Research Article Open Access
Increased Oxidative Damage in RNA in Alzheimer’s Disease Progression
Bradley-Whitman MA1 and Lovell MA1,2*1Sanders-Brown Center on Aging and Alzheimer’s Disease Center, University of Kentucky, USA
2Department of Chemistry, University of Kentucky, USA
- *Corresponding Author:
- Mark A Lovell, PhD
Sanders-Brown Center on Aging
University of Kentucky, Lexington, KY 40536, USA
Tel: 859-257-1412 (251)
Fax: 859-323-2866
E-mail: malove2@uky.edu
Received date: September 21, 2013; Accepted date: October 28, 2013; Published date: October 30, 2013
Citation: Bradley-Whitman MA, Lovell MA (2013) Increased Oxidative Damage in RNA in Alzheimer’s Disease Progression. J Anal Bioanal Tech S2:004. doi: 10.4172/2155-9872.S2-004
Copyright: © 2013 Bradley-Whitman MA, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Journal of Analytical & Bioanalytical Techniques
Abstract
Studies of oxidative damage during the progression of Alzheimer’s Disease (AD) suggest a central role in disease pathogenesis. To determine if RNA oxidation increases in the progression of AD levels of oxidized bases from RNA from the superior and middle temporal gyri (SMTG), inferior parietal lobule (IPL), and cerebellum (CER) throughout the progression of AD including subjects with mild cognitive impairment (MCI), preclinical AD (PCAD), late-stage AD (LAD), diseased control (DC) (Frontotemporal Dementia (FTD) and Dementia with Lewy Bodies (DLB)), and agematched normal control subjects (NC) were analyzed by gas chromatography mass spectrometry. Median levels of multiple RNA adducts were significantly (p<0.05) elevated in the SMTG, IPL, and CER in multiple stages of AD and in DC subjects. Elevated levels of 4,6-diamino-5-foramidopyrimidine (FapyA) suggest a hypoxic environment early in the progression of AD and in DC subjects. Overall, these data suggest oxidative damage is an early event not only in the pathogenesis of AD, but is present in neurodegenerative diseases in general.
Keywords
RNA; Mild cognitive impairment; Preclinical Alzheimer’s disease; Alzheimer’s disease; Neurodegenerative diseases; RNA oxidation; Gas chromatography-mass spectrometry
Introduction
Alzheimer’s Disease (AD), the most common form of dementia, is estimated to currently affect ~5 million (1 in 9) Americans aged 65 years or older [1]. Antemortem, AD is characterized by a progressive decline in both cognitive and non-cognitive domains following an insidious onset. Despite research endeavors by the scientific community, the etiology of AD remains unclear effectively hindering potential advancements. The study of the underlying disease etiology and pathogenesis of AD is complicated by the multifaceted nature of the disease on a molecular level.
Initial AD diagnostic criteria developed by the Neurological and Communicative Disorders Association (NINCDS) and the Alzheimer’s disease and Related Disorder Association (ADRDA) workgroup in 1984 proposed that the clinical manifestations were synonymous with the underlying pathogenesis of the disease. Subjects with mild cognitive impairment (MCI) experience a perceived and verifiable change in cognition but are not demented and maintain functional independence antemortem [2-4] and postmortem exhibit AD associated pathology but to a lesser extent than demented AD subjects [5]. More recently postmortem analyses have described the presence of senile plaques (SP) and neurofibrillary tangles (NFT) in the brains of cognitively normal subjects suggestive of an extended ‘preclinical AD’ (PCAD) stage of AD in which extensive pathology exists in the absence of obvious clinical manifestations [6-9].
Although multiple hypotheses have been proposed to explain the pathogenesis of AD, no single hypothesis fully encompasses both the clinical and pathological features of AD. There is the distinct possibility that a multifaceted hypothesis can more effectively account for pathogenesis of AD. One particular hypothesis that has received considerable interest is the oxidative damage hypothesis, an extension of the free radical theory of aging (FRTA) initially proposed by Harman [10,11] that attempts to unify themes of AD including the formation of SP and NFT, mitochondrial dysfunction, neuronal excitotoxicity, and neurodegeneration. Although the FRTA proposes that oxidative stress or free radical generation increases throughout the body, the brain is particular vulnerable due to the increased oxygen consumption/high energy demands and limited antioxidant defense relative to other tissue types. Increased generation or prolonged exposure to reactive oxygen species (ROS) is thought to play a role in a wide range of oxidative damage observed in both early and late stages of AD [12] resulting in compromised cellular function.
Aerobic oxidative phosphorylation by mitochondria is essential to meet energetic demands of neurons. Unfortunately, mitochondrial by-products of cellular respiration are considered to be the primary endogenous source of ROS, including hydrogen peroxide (H2O2), superoxide (O2•-), and hydroxyl (HO•) radicals [13]. It is estimated that ~2% of total oxygen consumed by the cell is converted to superoxide radical during respiration [14]. Although superoxide can be detoxified to water by sequential activities of superoxide dismutase and glutathione peroxidase, in the presence of reduced transition metals (copper and iron) hydrogen peroxide can generate hydroxyl radicals via Fenton [15] and/or Haber-Weiss reactions [16]. Increased oxidative damage to mitochondrial DNA (mtDNA),(~10X levels observed in nuclear DNA (nDNA)) is attributed to the close proximity to ROS generation and a lack of protective histones [14,17] and may contribute to the decreased glucose consumption and depletion of cellular energy noted in the earliest stages of AD [18-20]. Although multiple ROS are produced in the brain, the highly reactive hydroxyl radical is the primary initiator of damage to heterocyclic nucleic acids and associated sugar moieties at or near diffusion controlled rates [21,22]. Because markers of oxidative damage are an indirect measurement of oxidative damage, it remains unclear if the oxidative damage observed in AD is the direct result of increased generation or reduced efficiency in elimination or some combination thereoy.
Multiple by-products of ROS-mediated damage to lipids, proteins, and DNA have been studied during the pathogenesis of AD. Oxidation of nucleic acids, initiated by highly reactive hydroxyl radical is a result in over 20 potential adducts [23]. However, modification of guanine is considered the preeminent marker of nucleic acid oxidation because of its low reduction potential. Levels of oxidized guanine have been reported to be significantly increased in both nDNA [24-27] and mtDNA [26] in the vulnerable brain regions in AD.
In contrast, quantification of nucleic acid adducts in RNA has not been reported in either the early or late stages of AD. To fully characterize the extent of nucleic acid oxidation in RNA during the progression of AD, levels of multiple nucleic acid adducts were quantified from two vulnerable brain regions, the superior and middle temporal gyri (SMTG) and inferior parietal lobule (IPL), and a nonvulnerable brain region, the cerebellum (CER) of subjects with mild cognitive impairment (MCI), preclinical AD (PCAD), late-stage AD (LAD), diseased control (DC) subjects that included frontotemporal dementia (FTD) and dementia with Lewy bodies (DLB), and agematched normal control subjects (NC) by gas chromatography mass spectrometry (GC/MS) operated in selective ion monitoring mode (SIM).
Methods
Brain specimen sampling
Tissue specimens were obtained through the Neuropathology Core of the University of Kentucky Alzheimer’s Disease Center (UK-ADC) under University of Kentucky IRB approval protocols. Tissue specimens of short postmortem interval (PMI) autopsies from seven subjects with MCI (2 men [M]: 5 women [W]), fourteen PCAD subjects (3 M: 11 W), fifteen LAD subjects (7 M: 8 W), six DLB subjects (4 M: 2 W), six FTD subjects (3 M: 3 W), and fifteen age-matched NC subjects (4 M: 11 W) were flash frozen in liquid nitrogen and maintained at -80°C until processed for analysis.
