression of VEGF-A or -C had a significant correlation to poor prognosis even in early stages [5,6]. | |
| These results underline the clinical significance of angiogenesis in the progression of malignant disease. One of the most potent mediator of neoangiogensis is the mitogen VEGF. VEGF-A plays an essential role in vasculogenesis and angiogenesis. The protein is a glycosylated mitogen that specifically acts on endothelial cells and has various effects: mediating increased vascular permeability, inducing angiogenesis and endothelial cell growth, promoting cell migration and inhibiting apoptosis [7]. VEGF-A is also known as a Vascular Permeability Factor (VPF), other members of this growth factor family are Placenta Growth Factor (PlGF), VEGF-B, VEGF-C, VEGF-D and VEGF-E. | |
| VEGF acts as a potent survival factor for endothelial cells during physiological and tumor–induced angiogenesis and it has been shown to induce the expression of anti-apoptotic proteins in endothelial cells [8,9]. The instantaneous angiogenic effect of VEGF is the increase of vessel permeability and mitogenic stimulation of endothelial cells. According to its potential VEGF is also involved in pathophysiological processes like tumor growth; it is regarded as key regulator in tumor– induced neoangiogenesis. Mainly in hypoxic tumor regions raised VEGF levels could be scored [10]. The transition of tumor cells to an angiogenic phenotype, referred to as the “angiogenic switch”, contributes to the disruption of balance between angiogenic promoters and inhibitors. Experimental and clinical reports confirm that VEGF plays a pivotal role in regulating angiogenesis and vasculogenesis in solid tumors and is tightly associated with the angiogenic switch. This transition to a pro-angiogenic phenotype has been demonstrated to be crucial in the progression of oropharyngeal epithelial dysplasia to invasive head and neck squamous cell carcinoma [11]. | |
| In many tumors, angiogenesis is induced not by hypoxia, but by genetic alterations. The loss of function of tumor suppressor genes such as p53 or the activation of oncogenes including RAS result in increased expression and/or secretion of VEGF [12]. | |
| IL 8 (CXCL 8) is one of the major mediators of the inflammatory response and thus physiologically a strong promoter of vascular modulation as increased vessel permeability or angiogenesis. IL8 is able to initiate the intracellular VEGF pathway via NFκB activation. It controls the expression of VEGF in endothelial cells and thereby promotes the activation of VEGF receptors in an autocrine fashion. In cancer research, IL8 has already been identified as proangiogenic cytokine, which promotes cancer progression. VEGF and IL8, both bind to membranous endothelial cell receptors and contribute to angiogenesis [13,14]. | |
| RAS-related C3 botulinum toxin substrate 2 (RAC2) exerts the control of cell growth, cytoskeletal reorganization the activation of protein kinases and endothelial cell migration. Studies on RAC2 knockout mice revealed its vital importance for postnatal angiogenesis and suggested that RAC2 is involved in neovascularization by mobilizing bone marrow derived endothelial cells [15]. | |
| Phospholipase A2 group 10 (PLA2G10) induces the expression and release of VEGF-A and VEGF-C from macrophages. Via VEGF pathway, they play an important role in inflammatory and/or neoplastic angiogenesis and lymphangiogenesis [16]. | |
