Effects of Putrescine on Anti-Oxidative Enzymes in Two Rice Cultivars Subjected To Salinity
Received: 22-Jan-2016 / Accepted Date: 10-Feb-2016 / Published Date: 17-Feb-2016 DOI: 10.4172/2329-8863.1000210
Abstract
A study was conducted for elucidation of expression of isozymic profiles with respect to salinity along with putrescine application in two rice varieties cv. Nonabokra and cv. Swarna at 200mM NaCl alone and with 2mM putrescine. Preliminarily these two varieties displays differential pattern of accumulation of Na+ as revealed from SEM micrograph studies. There recorded significant variation in activities in-vitro as well as by in-gel studies of Guaiacol Peroxidase (GPX), Ascorbate Peroxidase (APX), Superoxide Dismutase (SOD), Catalase (CAT) and Glutathione Reductase (GR) enzymes. Activities for GPX, APX and SOD followed significant up regulation under salinity. In contrary CAT and GR enzymes were subdued in both varieties. Putrescine improved the activity for SOD, GPX and APX. CAT and GR maintained stable activity with putrescine. A number of isozymic bands were found with the induction of salinity and putrescine treatment. For SOD three distinct bands were recorded as Cu/Zn/Mn-Fe SOD. For GPX and APX multiple bands were revealed in activity gel. On the contrary CAT was insensitive with the putrescine induction and hardly there recorded any variation. GR isozyme was more prominent in band intensities both in salinity and putrescine. Conclusively different isozymic profiles have contributed to resist salinity along with putrescine.
Keywords: Anti-oxidative enzymes; Polyamine; Rice; Salinity; SEM
404100Abbreviations
CAT: Catalase; GPX: Guiacol Peroxidase; APX: Ascorbate Peroxidase; SOD: Superoxide Dismutase; GR: Glutathione Reductase; DCPIP: Dichlorophenol indophenol; MTT: 3-(4,5-dimethyl)-2 thiazolyl 2,5 diphenyl-H tetrazolium bromine; NBT: NitroTetrazolium Blue
Introduction
A variety of endogenous substances are found in plant system that essentially happens to be inducers or modulators for number of physiological processes. Polyamines being one of those are characterized by polycationic, low molecular weight, and straight chain aliphatic amines. Since, polyamines are positive in charge, it has an in built character to bind with different macromolecules like membrane phospholipids, nucleic acids, proteins, wall constituting residues and other moieties with negatively charged domain. In fact, the main functioning of polyamines is attributed for stabilizing the biological membrane those are prone to oxidative damages under various abiotic stresses. On the other hand, polyamines have also been implicated as a secondary messenger to perceive the stimuli under environmental stresses [1]. In salinity especially with regard to sodicity, a number of detrimental effects are experienced in plants when there absorbed an excess amount of sodium (Na+). In principle, plants lose a substantial amount of potassium (K+) as a direct effect of salinity and thus osmotically plants start malfunctioning [2]. Apart from this, salinity could induce the oxidative stress by accelerating some chemical reactions to generate some highly energized oxygen species commonly referred as Reactive Oxygen Species (ROS) [3]. It is now well established that ROS are the agents for membrane damages predominantly by inducing lipid peroxidation and protein carboxylation. In this context, polyamines are involved either way by moderation of ROS and free radical generation or by activating the scavenging reactions. Thus, a significant increase of polyamines with concomitant over expression of anti-oxidation pathway has been the feature in crop plant species [4]. Therefore, it would be needless to say that polyamines are effective for stress tolerance in plant breeding. It is practically exercised with two approaches: exogenous application of polyamines for evocation of anti-oxidation pathway and secondly use of transgenic plants with up regulation of polyamine biosynthetic genes. The former is more convenient for its direct implication of salinity tolerance in many crop species [5]. With this approach, different polyamines, mostly di, tri and tetra amines have been more in practice for salinity tolerance in plants [6]. Thus, it was found that one of the di-amine, putrescine when accumulated in excess becomes toxic to plants under salinity and finally may lead to loss of tissue viability. But, higher concentration of spermidine (tri-amine) and spermine (tetra-amine) could lead significant amelioration of salt stress in sensitive rice varieties [7]. Moreover, spermidine application have been reported in withdrawing the reversible damages of salt and chilling induced oxidative stress in rice [8], cucumber [9]. In Arabidopsis sp. spermine has been evident more effective in amelioration of electrolyte leakage, recovery of plasma membrane damage, sustaining of redox balance than spermidine [10]. However, putrescine was mostly found to be related with salt induced anti-oxidative pathways in both sensitive and tolerant varieties. Oxidative damages resulting in adjustment of cellular redox by both enzymatic and non-enzymatic pathways were reported in rice varieties with the application of putrescine [10]. Alteration of enzymatic activities is supposed to be an outcome of up/down regulation of stress induced enzyme synthesis de novo. This also circumvents the possibility of protein profiling study expressed differentially in sensitive and tolerant genotypes. There are many reports on expression of antioxidative enzymes like superoxide dismutase (SOD), peroxidase (PAX), catalase (CAT), glutathione reductase (GR) etc. Those were detected both qualitatively and quantitatively under oxidative stress in rice, maize and barley [11]. On the other hand use of putrescine sometimes may lead to accumulation of some ROS like H2O2. Therefore, the relationship between the use of putrescine and its derived H2O2 remains still questionable for stress tolerance. H2O2 however undoubtedly is proven as elicitors of induced tolerance under various stresses in plants. So application of putrescine for alleviation of oxidative damages in rice varieties are less explored, particularly, through development of H2O2 accumulation. On this background the present work aims to study the effects of putrescine on changes of anti-oxidative pathways with reference to enzymatic profiles in two indica rice varieties differing in sensitivity to salinity.