All subjects were characterized as previously described [28] based on existing criteria for MCI [4], PCAD [6,29], and LAD [30,31]. Postmortem DLB pathology was characterized by the presence of ubiquitin and α-synuclein positive Lewy bodies (LB) and by ubiquitin positive Lewy neurites (LN) in a semi-quantitative grading density in paralimbic and neocortical structures. DLB diagnosis was according to the current guidelines proposed by the International Workshop for the Consensus Guidelines for the Clinical and Pathological Diagnosis of DLB [32,33]. Postmortem DLB was characterized by symmetric focal atrophy of the frontal and/or temporal lobes, atrophy of the basal ganglia, and loss of pigmentation in the substantia nigra. Hematoxylin and eosin (H&E) staining of the cerebral cortex highlight microvacuolation and neuron loss, whereas immunostaining for α-β-crystallin reveal swollen cortical neurons. Pathological diagnosis of FTD was based on current standards [34,35]. Neuritic plaque (NP) and Braak staging scores for all subjects were provided by the Neuropathology Core of the UK-ADC.
RNA isolation
RNA samples were prepared as described by Chomczynski and Sacchi [36]. Briefly, tissue specimens (~100 mg) were homogenized in 1 ml of TRI ReagentR in a polypropylene microcentrifuge tube and incubated at room temperature for 5 min. Chloroform (200 μL) was added, mixed by hand, incubated for 5 min at room temperature, and centrifuged at 9000×g for 15 min at 4°C. Isopropyl alcohol (500 μL) was added to the resulting clear aqueous phase containing RNA, incubated at room temperature for 10 min and centrifuged at 9000×g for 10 min at 4°C. RNA pellets were washed in 75% ethanol, air dried, and stored at -80°C until analysis. Samples were resuspended in autoclaved distilled/ deionized water and the concentration and purity determined using a NanoDrop 1000 Spectrophotometer (NanoDrop, Wilmington, DE, USA).
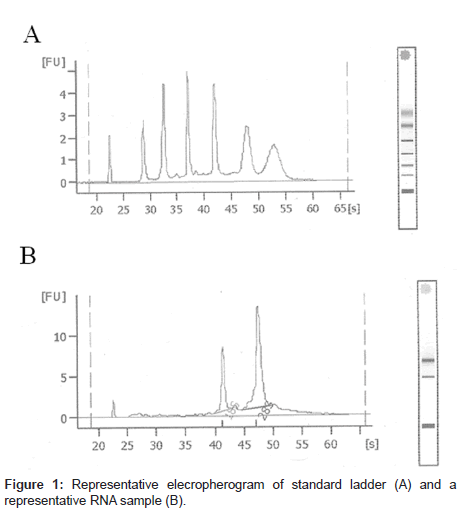
RNA integrity
Representative RNA isolates were submitted to the UK MircoArray Core Facility for electrophoretic analysis. Sharp peaks representing the eukaryotic 18S and 28S ribosomal subunits along with the absence of smaller well-defined peaks between the two ribosomes peaks are characteristics associated with high quality isolates. In addition, the RNA integrity number (RNI), an algorithm based product, designed to communicate the level of RNA degradation, on a scale of 1 (degraded) to 10 (intact) was also determined.
RNA derivatization
For analysis, RNA (100 μg) samples were prepared for GC/MS analysis as previously described by Wang et al. [26,37]. Ten nmol of each stable-labeled internal standard: 8-[8-13C, 7, 9-15N2] hydroxyguanine; 8-[8-13C, 6, 9-diamino-15N2] hydroxyadenine; 5-[2-13C, 1, 3-15N2] hydroxycytosine; [formyl-13C, diamino-15N2] fapyadenine; [formyl- 13C, diamino-15N2] fapyguanine were added for quantification of oxidized bases. Briefly, samples were subjected to acid hydrolysis with formic acid (250 μL, 90%) at 145°C for 30 min in evacuated tubes. Following lyophlization individual bases were derivatized with a pyridine:N,O-bis(trimethylsilyl)trifluoroacetamide (BSTFA)/1% trimethylchlorosilane (TMCS) solution (200 μL, 1:1) in evacuated sealed tubes for 2 hr at room temperature. The derivatized samples were dried under a stream of nitrogen in a water bath at 37°C, and 20 μL BSTFA/1% TMCS added immediately before GC/MS injection.
Gas chromatography/mass spectrometry (GC/MS) with selective ion monitoring (SIM)
Derivatized samples were analyzed using an Agilent 7890A gas chromatograph on a HP-5ms capillary column (0.25 mm internal diameter, 0.25 μm film thickness, and 30 m length; Hewlett Packard, Palo Alto, CA, USA) as previously described by Wang et al. [26,37]. Derivatized nitrogenous base spectra were acquired in SIM mode at the following m/z ratios: 443 (8-[8-13C,7,9-15N2] hydroxyguanine) and 440 (8-hydroxyguanine (8OHG)); 445 ([formyl-13C, diamino- 15N2] fapyguanine) and 442 (fapyguanine (FapyG)); 355 (8-[8-13C,6,9- diamino-15N2] hydroxyadenine) and 352 (8-hydroxyadenine (8OHA)); 357 ([formyl-13C, diamino-15N2] fapyadenine) and m/z 354 (fapyadenine (FapyA)); and 331 (5-[2-13C,1,3-15N2] hydroxycytosine) and m/z 328 (5-hydroxycytosine (5OHC)). Instrument response plots of integrated peak intensities for stable-labeled analyte signals were determined over a range of 0.5 nmol to 100 nmol per stable labeled isotope standard. The integrated area of each analyte was normalized with respect to the integrated area of the corresponding internal standard for all samples and corrected based on instrument response plots for a given internal standard.
Measurement of Aβ1-40 and Aβ1-42 levels
Total levels of Aβ1-40 and Aβ1-42 in phosphate buffered saline (PBS)-, sodium dodecyl sulfate (SDS)-, and formic acid (FA)-soluble fractions were provided by the Sanders-Brown Center on Aging Amyloid Core as previously described [38] for the SMTG, IPL, and CER. Briefly, tissue samples (200 mg) were homogenized in PBS with a complete protease inhibitor cocktail (PIC) via apolytron and centrifuged at 20,800×g for 30 min at 4°C. The supernatant (PBS-soluble pool) was collected and the remaining pellet sonicated (10×0.5 sec pulses at 100 W) in 2% (w/v) SDS with PIC and centrifuged at 20,800×g for 30 min at 14°C. The supernatant (SDS-soluble pool) was collected and the remaining pellet was sonicated as previously described in 70% (v/v) formic acid, and centrifuged at 20,800×g for 60 min at 4°C. The supernatant (FA-soluble pool) was collected. All samples were stored at -80°C until analysis.
Oligomeric Aβ was quantified as previously described using a single-site sandwich ELISA (4G8/4G8). Immulon 4HBX plates were coated with 0.5 μg/well antibody 4G8 (Covance, Princeton, NJ, USA) and incubated overnight at 4°C. Wells were blocked with Synblock® (AbD Seroctec; Oxford, UK) according to manufacturer’s instructions. Standard curves were prepared using synthetic Aβ peptide (rPeptide; Bogart, GA, USA). PBS samples were diluted 1:4 in Antigen Capture (AC) buffer (20 mM Na3PO4, 0.4% Block Ace (AbD Serotec), 0.05% NaN3, 2 mM EDTA, 0.4 M NaCl, 0.2% BSA, 0.05% CHAPS, pH 7). SDS samples were diluted between 1:20 and 1:100 in AC buffer and formic acid samples were first neutralized 1:20 in TP buffer (1.0 M Tris base, 0.5 M Na2HPO4) then further diluted between 1:5 and 1:20 in AC buffer. Standards and samples were analyzed in duplicate and were incubated at 4°C overnight. Biotinylated detection antibodies were Ab13.1.1 (Aβ1-40-end specific), 12F4 (Aβ1-42-end specific), or 4G8 (Aβ17-24-end specific) followed by the addition of NeurtAvidin-HRP (Pierce Biotechnologies, Rockford, IL, USA). Colorimetric detection used 3,3′,5,5′-tetramethylbenzidine reagent (TMB; Kirkegaard and Perry Laboratories; Gaithersburg, MD, USA). The reaction was stopped via acidification (6% o-phosphoric acid) and read using a BioTech Powerwave XS (Winooski, VT, USA) plate reader at 450 nm.