| It is a major regulator of the intracellular arachidonic acid metabolism: PLA2 hydrolyses arachidonic acid, which is the limiting substrate for prostaglandin production from membrane phospholipids. Angiogenesis is induced via prostaglandin E2 pathway, e.g. in colorectal cancer [17]. | |
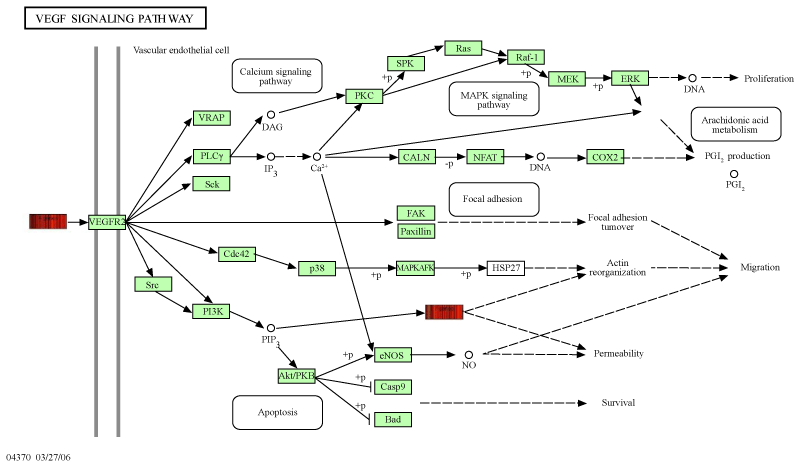
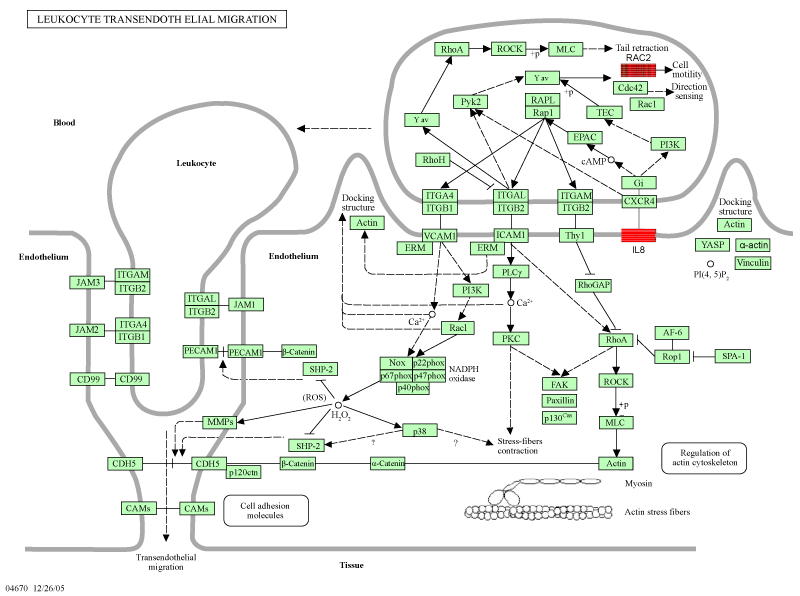
| The interconnection of these selected genes, exact location and mode of regulation is displayed in figure 2 and 3. They may come up as key regulators in tumor growth and precursors of hematogenous metastasis and where thus in the focus of our investigation. | |
| MicroRNAs (miRNAs, mi-Rs) play obviously a significant role in cell proliferation and differentiation and act as oncogenes or tumor supressors [18]. | |
| Not only is miRNA profiling a useful tool for clinical diagnostics, miRNA analysis can help to predict the cancer’s response to therapy and to consider and evaluate the benefit of alternative therapy regimens [19]. | |
| Specific miRNAs (angiomiRs) have recently been shown to regulate angiogenesis in vivo. miRNA-126, an EC-restricted miRNA, regulates vascular integrity and developmental angiogenesis [20]. AngiomiRNA therapeutics might provide a natural means of normalizing the expression of disease genes by interfering with their expression. | |
| Materials and Methods | |
| Patient data | |
| 83 tissue samples were taken during tumor surgery after informed consent of the patients. Included were patients with a histologically diagnosed (recurrent) squamous cell carcinoma of the oral cavity. These patients were over 18 years old and had not received any adjuvant radiation or chemotherapy. Healthy specimens were taken from patients without any malignant history in terms of traumatology or orthognathic surgery. The Ethics Committee of the medical faculty approved of the study setup. | |
| The tissue samples were snap-frozen in liquid nitrogen after surgery and stored at –80°C until further usage. | |
| RNA extraction and microarray assay | |
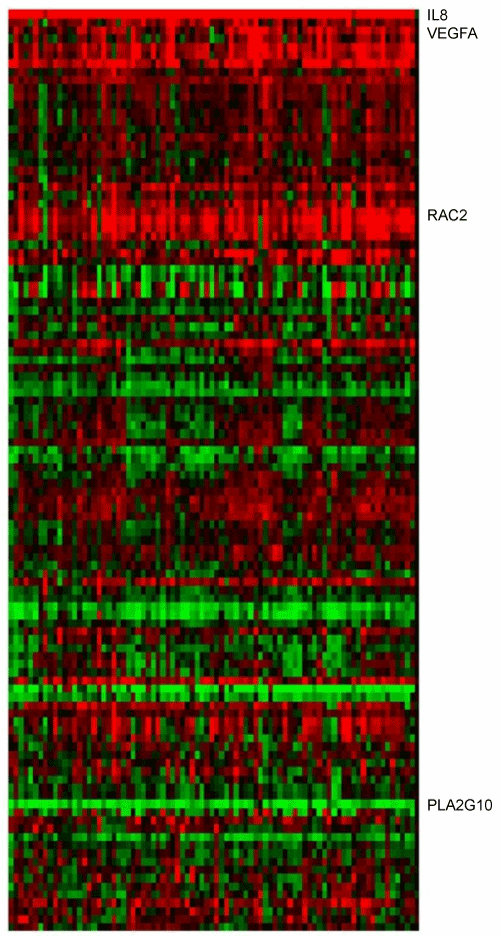
| Total RNA (including microRNA) was prepared by Qiazol (Qiagen) extraction and purification with the miRNeasy Mini Kit (Qiagen). Purity and integrity of the isolated total RNA were assessed on the Agilent 2100 bioanalyzer (Agilent Technologies). For microarray analyses, we used the Agilent Array platform employing the manufacturer’s standard protocols for sample preparation and microarray hybridization. Gene expression analysis was performed with the Whole Human Gene Expression Microarray (4x44K; GPL4133; Agilent Technologies) and microRNA expression analysis was performed with the human miRNA microarrays (V2; GPL8936; based on Sanger miRBase release 10.1; Agilent Technologies). Following the washing steps the arrays were scanned using the Agilent G2505B Microarray Scanner (Agilent Technologies) and feature extraction was performed with Feature Extraction software version 9.5 (Agilent Technologies) according to the Agilent quality control criteria. Data files from mRNA and miRNA microarrays were analyzed by GeneSpring GX 7.3.1 according to manufactures protocol (Agilent Technologies). Average values of the replicate spots of each mRNA and miRNA were due to background subtracted, normalized and analyzed. Samples of healthy oral mucosa were used as control references. The altered expression factors were correlated to the healthy mucosa pool that was defined as expression factor 1. In the hierarchical clustering, expression factor 1 is illustrated in black, expression factors > 1 are marked red and expression factors <1 are green. | |
| Statistical analysis | |
