Research Article Open Access
Mitochondrial DNA Haplogroups in Patients with Alzheimer Probable in Guadalajara, Mexico
Genaro Gabriel Ortiz1*, Verónica Loera-Castañeda1, Lucila Sandoval-Ramírez2, Miguel Ángel Macías-Islas3, Gracia Viviana González- Enríquez1, Luis J. Flores-Alvarado1,4, Irma E. Velázquez-Brizuela5,
Alfredo Celis de la Rosa4 and Fermín P. Pacheco-Moisés6
1Laboratorio de Desarrollo-Envejecimiento; Enfermedades Neurodegenerativas. División de Neurociencias. Centro de Investigación Biomédica de Occidente. Instituto Mexicano del Seguro Social. Guadalajara, Jalisco México
2Laboratorio de Bioquímica IV. División de Genética. Centro de Investigación Biomédica de Occidente. Instituto Mexicano del Seguro Social. Guadalajara, Jalisco. México
3Departamento de Neurología. UMAE-Hospital de Especialidades del Centro Médico Nacional de Occidente. Instituto Mexicano del Seguro Social. Guadalajara, Jalisco. México
4Centro Universitario de Ciencias de la Salud. Universidad de Guadalajara, Guadalajara, Jalisco. México.
5Instituto Jalisciense de Cancerología OPD-SSA. Guadalajara, Jalisco. México
6Departamento de Química, Centro Universitario de Ciencias Exactas e Ingenierías. Universidad de Guadalajara, Guadalajara, Jalisco. México
- Corresponding Author:
- Genaro Gabriel Ortiz
Laboratorio de Desarrollo- Envejecimiento
Enfermedades Neurodegenerativas
Centro de Investigación Biomédica de Occidente
Instituto Mexicano del Seguro Social
Sierra Mojada 800 cp 43440 Guadalajara, Jalisco, México
E-mail: genarogabriel@yahoo.com
Received date: October 20, 2011; Accepted date: November 09, 2011; Published November 11, 2011
Citation: Ortiz GG (2011) Mitochondrial DNA Haplogroups in Patients with Alzheimer Probable in Guadalajara, México. J Alzheimers Dis 1:103. doi:
Copyright: © 2011 Ortiz GG. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Journal of Alzheimers Disease & Parkinsonism
Abstract
Background and aims . It has been suggested that mitochondrial dysfunction is relevant to Alzheimer disease (AD) and some mitochondrial haplogroups could be related to the risk of AD. Therefore, in this work we have genotyped mtDNA haplogroups in Mexican patients with AD and their respective matched controls. Methods: Haplotyping of 33 AD patients and 49 healthy control subjects was performed using polymerase chain reaction/digest identification of key polymorphic sites in the mitochondrial genome. Results: Mitochondrial haplogroup A is the most frequent in AD patients and controls followed by haplogroup B. The haplogroup C was the least frequent in the control group and it was absent in the AD group. The frequency of haplogroup D was significantly higher in the AD group than in the control group (p = 0.03). Conclusion: There is clear association between the frequency of haplogroup D and Alzheimer disease in the analyzed population from Guadalajara, Mexico. Interestingly the haplogroup C was absent in the AD group. These data requires further study in a larger population to identify these differences more clearly.
Keywords
Alzheimer, Mitochondria, Haplogroups, PCR, Genome, Population
Abbreviations
AD: Alzheimer Disease; ATP: Adenosine Triphosphate; mtDNA: Mitochondrial DNA; Apo: Apolipoprotein
Introduction
Alzheimer’s disease (AD) is a genetically heterogeneous and insidiously progressive form of dementia in which more than 50 genetic loci are involved [1].Therefore, AD may represent a common phenotype that results from various genetic and environmental influences. Although the aetiology of AD remains largely unclear, mounting evidence suggests that mitochondrial dysfunction could be involved in AD pathogenesis because they can be both targets of oxidative damage and sources of reactive oxygen species [6].
Mitochondria are the cellular organelles that perform the metabolic reactions necessary to generate energy as adenosine triphosphate (ATP) and contain their own DNA in a single circular chromosome (mtDNA), and their own machinery for RNA and protein synthesis [14]. mtDNA has unique genetic features. It is independently replicated, transcribed, and translated, is maternally inherited, does not recombine, and has a very high mutation rate, which contributes to the variability [9]. The mtDNA single nucleotide polymorphisms have been accumulated sequentially along radiating female lineages, giving rise to a wide variety of haplogroups. Because of the central role of mitochondria in cell function and apoptosis it has been hypothesized that the characterisation of these haplogroups could contribute to the understanding of the expression of mitochondrial-related diseases, and carriers of a particular allele or haplogroup could be at a higher or lower risk for some multifactor diseases [9,11] i.e., the mitochondrial haplogroup in an individual may protect against or predispose AD. A prediction of this hypothesis is that the mitochondrial lineage represented by the mtDNA haplogroup is different in AD patients and controls.