Statistics
All data were tested for normality using the Wilkes-Shapiro test. Levels of oxidized RNA ribonucleic acid adducts, PBS-, SDS-, and FA-soluble Aβ1-40 and Aβ1-42, and Braak staging scores demonstrated non-normal distributions and were analyzed using the Mann-Whitney U-test and are reported as median values with range. All other data including, age and PMI, demonstrated a normal distribution and are reported as mean ± standard error of mean (SEM). No significant differences were observed for any measure between FTD and DLB subjects, therefore the two neurodegenerative diseases were pooled to create a diseased control (DC) subject pool. All statistical comparisons were carried out using Sigma Plot. Statistical significance was set at p ≤ 0.05 and trending toward significance was set at p ≤ 0.1.
The magnitude of Pearson’s r value was determined for levels of nucleic acid adducts and neuropathological markers including, NFT counts, SP counts, and levels of PBS-, SDS-, and FA- soluble Aβ1-40 and Aβ1-42. Type one errors arising from multiple comparisons per brain region studied were corrected through the use of the Bonferroni method. Recalculated P-values for significance for each brain region studied were 0.0045 (SMTG), 0.00625 (IPL), and 0.0045 (CER).
Results
Subject demographic data are shown in Table 1. Neither PMI nor age of subjects were significantly different except for a significant (P<0.05) increase in the age of PCAD (85.8 ± 1.7 y), MCI subjects (91.0 ± 1.9 y), LAD (80.8 ± 1.4 y) and NC subjects (86.3 ± 1.4 y) compared to DC subjects (68.9 ± 16.6 y) and a significant increase in the age of MCI subjects compared to LAD subject. Median Braak staging scores were significantly higher for MCI (IV), PCAD (V), LAD (VI), and DC subjects (II) compared to age-matched NC subjects (I).
| Group | Mean ± SEM age (years) | Sex | Mean ± SEM PMI (hr) | Median Braak Score |
| NC | 86.3 ± 1.4 | N=15; 4M, 11W | 2.8 ± 0.2 | I |
| PCAD | 85.8 ± 1.7a | N=14; 3M, 11W | 2.8 ± 0.2 | Vc |
| MCI | 91.0 ± 1.9a,b | N=7; 2M, 5W | 2.6 ± 0.2 | IVc |
| LAD | 80.8 ± 1.4a | N=15; 7M, 8W | 3.6 ± 0.4 | VIc |
| DC | 68.9 ± 4.8 | N=12; 7M, 5W | 3.5 ± 0.4 | IIc |
aP < 0.05 compared to DC
bP < 0.05 compared to LAD
cP ≤ 0.05 compared to NC
NC=Normal Control; MCI=Mild Cognitive Impairment; PCAD=Preclinical Alzheimer’s Disease; LAD=Late-stage Alzheimer’s Disease; DC=Diseased Control; PMI=Postmortem Interval.
Table 1: Subject demographic.
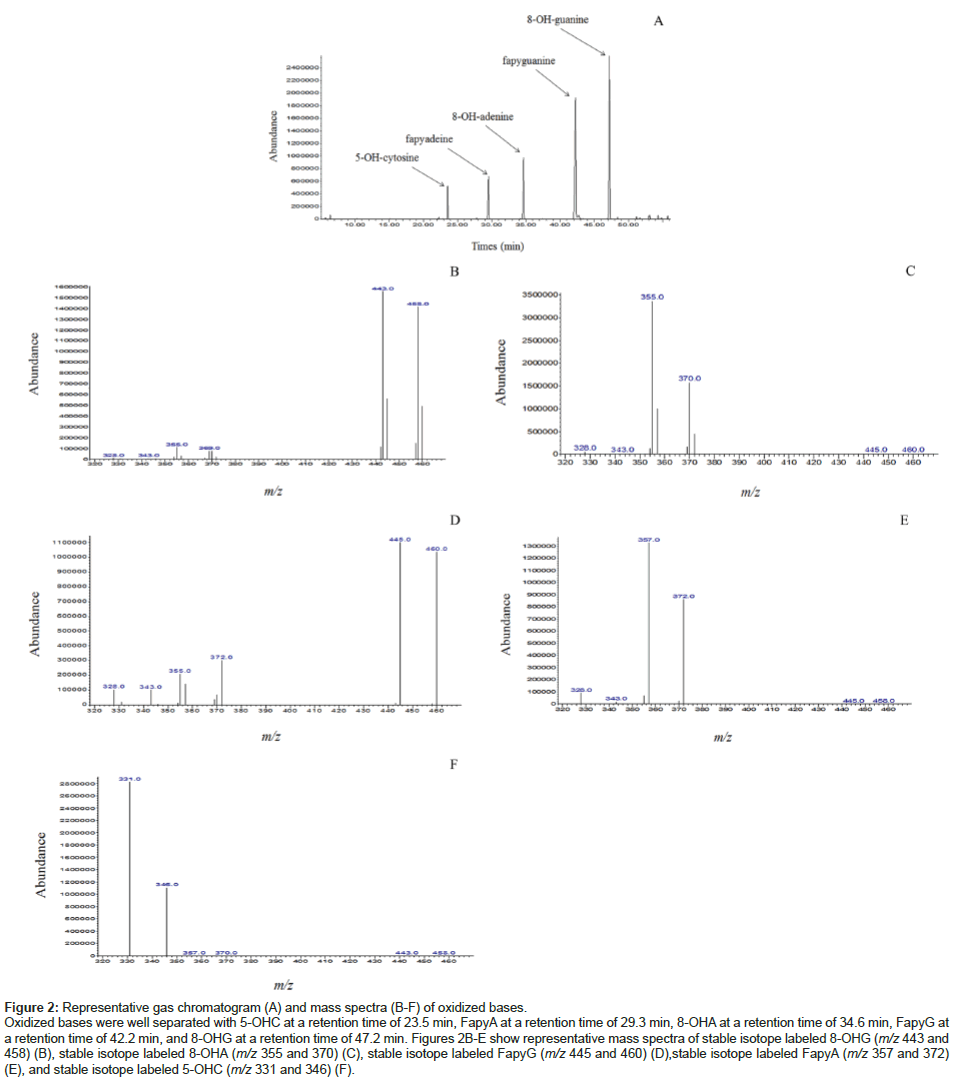
Representative RNA isolates exhibited signature characteristics, including 18S and 28S ribosomal RNA units and high RINs, indicating a highly pure and intact RNA (Figure 1). Oxidized RNA adducts quantified were based on previous studies that indicated significant changes in oxidized nucleic acids of nDNA and mtDNA between agematched NC subjects and LAD subjects. The five stable isotope labeled internal standards were well separated chromatographically at the following retention times: 8-[8-13C, 7, 9-15N2] hydroxyguanine (47.2 min), [formyl-13C, diamino-15N2] fapyguanine (42.2 min), 8-[8-13C, 6, 9-diamino-15N2] hydroxyadenine (34.6 min), [formyl-13C, diamino- 15N2] fapyadenine (29.3 min), and 5-[2-13C, 1, 3-15N2] hydroxycytosine (23.5 min) (Figure 2A).