| Analysis of Variance (ANOVA) was carried out using a p-value of 0.05. The multiple testing correcting of Bonferroni was taken as the level of significance. Hierarchical clustering was performed by an average linkage algorithm and Pearson correlation similarity measure. For the analysis according to tumor status, grading and lymph node status multiple testing corrections of Benjamini and Hochberg were taken as level of significance [21]. | |
| Quantitative real-time PCR | |
| Microarray data validation was performed for the selected candidate genes IL8 (NM_000584; IL8-F 5'-cttgtcattgccagctgtgt-3', IL8-R 5'-gccttgtatttaaaaatgcagtca-3'), VEGFA (NM_001025366; VEGFA-F 5'-ggtccctcttggaattggat-3', VEGF-R 5'-tgtatgtgggtgggtgtgtc-3'), RAC2 (NM_002872; RAC2-F 5'-tacgcctctggggatatctg-3', RAC2-R 5'-gcaggttgagttgggttttc-3') and PLA2G10 (NM_003561; PLA2G10-F 5'-gccaagaactgttgtgcaag-3', PLA2G10-R 5'-gagtccggctcacataggaa-3'). ACTG1 (NM_001614; ACTG1-F 5'-gaacaccgtgggctgttact-3', ACTGF1-R 5'-ctggggcctaatgttctcac-3'), GGNBP2 (NM_024835; GGNBP2-F 5'-tgccggttaaatgatcacaa-3', GGNBP2-R 5'-cagccacgcaactaaagaca-3') and ARF1 (NM_001658, ARF1-F 5'-atttggtgtcgtggaacctc-3', ARF1–R 5'-acagtgcttgtttgtcgaaat-3') were used as housekeeping genes. As cDNA templates one HNSCC tumor sample for each T status and a pool of healthy oral mucosa tissue was analyzed. Total RNA (1 μg) was isolated as described before and used as template for the synthesis of cDNA using the 1st strand cDNA Synthesis kit for RT-PCR (AMV), according to the manufacturer’s protocol (Roche Applied Science). cDNA was amplified on LightCycler 480 (Roche Applied Science) using commercially available primer in triplicates and the DyNAmo Color Flash SYBR Green qPCR Kit (Biozym). PCR efficiencies were determined using the LC480 control kit (Roche Applied Science). qPCR conditions were: initial denaturation at 95°C for 10 min was followed by 35 cycles of denaturation at 95°C for 10 s, annealing at 60°C for 20 s (GGNBP2, ARF1, IL8), annealing at 60°C for 15 sec (PLA2G10), annealing at 64°C for 20 sec (RAC2, VEGFA, ACTG1), and extension at 72°C for 20 s. Quantitative analysis was performed using the Light Cycler 480 Software (Roche Applied Science). | |
| Immunohistochemistry | |
| Serial formalin-fixed paraffin-embedded sections (4 μm) were incubated at 42°C over night and 1 h at 60°C. Afterwards deparaffinized in xylene and rehydrated through decreasing grades ethanol solution. Immunohistochemical procedure was carried out with Dako REALT Detection kit according to the manufactures protocols (Dako, Germany). VEGF Ab-3 primary antibodies with a 1:50 dilution were used (NeoMarkers, Germany). | |
| Results | |
| Patient data | |
| The clinical and pathologic features of the 83 patients in the final expression profiling data set were the following: the median age was 63 years (range: 31-93 years). There were 15 patients under 50 years of age. There were 32 women and 51 men. The T status was T1 in 23 patients (28%), T2 in 32 patients (39%), T3 in 9 patients (11%) and T4 in 18 patients (22%). The tumors were well (4%), moderately (81%) and poorly (15%) differentiated. 37.5% of patients were diagnosed with lymph node metastasis. 15 patients died within 20 months after surgery, 44 patients were smokers and 75% of them drank as well. 20 patients did not smoke and 25 did not drink, 12 patients neither drank nor smoked, 19 patients made no specifications. Healthy tissue controls were taken from mucosa samples during orthognathic of traumatologic surgery after patients´ informed consent. | |