To our knowledge no studies analysing the role of common mitochondrial polymorphisms in the risk of developing AD have been published for the Mexican population. On the other hand, four major founding mtDNA haplogroups (A, B, C, D), which account together for 97% of modern American indian mtDNA´s has been reported. Therefore the aim of the present work was to investigate the frequency and possible association of these mitochondrial haplogroups in AD patients and control groups from Guadalajara, México.
Materials and Methods
Patients
33 AD subjects were recruited from the Clínica de Trastornos Cognitivos y Demencias at the Unidad Médica de Alta Especialidad del Hospital de Especialidades del Centro Médico Nacional de Occidente at Guadalajara, Jalisco, México. All patients met NINCDS-ADRDA criteria for probable AD and they did not manifest signs or symptoms of an alternative neurodegenerative disorder. An age-matched control group of 49 healthy subjects were also examined. Informed assent and consent were obtained from the study participants or their caregiver to collect DNA for genetic studies. This study was approved by the Comisión Nacional de Investigación Científica Review board and was carried out in accordance with the principles of the Declaration of Helsinki as revised in 2000.
DNA purification and haplogroups
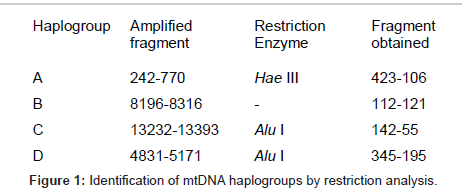
DNA was obtained from 5 ml of venous blood following the Miller procedure [8] and purified by ethanol precipitation and its concentration was determined by spectrophotometry UV absorption at 260 nm/280 nm. mtDNA haplogroup typing was carried out by restriction analyses. Briefly, mtDNA specific primers were used to amplify by polymerase chain reaction the mtDNA regions containing the polymorphic restriction sites that characterize each haplogroup. The amplified fragments, after digestion by appropriate restriction enzymes (Figure 1), were separated by agarose gel electrophoresis. Fragments visualised after ethidium bromide staining.
Haplogroups are defined by nucleotides at specific known polymorphic sites in the mtDNA. The polymorphisms analysed in this study define the four major haplogroups that have proved to be the progenitors of virtually all Native American mtDNA´s. After genotyping these polymorphisms, the haplogroup of each patient and control was determined.
Statistical analysis
The binomial analysis between groups was analyzed by Pearson correlation. The significance level was (p<0.05).
Results
It has been suggested that the assessment of global mtDNA variation is essential for elucidating the role of mtDNA mutations in disease. Therefore we genotyped the four major mtDNA haplogroups of Native Americans in a sample from Guadalajara, Jalisco. México. The prevalence of haplogroups A, B, C, and D mtDNA´s in Native American populations has now been confirmed [10]. In addition it has been reported that in healthy people of the North-Central Mexico the haplogroup A is the most frequent, followed in descending order, by the B, C and D haplogroups [5]. While Torroni et al., [12] in 4 native populations from Southern México, reported only the haplogroups A, B and C. In the population analyzed from the west of México, we found the four major haplogroups of Asian origin.
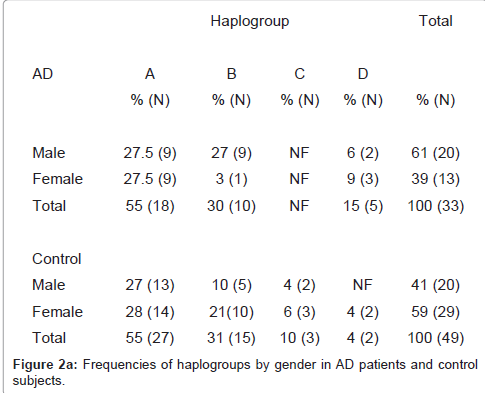
A total of 33 AD patients and 49 healthy control subjects were studied. In the AD group, there were 13 (39.4%) women and 20 (60.6%) men, whereas 29 (59.2%) women and 20 (41) men where control subjects. The mean age of the AD population was 65 ± 13.6 years. (47-89 years) whereas that the control group was 65 ± 13.2 years. (47-89 years) (Figure 2a)
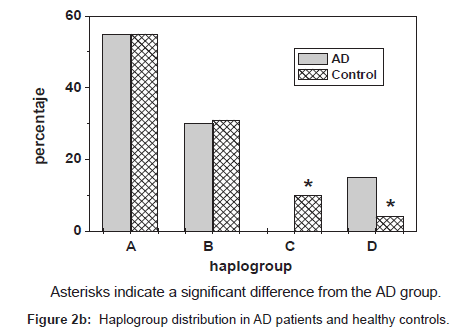
In the healthy population we found the haplogroups A, B, C and D. Frequency of haplogroup A was higher than haplogroups B and D in AD patients (55%, 30%, and 15%. respectively). Similar frequencies for haplogroups A and B were found in healthy control subjects (55% and 31%, respectively). Haplogroup D showed higher frequency in AD patients than healthy controls (15% and 4%, respectively). Frequency of haplogroup C in the healthy control group was 10% and was absent in the AD population. In addition haplogroup D was absent in healthy control male gender (Figure 2b).