Three replicate analyses were performed to determine the dynamic range of each stable isotope labeled internal standard. Positive significant correlations were observed between the instrument response over a range of 500 pmol to 100 nmol for: 8-[8-13C, 7, 9-15N2] hydroxyguanine (r=0.99), [formyl-13C, diamino-15N2] fapyguanine (r=0.99), 8-[8-13C, 6, 9-diamino-15N2] hydroxyadenine (r=0.97), [formyl-13C, diamino- 15N2] fapyadenine (r=0.99), 5-[2-13C, 1,3-15N2] hydroxycytosine (r=0.99). Representative mass spectra for each stable labeled adduct of interest are shown in Figures 2B-2F. For quantitative purposes the dominant daughter ion for each oxidized ribonucleic acid adducts were monitored. To illustrate the percentage of change, levels of ribonucleic acid oxidation are expressed levels as median [range] % of NC subjects. Median levels of oxidized bases measured in each identifiable stage of AD and in DC subjects are shown in Table 2.
| Brain Region | ||||
| Lesion | Group | SMTG | IPL | CER |
| 8OHG | NC | 100.0 [33.6-205.3] | 100.0 [12.3-293.5] | 100.0 [38.0-109.3] |
| PCAD | 113.0 [38.3-388.8] | 96.2 [23.0-254.9] | 133.6 [87.5-165.0] | |
| MCI | 230.5 [53.8-573.1]* | 124.5 [26.6-276.8] | 117.3 [69.6-261.4] | |
| LAD | 599.3 [106.6-2174.3]* | 117.5 [27.6-297.0] | 89.3 [66.1-196.8] | |
| DC | 344.6 [32.9-841.3]ŧ | 100.5 [30.2-280.6] | 330.4 [113.3-1067.1]* | |
| FapyG | NC | 100.0 [24.9-202.8] | 100.0 [45.0-199.4] | 100.0 [57.6-263.5] |
| PCAD | 113.5 [61.5-196.2] | 118.4 [12.4-200.3] | 150.1 [82.2-548.7]* | |
| MCI | 60.2 [24.2-105.5] | 121.7 [85.0-148.9] | 119.2 [57.2-611.4] | |
| LAD | 102.5 [52.8-212.6] | 141.8 [105.7-517.3]* | 171.4 [83.3-876.2]* | |
| DC | 84.2 [47.6-131.4] | 115.8 [80.4-139.0] | 193.6 [83.2-341.8]* | |
| 8OHA | NC | 100.0 [44.3-208.4] | 100.0 [50.0-522.7] | 100.0 [19.0-274.2] |
| PCAD | 169.5 [84.8-252.9]* | 152.4 [57.5-799.5] | 196.2 [44.5-3893.9] | |
| MCI | 187.5 [47.8-884.6]ŧ | 93.7 [27.9-491.6] | 120.9 [108.2-168.8] | |
| LAD | 138.4 [81.5-397.7]ŧ | 97.9 [54.7-172.9] | 143.3 [56.1-2372.8] | |
| DC | 120.2 [57.6-458.7]ŧ | 97.6 [10.8-579.7] | 237.4 [53.8-2716.6] | |
| FapyA | NC | 100.0 [45.1-159.8] | 100.0 [27.3-364.0] | 100.0 [41.4-238.9] |
| PCAD | 70.7 [45.2-160.1] | 276.8 [206.4-366.6]* | 133.7 [6.7-543.0] | |
| MCI | 60.0 [27.9-186.5] | 328.5 [215.7-370.5]* | 106.3 [27.0-554.5] | |
| LAD | 72.0 [40.3-168.6] | 316.7 [182.7-810.2]* | 71.6 [23.9-239.8] | |
| DC | 72.8 [44.6-107.1] | 288.1 [196.2-375.3]* | 82.6 [44.9-307.6] | |
| 5OHC | NC | 100.0 [76.6-160.3] | 100.0 [20.4-233.8] | 100.0 [31.6-246.9] |
| PCAD | 137.4 [71.6-328.5] | 94.4 [71.0-170.3] | 122.9 [77.7-264.4] | |
| MCI | 75.3 [48.5-216.3] | 81.8 [71.4-91.4] | 108.7 [74.0-175.4] | |
| LAD | 100.9 [71.0-220.1] | 105.9 [55.2-235.1] | 107.4 [66.0-161.8] | |
| DC | 85.5 [69.0-177.4] | 112.1 [61.4-237.9] | 190.6 [138.6-513.0]* | |
*P ≤ 0.05 compared to NC
ŧP ≤ 0.1 compared to NC
Table 2: Levels of oxidized nucleic acid base adducts in RNA expressed as % of NC (Median [Range]).
Figure 2: Representative gas chromatogram (A) and mass spectra (B-F) of oxidized bases.
Oxidized bases were well separated with 5-OHC at a retention time of 23.5 min, FapyA at a retention time of 29.3 min, 8-OHA at a retention time of 34.6 min, FapyG at a retention time of 42.2 min, and 8-OHG at a retention time of 47.2 min. Figures 2B-E show representative mass spectra of stable isotope labeled 8-OHG (m/z 443 and 458) (B), stable isotope labeled 8-OHA (m/z 355 and 370) (C), stable isotope labeled FapyG (m/z 445 and 460) (D),stable isotope labeled FapyA (m/z 357 and 372) (E), and stable isotope labeled 5-OHC (m/z 331 and 346) (F).
Significant (p ≤ 0.05) elevations in median levels of 8-OHG were observed in the SMTG of MCI and LAD subjects compared to NC subjects. Although median levels of 8-OHG in DC subjects were not significantly different, levels trended toward significance (p ≤ 0.1) compared to NC subjects. In contrast, no significant changes were detected in PCAD in the SMTG. No significant changes in median levels of 8-OHG were observed in the IPL in any stage of AD pathogenesis or in DC subjects. Median levels of 8-OHG in the CER were not significantly altered during the pathogenesis of AD but were significantly (p ≤ 0.05) increased in DC subjects compared to NC subjects. Unlike median levels of 8-OHG, no significant alterations in median levels of FapyG were detected in the SMTG during the pathogenesis of AD or in DC subjects compared to NC subjects. In contrast, FapyG median levels were significantly elevated (p ≤ 0.05) in the IPL of LAD subjects compared to NC subjects. In the CER, median levels of FapyG were significantly (p ≤ 0.05) elevated in both PCAD, and LAD subjects compared to NC subjects. Furthermore, median levels of FapyG were significantly increased in the CER of DC subjects compared to NC subjects.
Significant (p ≤ 0.05) elevations of median levels of 8-OHA were observed in only PCAD subjects, although they trended toward significance (p ≤ 0.1) in MCI, LAD, and DC subjects compared to NC subjects. Similarly to 8-OHG, no significant changes in the median level of 8-OHA were noted during the progression of AD or in DC subjects compared to NC subjects in the IPL or CER. Median levels of FapyA were not significantly different in the SMTG of any disease stage studied but were significantly elevated in the two early stages (PCAD and MCI) as well as LAD and DC subjects compared to NC subjects in the IPL. No significant changes were noted in the CER during AD pathogenesis or in DC subjects compared to NC subjects.
Median levels of 5-OHC were not significantly different in any brain region studied, although median levels were significantly (p ≤ 0.05) elevated in the CER only of DC subjects compared to NC subjects.
Median levels of PBS-, SDS-, and FA soluble Aβ1-40 and Aβ1-42 in each identifiable stage of AD and in DC subjects are shown in Tables 3-5. In the SMTG, levels of PBS-soluble Aβ1-40 were significantly (p ≤ 0.05) elevated in both MCI and LAD subjects compared to NC subjects. Median levels of PBS-soluble Aβ1-42 were significantly (p ≤ 0.05) elevated in only LAD subjects compared to NC subjects. No significant changes in median levels of PBS-soluble Aβ1-40 or Aβ1-42 were detected in the IPL or CER. In the SMTG, median levels of SDS-soluble Aβ1-42 were significantly (p ≤ 0.05) increased in only LAD subjects compared to NC subjects. No significant changes in the median levels of SDSsoluble Aβ1-42 were detected in either the IPL or CER. Furthermore, no significant changes in median levels of SDS-soluble Aβ1-40 were detected in any brain region studied during the progression of AD or in DC subjects compared to NC subjects. Median levels of FA-soluble Aβ1-40 were significantly (p ≤ 0.05) elevated in only the IPL of LAD subjects but no other brain region studied compared to NC subjects. In contrast, median levels of FA-soluble Aβ1-40 were significantly (p ≤ 0.05) elevated in the SMTG of MCI subjects and the each brain region studied in LAD subjects compared to NC subjects.