| Microarray analysis | |
| The selected candidate genes showed high significance in their altered expression. Expression factors for the candidate genes are listed in table 1. VEGF-A (p-value 8.06E-08) was over-expressed in ascending order of 74% in T1 status, 75% in T2, 78% in T3 and 83% in T4 status HNSCC tumor specimens. The expression factors ranged from 2.5 to 4.5. RAC2 (p-value 4.34E-09) was over-expressed significantly in 95% of T1 status specimens and in 100% of the T2, T3 and T4 tumor samples. The expression factors ranged here from 3.7 to 6.0. IL8 (p-value 6.44E-11) was over-expressed significantly in 100% of the 83 analyzed specimens with an expression factors from 74 to 100. PLA2G10 (p-value 4.07E-6) was down regulated in 100% of the T1, T2 and T4 tumor samples. In T3 status, 90% of the specimens were down regulated. The expression factors ranged from 0.06 to 0.22. The hierarchical clustering is summarized in figure 1. Figures 2 and 3 illustrate the altered gene expression in the context of their respective pathway. | |
| Real time PCR | |
| ACTG1, ARF1 and GGNBP2 showed similar gene expression in microarray analysis in HNSCC tumor specimens as well as in healthy oral mucosa tissue and thus were chosen as housekeeping genes. The microarray probe sequences were considered when designing the primers. Table 1 presents the results with corresponding expression factors. Real time PCR results confirmed the results of the microarray analysis. In one case of chosen T3 status sample we found in real time PCR an over-expression of PLA2G10 in correlation to healthy oral mucosa tissue. In microarray analysis this sample showed an absent flag signal. | |
| Immunohistochemistry | |
| In an HE staining the tumor specimens were selected for immunohistochemistry. 12 of the 14 stained tissue samples showed positive IHC signals with a range of stained cells of 20% to 100% with comparable intensity. Figure 4 summarizes the staining results of three cases from grading G1 to G3. Immunohistochemical results underlined the microarray results. The intensity of the immunohistochemical staining did not correspond linearly with the over expression of our marker gene. | |
| AngiomiRNA | |
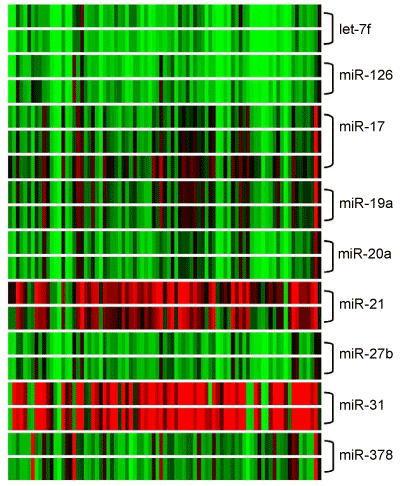
| We analyzed a set of angiomiRs by microarray analysis and found out that few of them were significantly altered in expression. The expression factors for selected angiomiRs are listed in table 2. The expression factors are related to a healthy skin mucosa pool with an expression factor of 1. A correlation to T status, lymph node status or grading could not be observed. | |
| From miR17-92 cluster miR17 (p-value 1.92E-2), miR-19a (p-value 1.7E-08) and miR-20a (p-value 7.44E-08) showed a consistent expression. MiR-17 and miR-19 were down regulated in about 79% of all 83 HNSCC tumor specimens and miR-20a in 91%. MiR-378 (p-value 3.61E-86) and miR-126 (p-value 2.55E-14) were down regulated in 83% and 96%, respectively. MiR-27b (p-value 5.91E-09) and let-7f (p-value 7.9E-15) were down regulated in 93% and 99% of all 83 HNSCC tumor specimens. Expression factors ranged from 0.18 to 0.62. | |