Binomial analysis of the data showed significant differences between the haplogrous found in patients with AD compared with the control group (p=0.03) (Figure 2b). No significant differences between haplogroups and gender in each of the groups studied were found.
Discussion
The role of mtDNA haplogroups in the neurodegenerative cascade leading to AD is still a matter of controversy and it has been hypothesized that patients with certain mitochondrial haplogroups are more susceptible to malignant mitochondrial diseases (e.g. Leber’s hereditary optic neuropathy, Alzheimer’s disease, multiple sclerosis and Parkinson’s disease in Caucasians [11]. For example, Carrieri et al. [10] suggested that K and U haplogroups may act by neutralising the effect of the major AD risk factor apolipoprotein E (ApoE) 4 allele, as they are present at a lower frequency in E4 carriers than in noncarrier AD patients, while in controls there is association between E4 allele and mtDNA Haplogroups [2]. Van der Walt JM, found that males classified as haplogroups U had a significant increase in risk of AD [13]. On the other hand, Chinnery PF, Elson et al. and Mancuso [12-14] found no significant haplogroup associations in the AD and normal control groups [3,4,7].
The present data suggest that mitochondrial haplogroup C was a protective factor for AD and males classified as haplogroup D showed an increase in risk of AD. However, our study was based on a limited number of patients, and other studies with larger samples and other populations should be necessary to fully understand the role of mtDNA on AD. In addition, a complete sequencing of the mitochondrial genome in individuals with haplogroup C should be necessary to define the nucleotide changes directly responsible for this effect. These doubts will be resolved in subsequent work in our laboratory.
Acknowledgements
This Work was supported by CONACyT-México. Fondo Sectorial Salud 1419- 5 and 7479-5.
References
- Cacabelos R (2003) The application of functional genomics to Alzheimer's disease. Pharmacogenomics 4: 597-621.
- Mancuso M, Coppede F, Murri L, Siciliano G (2007) Mitochondrial cascade hypothesis of Alzheimer's disease: myth or reality? Antioxid Redox Signal 9: 1631-1646.
- Wallace DC (1994) Mitochondrial DNA sequence variation in human evolution and disease. Proc Natl Acad Sci USA 91: 8739-8746.
- Parsons TJ, Muniec DS, Sullivan K, Woodyatt N, Alliston-Greiner R, et al. (1997) A high observed substitution rate in the human mitochondrial DNA control region. Nat Genet 15: 363-368.
- Shoffner JM, Brown MD, Torroni A, Lott MT, Cabell MF, et al. (1993) Mitochondrial DNA variants observed in Alzheimer disease and Parkinson disease patients. Genomics 17: 171-184.
- Wallace DC (1995) Mitochondrial DNA variation in human evolution, degenerative disease and aging. Am J Hum Genet 57: 201-223
- Miller SA, Dykes DD, Polesky HF (1998) A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res 16: 1215.
- Santos M, Barrantes R (1994) D-Loop mtDNA deletion is a unique marker of Chibchan Americans. Am J Hum Genet 55 :413-414
- Torroni A, Chen YS, Semino O, Santachiara-Beneceretti AS, Scott CR, et al. (1994) mtDNA and Y-chromosome polymorphisms in four Native American populations from southern Mexico. Am J Hum Genet 54: 303-318.
- Carrieri G, Bonafe M, De Luca M, Rose G, Varcasia O, et al. (2001) Mitochondrial DNA haplogroups and APOE4 allele are non-independent variables in sporadic Alzheimer's disease. Hum Genet 108(3): 194-198.
- Van der Walt JM, Dementieva YA, Martin ER, Scott WK, Nicodemus KK, et al. (2004) Analysis of European mitochondrial haplogroups with Alzheimer disease risk. Neurosci Lett 365: 28-32.
- Chinnery PF, Taylor GA, Howell N, Andrews RM, Morris CM, et al. (2000) Mitochondrial DNA haplogroups and susceptibility to AD and dementia with Lewy bodies. Neurology 55: 302-304.
- Elson JL, Herrnstadt C, Preston G, Thal L, Morris CM, et al. (2006) Does the mitochondrial genome play a role in the etiology of Alzheimer's disease? Hum Genet 119: 241-254.
- Mancuso M, Nardini M, Micheli D, Rocchi A, Nesti C, et al. (2007) Lack of association between mtDNA haplogroups and Alzheimer's disease in Tuscany. Neurol Sci 28: 142-147.
Relevant Topics
- Advanced Parkinson Treatment
- Advances in Alzheimers Therapy
- Alzheimers Medicine
- Alzheimers Products & Market Analysis
- Alzheimers Symptoms
- Degenerative Disorders
- Diagnostic Alzheimer
- Parkinson
- Parkinsonism Diagnosis
- Parkinsonism Gene Therapy
- Parkinsonism Stages and Treatment
- Stem cell Treatment Parkinson
Recommended Journals
Article Tools
Article Usage
- Total views: 14795
- [From(publication date):
November-2011 - Nov 26, 2025] - Breakdown by view type
- HTML page views : 10097
- PDF downloads : 4698