| Brain Region | ||||
|---|---|---|---|---|
| Aβ | Group | SMTG | IPL | CER |
| PBS Aβ1-40 | NC | 0.0 [0.0-179.0] | 52.7 [0.0-103.5] | 3.4 [0.0-30.6] |
| PCAD | 12.8 [0-122.6] | 0.0 [0.0-257.4] | 14.75 [0.0-66.5] | |
| MCI | 50.6 [0-66.4]* | 29.9 [0.0-117.1] | 0.0 [0.0-10.8] | |
| LAD | 80.6 [10.8-281.1]* | 163.4 [125.2-1412.3] | 58.2 [0-601.6] | |
| DC | 22.7 [6.8-34.6] | 0.0 [0-173.2] | 0.0 [0.0-10.8] | |
| PBS Aβ1-42 | NC | 75.8 [0.0-209.7] | 0.0 [0.0-118.6] | 131.6 [60.2-298.2] |
| PCAD | 175.6 [31.9-249.0] | 0.0 [0.0-88.2] | 136.7 [57.6-327.9] | |
| MCI | 118.9 [12.7-347.8] | 0.0 [0.0-825.8] | 0.0 [0.0-157.7] | |
| LAD | 196.7 [154.4-255.5]* | 0.0 [0.0-454.8] | 213.0 [161.8-292.9] | |
| DC | 3.1 [0-252.3] | 0.0 [0.0-254.3] | 151.2 [44.8219.5] | |
*P ≤ 0.05 compared to NC
Table 3: Levels of PBS Aβ1-40 and Aβ1-42 expressed as Median [Range] in pM.
| Brain Region | ||||
| Aβ | Group | SMTG | IPL | CER |
| SDS Aβ1-40 | NC | 1367.4 [0-2884.7] | 1454.5 [0.0-3594.0] | 1169.3 [346.3-4707.3] |
| PCAD | 207.0 [0-945.4] | 1972.2 [0.0-5634.8] | 1085.8 [665.4-1668.7] | |
| MCI | 286.5 [107.5-625.5] | 1258.3 [0.0-18587.3] | 1025.6 [865.4-1427.1] | |
| LAD | 6966.2 [169.4-22297.8] | 19241.1 [3740.9-27165.0] | 5645.0 [0-9880.0] | |
| DC | 306.4 [28.0-505.8] | 1546.2 [1125.4-4587.6] | 905.4 [545.6-1025.6] | |
| SDS Aβ1-42 | NC | 1456.9 [0.0-4509.1] | 0.0 [0.0-60988.6] | 1285.8 [0-2483.0] |
| PCAD | 1415.2 [0.0-5349.2] | 0.0 [0.0-29160.9] | 0.0 [0.0-127.2] | |
| MCI | 737.4 [0.0-3312.2] | 658.5 [0.0-54992.3] | 0.0 [0.0-95.3] | |
| LAD | 61694.1 [12198.5-111060.8]* | 116829.1 [0.0-124769.7] | 6634.3 [5162.2-7106] | |
| DC | 0.0 [0.0-0.0] | 75.6 [0.0-125.6] | 0.0 [0.0-0.0] | |
*P ≤ 0.05 compared to NC
Table 4: Levels of SDS-soluble Aβ1-40 and Aβ1-42 expressed as Median [Range] in pM.
| Brain Region | ||||
| Aβ | Group | SMTG | IPL | CER |
| FA Aβ1-40 | NC | 0.0 [0.0-1307.8] | 1945.1 [0.0-22562.5] | 0.0 [0.0-1960.8] |
| PCAD | 1226.3 [737.5-2206.0] | 41694.6 [0.0-88404.7] | 1226.3 [900.4-2124.0] | |
| MCI | 2371.1 [1637.8-4575.4] | 21804.2 [0.0-140836.8] | 1800.8 [661.4-2615.6] | |
| LAD | 13081.6 [3470.4-572858.8] | 400792.0 [253539.4-955235.6]* | 5855.8 [0-114149.1] | |
| DC | 1715.8 [1226.3-2287.7] | 1432.1 [0.0-2546.2] | 818.9 [574.8-1063.3] | |
| FA Aβ1-42 | NC | 3662.0 [0.0-61454.0] | 28757.8 [0.0-111658.2] | 3912.4 [0-4079.4] |
| PCAD | 799.4 [0.0-18514.6] | 77825.6 [0.0-437983.8] | 0.0 [0.0-1245.0] | |
| MCI | 4721.9 [0.0-60228.4]* | 222308.0 [17327.6-51564.44] | 0.0 [0.0-0.0] | |
| LAD | 81655.3 [169.4-153773.5]* | 1009392.9 [911145.7-1703310.5]* | 19994.7 [0.0-23007.5]* | |
| DC | 0.0 [0.0-7520.4] | 15243.2 [0.0-256462] | 0.0 [0.0-0.0] | |
Table 5: Levels of FA soluble Aβ1-40 and Aβ1-42 expressed as Median [Range] in pM.
Levels of oxidized bases measured were correlated with PBS-, SDS-, and FA soluble Aβ1-40 and Aβ1-42 in each brain region studied and a summary of significant correlations are shown in Table 6. In the SMTG, significant positive correlations between levels of median levels of 8-OHA and PBS-soluble Aβ1-42 (r=0.526: p=0.002) and median levels of 8-OHG and FA-soluble Aβ1-42 (r=0.592: p=0.0004) were observed. In contrast, there was a significant negative correlation between levels of FapyG and PBS-soluble Aβ1-42 (r=-0.508: p=0.004). In the IPL, median levels of 8-OHG were significantly correlated with FA-soluble Aβ1-40 (r=0.55: p=0.005) and FA-soluble Aβ1-42 (r=0.6: p=0.006). Median levels of FapyG were significantly correlated with SDS-soluble Aβ1-42 (r=0.653: p=0.002), SDS-soluble Aβ1-42 (r=0.773: p=0.00006), FA-soluble Aβ1-40 (r=0.887: p=0.0000002), and FA-soluble Aβ1-42 (r=0.730: p=0.0008). In the CER there was a significant positive correlation between the median levels of FapyG and levels of FA soluble Aβ1-42 (r=0.645: P=0.00005).
| Brain Region | |||
| Aβ | SMTG | IPL | CER |
| PBS Aβ1-40 | - | - | - |
| PBS Aβ1-42 | 8-OHA r = 0.526 p = 0.002; FapyG r = -0.508 p = 0.004 | - | - |
| SDS Aβ1-40 | - | FapyG r = 0.653 p = 0.002 | - |
| SDS Aβ1-42 | - | FapyG r = 0.773 p = 0.00006 | - |
| FA Aβ1-40 | - | 8-OHG r = 0.55 p = 0.005; FapyG r = 0.887 p = 0.0000002 |
- |
| FA Aβ1-42 | 8-OHG r = 0.592 p = 0.0004 | 8-OHG r = 0.60 p = 0.006; FapyG r = 0.730 p = 0.0008 |
FapyG r = 0.645 p=0.00005 |
Table 6: Summary of Significant Correlations between levels of PBS-, SDS- and FA-soluble Aβ1-40 and Aβ1-42 and Individual RNA Adducts.
Discussion
Despite extensive study, the etiology of AD remains unclear. The indirect relationship between clinical manifestations and pathological features and an extended prodromal phase challenges researchers. In the current study, two distinct early stages of AD progression were studied including subjects who demonstrate substantial AD pathology but remain cognitively normal (PCAD) and the earliest detectable clinical stage (MCI) concurrent with LAD subjects in both vulnerable and non-vulnerable brains regions. Furthermore, inclusion of FTD and DLB subjects as a pooled diseased control group allowed us to assess RNA oxidation in additional neurodegenerative diseases to determine if RNA oxidation is unique to AD.