| The so-called pro-angiomiRs miR-21 (p-value 5.7E-66) and miR-31 (p-value 1.15E-78) showed an over-expression in microarray analysis upto 87% and upto 89% respectively. Expression factors ranged from 3.27 to 13.5. A statistical correlation to T status, lymph node status or grading could not be observed for miR-21. The outstanding exception is miR-31, which shows a significantly higher expression according to higher grading (G2 and G3), miR-31 expression factors range from 0.75 (p value 0.003) in G1 tumors, over 13.6 in G2 tumors, upto 16.6 in G3 tumors. With increasing de-differentiation miR-31 expression ascends. The hierarchical clustering of angiomiRs is summarized in figure 5. There is no correlation of gene expression pattern or immunohistochemical detection of VEGF to clinical parameters as long time survival or recurrent disease. Poor prognosis was associated with increasing T stadium. | |
| Discussion | |
| The elucidation of the molecular biology of cancer cells has identified various molecular pathways that are altered in different cancers. This information is currently used to identify oncological biomarkers and to develop potential therapies that target molecules in these pathways. | |
| Several discrete steps are discernible in the biological cascade leading to tumor growth, neo-angiogenesis and metastasis: loss of cellular adhesion, increased motility and invasiveness, entry and survival into the circulation, entrance into new tissue, and eventual colonization of a distant site [22]. | |
| In our study we describe a gene expression profile regarding VEGF and its pathways in HNSCC. We investigated tissue samples of 83 patients in selected marker genes and we detected a significantly altered expression pattern compared to healthy mucosa. | |
| We choose VEGF-A, PLA2G10 an RAC2 for their position and their interconnection in the VEGF signaling pathway (Figures 2 and 3). | |
| The analyzed genes represent landmarks in angiogenesis and act on different levels. An altered expression of these genes has been observed in other tumor entities and they represent potential targets for specific anti-angiogenic therapy strategies on a post-transcriptional level. | |
| VEGF expression has been reported to be elevated in both oral dysplasia and head and neck squamous cell carcinomas; Denhart et al. [23] observed in 45 samples that 50% of premalignant and 75% of malignant oral lesions expressed increased levels of either VEGF or its receptors by in situ hybridization. | |
| In breast cancer VEGF expression has prognostic value: Downregulation of VEGF expression induced apoptosis and decreased cell proliferation in different breast cancer cell lines and was correlated with reduced malignancy and prolonged survival [24]. Ghosh et al. [25] found a significant over-expression of VEGF and its receptors in breast tumors with a worse outcome. VEGF has underlined its reliability as biomarker in many tumor entities. | |
| IL8 in our data is the analyzed gene that shows the strongest alteration, e.g. the highest levels of over-expression. IL8 has been identified as an essential regulator of cross talk between endothelial cells and tumor cells creating a chemotactic gradient, which initiates tumor invasion and finally field cancerization. In studies with HNSCC tumor cell lines IL8 and IL1 acted as potent chemo attractant for tumor cells [26]. | |
| Chiang et al. [27] differentiated expression profiles of metastasisrelated genes by suppression subtractive hybridization and microarray analysis and found an over-expression of IL8 in invasive oral cancer together with Matrix Metalloproteinases (MMPs); the increased level of IL8 was correlated with reduced survival. | |
| In oral dysplasia, the synergistic influence of IL8 and VEGF has already been described: Rao et al. [28] deduced in 2010 from their analysis of pro-inflammatory genes in oral squamous cell carcinoma a cross talk between IL8 signal cascading and VEGF-dependent tumor progression. In their data they found a distinct over-expression of IL8 and VEGF in microarray analysis. | |