To our knowledge, this study is the first to quantify levels of multiple oxidized nucleic acids in RNA throughout the pathogenesis of AD as well as in DC subjects (DLB and FTD). In an effort to minimize potential analytical biases, subjects from each group and multiple brain regions were analyzed in parallel. We analyzed nucleotides isolated from bulk tissue composed of a mixture of glia, neurons and vasculature, however previous immunohistochemical studies by Nunomura et al. [39] demonstrated that 8-hydroxyguanine immunoreactivity was restricted to neuronal cells. Yet, the detection of multiple mRNA species sequestered in SPs of AD [40] suggest total RNA isolated during that bulk tissue processing includes remnant RNA incorporated in SP following neuron degeneration. Furthermore, the current study did not distinguish between different types of RNA isolated. Therefore median levels reported here represent total RNA including messenger (mRNA), ribosomal (rRNA), and transfer (tRNA). Previous studies by Shan and Lin [41] reported mRNA accounts for less than 5% of total RNA isolates and reported significant oxidation of polyribosomal RNA fractions 40S, 60S, and 80S in LAD [42] suggesting changes observed in their study likely reflect rRNA and tRNA oxidation.
Although pure intact RNA as indicated by a RIN of 9.8 was analyzed, nucleic acid adduct quantification by GC/MS as described in the current study, has the potential to introduce artifactual oxidation during isolation of as well as derivitization [43-45]. Although the use of phenol during nucleic acid isolation has been suggested to contribute to artifactual oxidation, Wang et al. [26] demonstrated that detectable DNA lesions were not significantly different when extracted with a phenol:chloroform extraction compared to the popular NaI based ‘salting out’ method consistent with the findings of Rehamn et al. [46]. Furthermore, atmospheric oxygen was excluded during nucleic acid derivatization and carried out at room temperature to minimize artifactual oxidation [46-48].
Similar to previous studies that demonstrated increased nucleic acid in both nuclear and mitochondrial DNA [25,37,49], we found that median levels of multiple oxidized RNA base adducts were significantly elevated in vulnerable brain regions in LAD consistent with Weidner et al. [38] who observed increased levels of 8-hydroxyadenine in RNA from LAD subjects. Additionally, median levels of 8-OHG and 8-OHA were significantly increased or trended toward significance in the SMTG. These findings are consistent with those of Nunomura et al. [50] who demonstrated increased RNA oxidation in vulnerable neurons of MCI subjects in temporal cortex pyramidal neurons suggesting the earliest clinically identifiable stage of AD is characterized by nucleic acid oxidation. Additionally, median levels of 8-OHA were significantly elevated in PCAD subjects suggesting RNA oxidation may be a feature of the prodromal phase of AD (PCAD) consistent with Nunomura et al. [50] who reported RNA oxidation is an event preceding a transition from normal aging during the extended prodromal phase of AD.
While 8-hydroxyguanine is considered to be the most common marker of oxidative damage, our study suggests levels of 8-hydroxyguanine are not significantly different than levels of 8-hydroxyadenine. Although elevated levels of 8OHG and FapyA were detected in both early stages of AD (PCAD and MCI), levels were not significantly different from those in LAD subjects. These findings suggest RNA oxidation reaches a peak early in AD progression without further significant elevation later in disease. Furthermore, significantly elevated levels of 8-hydroxyguaine and 8-hydroxyadenine observed in DC subjects were comparable to levels observed in both early and late stages of AD suggesting oxidation is a common feature of neurodegeneration, rather than an AD specific event. These findings are consistent with a previous study of nuclear and mitochondrial DNA nucleic acid oxidation in the same subject pool [51].
Interestingly, levels of FapyA were only significantly increased in the IPL. Formation of FapyA following the initial insult is dependent on environment conditions and elevation suggests damage occurred in a hypoxic environment. Again median levels of nucleic acid oxidation observed in PCAD and MCI subjects were not significantly different from levels observed in LAD subjects. Additionally, levels of FapyA in the IPL were also significantly increased in DC subjects. Collectively these data suggest a hypoxic environment may be an early event in AD pathogenesis but is not an exclusive feature of AD but rather a common feature of neurodegenerative disease consistent with reports of RNA oxidation in Down syndrome [52,53], familial AD [54], DLB [55], and Parkinson’s disease [56].
The cerebellum is widely regarded as an internal control with minimal changes associated with AD. However, in the current study levels of FapyG were significantly elevated in the CER of PCAD, LAD, and DC brain. While it remains unclear why significant oxidation was detected in the CER, studies of nuclear DNA and mitochondrial DNA isolated from the same subject pool also exhibited elevated levels of multiple markers of oxidative damage in the CER [51]. Furthermore, Bradley et al. [28] showed levels of extractable 4-hydroxyhexenal were significantly increased in the CER of LAD subjects compared to age-matched NC subjects ina subset of subjects described in the current study. Additionally, Braak et al. [57] and Yamaguchi et al. [58] described the presence of diffuse amyloid plaques in the molecular layer of the cerebellar cortex of AD subjects and Wegiel et al. [59] reported a volumetric loss in both the molecular and granular layers. Shrinkage of the posterior cerebellar region of AD has been associated with poorer cognitive performance [60]. Furthermore, elevated levels of 8-hydroxyguanine and FapyG were observed in the CER of DC subjects, consistent with FTD associated loss of the Purkinje cells [61] and detection of α-synuclein positive Purkinje cells in DLB subjects [62]. Significant oxidation of nucleic acids in the CER of DC subjects suggests the phenomenon is not AD specific.
Correlation analyses between median levels of RNA base adducts and neuropathological features including NP and NFT densities, and PBS-, SDS-, and FA- soluble Aβ1-42 and Aβ1-40 were determined for the SMTG, IPL, and CER. No significant correlations were detected between NP or NFT counts. Although multiple significant correlations between RNA adducts and specific pools of Aβ1-40 and Aβ1-42 in AD levels of multiple RNA lesions were also found to be significantly elevated in DC subjects whose levels of Aβ1-40 and Aβ1-42 were comparable to NC subjects, suggesting that the observed relationships are not a direct cause and effect consistent with previous reports that suggest nucleic acid oxidation is independent of amyloid deposition [51,52]. However, because the current study quantified median levels of RNA oxidation determined from bulk tissue, we were unable to address the hypothesis that intraneuronal non-oligomeric Aβ peptide may act as an antioxidant as supported by Nunomura et al. [63,64] who found an inverse relationship between staining intensities of oxidized guanine and levels of intraneuronal non-oligomeric Aβ peptide and NFT tangle formation.
It has been proposed that the transient nature of mRNA may function as an oxidative sponge. However, because preparations of RNA analyzed are attributed to longer lived ribosomal RNA, this hypothesis was not addressed. Yet, if most RNA lesions detected in the current study are a direct result of ribosomal and transfer RNA oxidation rather than messenger RNA, it would suggest that RNA repair mechanisms may exist. Unfortunately, while expression and associated activity of multiple substrate-specific DNA glycosylases involved in DNA base excision repair have been widely studied, few have focused on the repair of oxidative damage in RNA [15,65-68]. However, it would be expected that oxidized RNA would not be tolerated under normal cellular conditions without deleterious effects. For example, direct oxidation of the firefly luciferase messenger RNA resulted in the premature termination of translation and subsequent decreased expression following transfection in HEK293 cells [69]. However, decreased levels of total RNA oxidation 24-72 hours after oxidative insults have been reported [70,71] suggesting that, although the mechanism is unclear, repair of RNA lesions does occur. Furthermore, LAD associated ribosomal dysfunction was related to decreased levels of rRNA and tRNA and increased levels of oxidation [72].
Conclusion
Consistent with our previous study of nuclear and mitochondrial DNA, the current data suggest RNA oxidative damage is an early event in the pathogenesis of AD. Furthermore, levels of RNA oxidation are comparable between both early stages of AD progression and LAD and DC subjects suggesting that nucleic acid oxidation is a common feature of neurodegeneration rather than an AD exclusive feature.
Conflict of Interest
The authors have no conflicts of interest.
Acknowledgements
This research was supported by NIH grants 5P01-AG05119 and P30- AG028383. The authors would like to thank Dr. M. Paul Murphy of Sanders-Brown Center on Aging Amyloid Core for Aβ1-40 and Aβ1-42 measurements. The authors would also like to thank Ms. Sonya Anderson for subject demographic data and Ms. Paula Thomason for editorial assistance.