| The myeloid-specific Rac2 belongs to the Rac family, a group of small GTPases, which bind and hydrolyze GTP, thus switching between GDP–the inactive form–and GTP as active metabolite. As members of the RAS-related oncogene family they are supposed to play a significant role in tumorigenesis. The role of Rac2 is so far widely unknown; it has effects on chemotaxis and cell adhesion [29]. | |
| In an investigation of motility-related proteins as Rho A, Cdc42, Rac2 and others by immunohistochemistry and western blot, an overexpression of Rac2 in malignant transformed tissue in comparison to healthy mucosa could be verified. The authors conclude that Rac2 identifies itself not only as molecular tumor marker in HNSCC but also as potential therapeutical target as to motility, cell migration, invasion and metastasis [30]. | |
| PLA2G10 was the only gene in our investigation where we observed a significant down-regulation; this observation leads to the assumption that PLA2 has a protective or controlling function in neo-angiogenesis. | |
| The PLA2 family has underlined its role as tumor biomarker and potential therapeutical point of attack in many cancer entities. An over-expression of PLA2 was observed in prostate cancer, where its high levels came with poor survival rates [31], an observation that contradicts our results. | |
| In contrast, in gastric adenocarcinoma high levels of PLA2 were associated with reduced metastasis and prolonged survival. Downregulation on the other hand came with more aggressive malignancy [32]. The biology of this tumor entity seems to share more parallels with HNSCC. | |
| Secreted phospholipases A2 have been discussed as promising point of attack for anti-cancer drugs: PLA2 inhibitors are supposed to reduce the release of arachidonic acid and lysophosphatidic acid and thus to inhibit cell growth, differentiation, invasion and angiogenesis. The clue is obviously to filter the specific target in the PLA2 family in the respective tumorigenesis [33]. | |
| The altered expression of PLA2 seemed closely related to the overexpression of IL8. Mournier et al. [34] analyzed the expression pattern of secreted phospholipases in normal colon and colon adenocarcinoma by PCR and immunohistochemistry. They observed an overexpression of PLA2G3 together with a down-regulation of PLA2G2D and PLA2G5. This expression profile came with an over-expression of IL8. The corresponding conclusion from our data might be that the dependent contrary alteration of PLA2G10 and IL8 is a characteristic feature in HNSCC [34]. | |
| MiRNA | |
| MiRNAs regulate the expression of their target genes posttranscriptionally. They seem to be involved in cancer initiation and tumor progression: the first correlation was verified by Calin et al. [35] who found a general deletion or down-regulation of mi-R15 and mi- R16 in chronic lymphatic leukemia in 2002. Relevant oncomiRNAs are the Mi-R17-92 cluster (OncomiR-1) and let-7. | |
| VEGF induces time-dependent expression of miR-191, -155, -31, -17-5p, -18a and miR-20a, with little change in miR-126 and miR-222. Interestingly, this set of VEGF-regulated miRNAs is commonly overexpressed in human tumors and has been implicated in the control of tumor growth, survival and angiogenesis [36]. | |