References
- Hebert LE, Weuve J, Scherr PA, Evans DA (2013) Alzheimer disease in the United States (2010-2050) estimated using the 2010 census. Neurology 80: 1778-1783.
- Albert MS, DeKosky ST, Dickson D, Dubois B, Feldman HH, et al. (2011) The diagnosis of mild cognitive impairment due to Alzheimer's disease: recommendations from the National Institute on Aging-Alzheimer's Association workgroups on diagnostic guidelines for Alzheimer's disease. Alzheimers Dement 7: 270-279.
- Jack CR Jr, Albert MS, Knopman DS, McKhann GM, Sperling RA, et al. (2011) Introduction to Revised Criteria for the Diagnosis of Alzheimer’s Disease: National Institute on Aging and the Alzheimer Association Workgroups. Alzheimers Dement 7: 257-262.
- Petersen RC, Smith GE, Waring SC, Ivnik RJ, Tangalos EG, et al. (1999) Mild cognitive impairment: clinical characterization and outcome. Arch Neurol 56: 303-308.
- Markesbery WR, Schmitt FA, Kryscio RJ, Davis DG, Smith CD, et al. (2006) Neuropathologic substrate of mild cognitive impairment. Arch Neurol 63: 38-46.
- Schmitt FA, Davis DG, Wekstein DR, Smith CD, Ashford JW, et al. (2000) "Preclinical" AD revisited: neuropathology of cognitively normal older adults. Neurology 55: 370-376.
- Davis DG, Schmitt FA, Wekstein DR, Markesbery WR (1999) Alzheimer neuropathologic alterations in aged cognitively normal subjects. J Neuropathol Exp Neurol 58: 376-388.
- Knopman DS, Parisi JE, Salviati A, Floriach-Robert M, Boeve BF, et al. (2003) Neuropathology of cognitively normal elderly. J Neuropathol Exp Neurol 62: 1087-1095.
- Price JL, Morris JC (1999) Tangles and plaques in nondemented aging and "preclinical" Alzheimer's disease. Ann Neurol 45: 358-368.
- Harman D (1956) Aging: a theory based on free radical and radiation chemistry. J Gerontol 11: 298-300.
- Harman D (2006) Alzheimer's disease pathogenesis: role of aging. Ann N Y Acad Sci 1067: 454-460.
- Markesbery WR (1997) Oxidative stress hypothesis in Alzheimer's disease. Free Radic Biol Med 23: 134-147.
- Wallace DC (1999) Mitochondrial diseases in man and mouse. Science 283: 1482-1488.
- Ames BN, Shigenaga MK, Hagen TM (1993) Oxidants, antioxidants, and the degenerative diseases of aging. Proc Natl Acad Sci U S A 90: 7915-7922.
- Imam SZ, Karahalil B, Hogue BA, Souza-Pinto NC, Bohr VA (2006) Mitochondrial and nuclear DNA-repair capacity of various brain regions in mouse is altered in an age-dependent manner. Neurobiol Aging 27: 1129-1136.
- Mello Filho AC, Meneghini R (1984) In vivo formation of single-strand breaks in DNA by hydrogen peroxide is mediated by the Haber-Weiss reaction. Biochim Biophys Acta 781: 56-63.
- Wallace DC (1992) Mitochondrial genetics: a paradigm for aging and degenerative diseases? Science 256: 628-632.
- Arnáiz E, Jelic V, Almkvist O, Wahlund LO, Winblad B, et al. (2001) Impaired cerebral glucose metabolism and cognitive functioning predict deterioration in mild cognitive impairment. Neuroreport 12: 851-855.
- Drzezga A, Lautenschlager N, Siebner H, Riemenschneider M, Willoch F, et al. (2003) Cerebral metabolic changes accompanying conversion of mild cognitive impairment into Alzheimer's disease: a PET follow-up study. Eur J Nucl Med Mol Imaging 30: 1104-1113.
- Fukuyama H, Ogawa M, Yamauchi H, Yamaguchi S, Kimura J, et al. (1994) Altered cerebral energy metabolism in Alzheimer's disease: a PET study. J Nucl Med 35: 1-6.
- Dizdaroglu M, Jaruga P (2012) Mechanisms of free radical-induced damage to DNA. Free Radic Res 46: 382-419.
- Evans MD, Dizdaroglu M, Cooke MS (2004) Oxidative DNA damage and disease: induction, repair and significance. Mutat Res 567: 1-61.
- Cooke MS, Evans MD, Dizdaroglu M, Lunec J (2003) Oxidative DNA damage: mechanisms, mutation, and disease. FASEB J 17: 1195-1214.
- Gabbita SP, Lovell MA, Markesbery WR (1998) Increased nuclear DNA oxidation in the brain in Alzheimer's disease. J Neurochem 71: 2034-2040.
- Lovell MA, Gabbita SP, Markesbery WR (1999) Increased DNA oxidation and decreased levels of repair products in Alzheimer's disease ventricular CSF. J Neurochem 72: 771-776.
- Wang J, Markesbery WR, Lovell MA (2006) Increased oxidative damage in nuclear and mitochondrial DNA in mild cognitive impairment. J Neurochem 96: 825-832.
- Mecocci P, MacGarvey U, Kaufman AE, Koontz D, Shoffner JM, et al. (1993) Oxidative damage to mitochondrial DNA shows marked age-dependent increases in human brain. Ann Neurol 34: 609-616.
- Bradley MA, Xiong-Fister S, Markesbery WR, Lovell MA (2012) Elevated 4-hydroxyhexenal in Alzheimer's disease (AD) progression. Neurobiol Aging 33: 1034-1044.
- Schmitt FA, Nelson PT, Abner E, Scheff S, Jicha GA, et al. (2012) University of Kentucky Sanders-Brown healthy brain aging volunteers: donor characteristics, procedures and neuropathology. Curr Alzheimer Res 9: 724-733.
- Hyman BT, Phelps CH, Beach TG, Bigio EH, Cairns NJ, et al. (2012) National Institute on Aging-Alzheimer's Association guidelines for the neuropathologic assessment of Alzheimer's disease. Alzheimers Dement 8: 1-13.
- Montine TJ, Phelps CH, Beach TG, Bigio EH, Cairns NJ, et al. (2012) National Institute on Aging-Alzheimer's Association guidelines for the neuropathologic assessment of Alzheimer's disease: a practical approach. Acta Neuropathol 123: 1-11.
- McKeith IG, Dickson DW, Lowe J, Emre M, O'Brien JT, et al. (2005) Diagnosis and management of dementia with Lewy bodies: third report of the DLB Consortium. Neurology 65: 1863-1872.
- McKeith IG, Galasko D, Kosaka K, Perry EK, Dickson DW, et al. (1996) Consensus guidelines for the clinical and pathologic diagnosis of dementia with Lewy bodies (DLB): report of the consortium on DLB international workshop. Neurology 47: 1113-1124.
- Cairns NJ, Bigio EH, Mackenzie IR, Neumann M, Lee VM, et al. (2007) Neuropathologic diagnostic and nosologic criteria for frontotemporal lobar degeneration: consensus of the Consortium for Frontotemporal Lobar Degeneration. Acta Neuropathol 114: 5-22.
- Neary D, Snowden JS, Gustafson L, Passant U, Stuss D, et al. (1998) Frontotemporal lobar degeneration: a consensus on clinical diagnostic criteria. Neurology 51: 1546-1554.
- Chomczynski P, Sacchi N (1987) Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal Biochem 162: 156-159.
- Wang J, Xiong S, Xie C, Markesbery WR, Lovell MA (2005) Increased oxidative damage in nuclear and mitochondrial DNA in Alzheimer's disease. J Neurochem 93: 953-962.
- Weidner AM, Bradley MA, Beckett TL, Niedowicz DM, Dowling AL, et al. (2011) RNA oxidation adducts 8-OHG and 8-OHA change with Aβ42 levels in late-stage Alzheimer's disease. PLoS One 6: e24930.
- Nunomura A, Perry G, Pappolla MA, Wade R, Hirai K, et al. (1999) RNA oxidation is a prominent feature of vulnerable neurons in Alzheimer's disease. J Neurosci 19: 1959-1964.
- Ginsberg SD, Crino PB, Hemby SE, Weingarten JA, Lee VM, et al. (1999) Predominance of neuronal mRNAs in individual Alzheimer's disease senile plaques. Ann Neurol 45: 174-181.
- Shan X, Lin CL (2006) Quantification of oxidized RNAs in Alzheimer's disease. Neurobiol Aging 27: 657-662.
- Ding Q, Markesbery WR, Cecarini V, Keller JN (2006) Decreased RNA, and increased RNA oxidation, in ribosomes from early Alzheimer's disease. Neurochem Res 31: 705-710.
- Cadet J, Douki T, Gasparutto D, Ravanat JL (2003) Oxidative damage to DNA: formation, measurement and biochemical features. Mutat Res 531: 5-23.
- Halliwell B, Dizdaroglu M (1992) The measurement of oxidative damage to DNA by HPLC and GC/MS techniques. Free Radic Res Commun 16: 75-87.
- Collins A, Cadet J, Epe B, Gedik C (1997) Problems in the measurement of 8-oxoguanine in human DNA. Report of a workshop, DNA oxidation, held in Aberdeen, UK, 19-21 January, 1997. Carcinogenesis 18: 1833-1836.
- Rehman A, Jenner A, Halliwell B (2000) Gas chromatography-mass spectrometry analysis of DNA: optimization of protocols for isolation and analysis of DNA from human blood. Methods Enzymol 319: 401-417.
- Dizdaroglu M, Gajewski E (1990) Selected-ion mass spectrometry: assays of oxidative DNA damage. Methods Enzymol 186: 530-544.
- Jenner A, England TG, Aruoma OI, Halliwell B (1998) Measurement of oxidative DNA damage by gas chromatography-mass spectrometry: ethanethiol prevents artifactual generation of oxidized DNA bases. Biochem J 331 : 365-369.
- Gabbita SP, Aksenov MY, Lovell MA, Markesbery WR (1999) Decrease in peptide methionine sulfoxide reductase in Alzheimer's disease brain. J Neurochem 73: 1660-1666.
- Nunomura A, Tamaoki T, Motohashi N, Nakamura M, McKeel DW Jr, et al. (2012) The earliest stage of cognitive impairment in transition from normal aging to Alzheimer disease is marked by prominent RNA oxidation in vulnerable neurons. J Neuropathol Exp Neurol 71: 233-241.
- Bradley-Whitman MA, Timmons MD, Beckett TL, Murphy MP, Lynn BC, et al. (2013) Nucleic acid oxidation: an early feature of Alzheimer's disease. J Neurochem .
- Nunomura A, Perry G, Pappolla MA, Friedland RP, Hirai K, et al. (2000) Neuronal oxidative stress precedes amyloid-beta deposition in Down syndrome. J Neuropathol Exp Neurol 59: 1011-1017.
- Nunomura A, Perry G, Hirai K, Aliev G, Takeda A, et al. (1999) Neuronal RNA oxidation in Alzheimer's disease and Down's syndrome. Ann N Y Acad Sci 893: 362-364.
- Nunomura A, Chiba S, Lippa CF, Cras P, Kalaria RN, et al. (2004) Neuronal RNA oxidation is a prominent feature of familial Alzheimer's disease. Neurobiol Dis 17: 108-113.
- Nunomura A, Chiba S, Kosaka K, Takeda A, Castellani RJ, et al. (2002) Neuronal RNA oxidation is a prominent feature of dementia with Lewy bodies. Neuroreport 13: 2035-2039.
- Zhang J, Perry G, Smith MA, Robertson D, Olson SJ, et al. (1999) Parkinson's disease is associated with oxidative damage to cytoplasmic DNA and RNA in substantia nigra neurons. Am J Pathol 154: 1423-1429.
- Braak H, Braak E, Bohl J, Lang W (1989) Alzheimer's disease: amyloid plaques in the cerebellum. J Neurol Sci 93: 277-287.
- Yamaguchi H, Hirai S, Morimatsu M, Shoji M, Nakazato Y (1989) Diffuse type of senile plaques in the cerebellum of Alzheimer-type dementia demonstrated by beta protein immunostain. Acta Neuropathol 77: 314-319.
- Wegiel J, Wisniewski HM, Dziewiatkowski J, Badmajew E, Tarnawski M, et al. (1999) Cerebellar atrophy in Alzheimer's disease-clinicopathological correlations. Brain Res 818: 41-50.
- Thomann PA, Schläfer C, Seidl U, Santos VD, Essig M, et al. (2008) The cerebellum in mild cognitive impairment and Alzheimer's disease - a structural MRI study. J Psychiatr Res 42: 1198-1202.
- Hayashi S, Toyoshima Y, Hasegawa M, Umeda Y, Wakabayashi K, et al. (2002) Late-onset frontotemporal dementia with a novel exon 1 (Arg5His) tau gene mutation. Ann Neurol 51: 525-530.
- Mori F, Piao YS, Hayashi S, Fujiwara H, Hasegawa M, et al. (2003) Alpha-synuclein accumulates in Purkinje cells in Lewy body disease but not in multiple system atrophy. J Neuropathol Exp Neurol 62: 812-819.
- Nunomura A, Tamaoki T, Tanaka K, Motohashi N, Nakamura M, et al. (2010) Intraneuronal amyloid beta accumulation and oxidative damage to nucleic acids in Alzheimer disease. Neurobiol Dis 37: 731-737.
- Nunomura A, Perry G, Aliev G, Hirai K, Takeda A, et al. (2001) Oxidative damage is the earliest event in Alzheimer disease. J Neuropathol Exp Neurol 60: 759-767.
- Szczesny B, Bhakat KK, Mitra S, Boldogh I (2004) Age-dependent modulation of DNA repair enzymes by covalent modification and subcellular distribution. Mech Ageing Dev 125: 755-765.
- Mao G, Pan X, Zhu BB, Zhang Y, Yuan F, et al. (2007) Identification and characterization of OGG1 mutations in patients with Alzheimer's disease. Nucleic Acids Res 35: 2759-2766.
- Shao C, Xiong S, Li GM, Gu L, Mao G, et al. (2008) Altered 8-oxoguanine glycosylase in mild cognitive impairment and late-stage Alzheimer's disease brain. Free Radic Biol Med 45: 813-819.
- Weissman L, Jo DG, Sørensen MM, de Souza-Pinto NC, Markesbery WR, et al. (2007) Defective DNA base excision repair in brain from individuals with Alzheimer's disease and amnestic mild cognitive impairment. Nucleic Acids Res 35: 5545-5555.
- Tanaka M, Chock PB, Stadtman ER (2007) Oxidized messenger RNA induces translation errors. Proc Natl Acad Sci U S A 104: 66-71.
- Kajitani K, Yamaguchi H, Dan Y, Furuichi M, Kang D, et al. (2006) MTH1, an oxidized purine nucleoside triphosphatase, suppresses the accumulation of oxidative damage of nucleic acids in the hippocampal microglia during kainate-induced excitotoxicity. J Neurosci 26: 1688-1698.
- Shen Z, Wu W, Hazen SL (2000) Activated leukocytes oxidatively damage DNA, RNA, and the nucleotide pool through halide-dependent formation of hydroxyl radical. Biochemistry 39: 5474-5482.
- Ding Q, Markesbery WR, Chen Q, Li F, Keller JN (2005) Ribosome dysfunction is an early event in Alzheimer's disease. J Neurosci 25: 9171-9175.
Relevant Topics
Recommended Journals
Article Tools
Article Usage
- Total views: 16505
- [From(publication date):
specialissue-2014 - Dec 21, 2025] - Breakdown by view type
- HTML page views : 11737
- PDF downloads : 4768