| An altered expression of let-7 has also been described in cancer research. Down-regulation of let-7 was observed in human lung cancer and reduced levels of let-7 were significantly associated with poor survival [37]. In pancreatic cancer, miRNA have already confirmed their function as diagnostic biomarkers: miR-221, miR-301 and miR- 376a were expressed in tumor cells but not in healthy pancreatic tissue. On the basis of the aberrantly miRNA expression, the authors developed an algorithm to classify the pancreatic pathologies [38]. An increased miR-31 expression in the analyzed tumor tissue hints not only to a malignant transformation but the extent of the upregulation might emerge as a reliable prognostic factor [39]. | |
| With the current knowledge of tumor biology there are different levels of attack. One way is to interfere with tumor growth by inhibiting angiogenesis extracellularly. | |
| Our results strengthen the pharmaceutical advances e.g. with bevacizumab, a monoclonal antibody against VEGF to inhibit tumorinduced angiogenesis. However, it has been observed, that tumor cells side-step its attack by simply up-regulating VEGF [40]. | |
| Finally, the identification of marker genes in key positions of tumorigenesis reveals new targets for a specific therapeutical approach on a genetic level, for example by RNA interference. | |
| RNA interference (RNAi) technology is being used in vitro and in animal models with promising results. SiRNA, bound in an RNAinduced Silencing Complex (RISC) leads to the degradation of the target mRNA, for example VEGF or IL8. The delivery of these specific silencers is in vivo not yet technically mature [41]. | |
| Conclusion | |
| Our research represents a crucial step in the revelation and understanding of molecular carcinogenesis and especially biomolecular neo-angiogenesis in HNSCC. | |
| VEGF shows an altered expression in malignant tissue in genetic and protein-associated aspects. The conclusive results of microarray analysis and immunohistochemistry suggest that in the majority of the analyzed samples there is no further posttranscriptional regulation mechanism. | |
| MiR-31 as prominent proangiomiR unmasked itself as a potent marker molecule for diagnosis and beyond that as an approbate prognosis forecast. | |
| Identification of new relevant oncogenes and insight into the process of tumor vasculogenesis with a highly sensitive method will lead to effective diagnostics and finally individually adapted therapeutical strategies with respect for the special biological characteristics of the respective tumor. | |
| To use the knowledge of the point of attack and to combine it with developing molecular strategies will be the next step in the development of innovative bio-molecular approaches, which will replenish and support well-established therapeutic regimens. | |
| Acknowledgements | |
| We thank arrows biomedical Deutschland GmbH for the realization of the laboratory work of our project. This work was supported by a grant from the Pro Inno II Project, endorsed by the working group of industrial research federations "Otto von Guericke" (AiF), Germany and BMWi, the German Federal Ministry of Economics and Technology. The research project is registered with the WHO clinical trial registry and the clinical trial register of the university Muenster. | |
| References | |
|
|
Tables and Figures at a glance
| Table 1 |
Figures at a glance
 |
 |
 |
 |
 |
||||
| Figure 1 | Figure 2 | Figure 3 | Figure 4 | Figure 5 |
Relevant Topics
Recommended Journals
Article Tools
Article Usage
- Total views: 13459
- [From(publication date):
January-2013 - Nov 09, 2025] - Breakdown by view type
- HTML page views : 8874
- PDF downloads : 4585