Regarding the salinity tolerance of crop plants, rice exhibits a wider adaptability in varying potentials in different cultivars. The traditional tall indica varieties like Pokkali is recommended as highly tolerant but less yielder. The reverse is true for high yielding varieties like IR- 72, IR-36, IR-29, etc. [12]. Likewise, those recorded significant loss of turgidity, photosynthetic potential, high rate of lipid peroxidation and protein oxidation, inadequate membrane transport but more with synthesis of compatible solutes [13]. In the present experiment, we have explored through biochemical analysis of some anti-oxidative enzymes of SOD, GPX, CAT and GR and their differential expression. This study might be additive over existing information in salinity tolerance of land races in rice with varying potential and its modulation with polyamine application.
Materials and Methods
Raising of the plants under treatments
Two rice varieties (Oryza sativa L.) cvs. Nonabokra and Swarna were selected for the experiment to access their tolerance in the laboratory of Plant physiology and Plant Molecular Biology Research unit, Department of Botany, University of Kalyani, Kalyani, West Bengal, India. The seeds of these two varieties (both are reported as 110-120 days duration, photo insensitive and moderately nitrogen responsive) were collected from Central Soil Salinity Research Institute (CSSRI), Canning Town- 743329, West Bengal, India. Seeds were allowed to germinate after properly disinfection with 0.1% sodium hypochloride followed by repeated washing. After testing the viability as suggested [14] seeds were germinated in a seed germinator with optimum temperature and relative humidity as 27 ± 1°C and 80% respectively. The fully germinated seedlings of 15 days old were transferred to nutrient solution [15] of 5 lit capacity in a non-absorbing container for 10 days in open air to acclimatize with natural condition. During this period the average day and night period (during the month of June), average temperature, relative humidity and photoperiod (light/ dark) were 30°C, 70-80%, 14/10 hour respectively. This condition was maintained upto 15 days with frequency of nutrient solution renewal at every five days. After completion the seedlings were divided into three treatments in the same solution of 0mM NaCl (control), 200mM NaCl (18 ds/m), 200mM+2mM putrescine. The salinity level was titred for range of tolerance in rice varieties was adopted from earlier works [16]. All the sets were replicated thrice under same condition and continued upto 10 days. Plants were harvested into roots and shoots, immediately freezed in liquid nitrogen and cryo preserved in -80°C (Thermo USA). For each analysis equal amount of plant leaf tissues were taken from both control and treatment with three replications for the biochemical estimations under appropriate conditions following standard protocols.
Histochemical detection of Na in the tissues
For detection of the Na salt deposition in the tissue, the root samples were preserved in fixative solution for 4 hr and then microtome sections were done. The sections were coated with gold (5 mA, 200 Å gold coating) using IB2 ion coater, Eiko engineering. The SEM photographs/micrographs were taken using Hitachi S-530 scanning electron microscope attached with energy dispersed X-ray (EDX) unit.
Assay of anti-oxidative enzyme activity in vitro and in-gel
To study the enzymatic anti-oxidative pathway of rice plants under control and NaCl treatment, enzymes viz. guaiacol peroxidase (GPX), ascorbate peroxidase (APX), catalase (CAT), glutathione reductase (GR) and superoxide dismutase (SOD) were studied following appropriate protocol for extraction, purification and assay reaction. For each type of enzyme 1.0 g of leaf tissue was taken and crushed in liquid nitrogen followed by homogenization with extraction buffer (3.0 ml) containing 1.0 mM Tris–Cl (pH 7.7), 10 mM MgCl2, 1.0 mM DTT, 0.1 mM PMSF, 0.1 mM EDTA, 0.1 mM leupeptin, 0.1 mM BSA, 2% PVP and 0.1% protease inhibitor under cold condition. The homogenate was centrifuged at 15,000 × g, at 4°C for 15 min. The supernatant containing the crude enzyme was precipitated by 80% ammonium sulphate followed by purification in dialysis bag with suitable buffer (10 mM KCl, 0.1 mM EDTA, 1 mM PMSF, 1 mM β-ME) under 4°C for 6-8 h. The protein was concentrated under vacuum and dissolved in 0.5 ml of the dialysis buffer. The purified protein was used for assay of GPX, CAT and GR.
The activity of GPX (EC 1.11.1.7) was assayed spectrophotometrically using O-dianisidine as electron donor and hydrogen peroxide (H2O2) as substrate according to Hu et al. [17] In a reaction mixture containing (0.1 M phosphate buffer, pH 6.5; 1.5 mM O-dianisidine; 0.2 M H2O2; 0.1% protease inhibitor and equivalent amount of enzyme extract containing 50 μg of protein) The protein was measured as per Bradford et al. [18] The change of absorbance was monitored at 430 nm. The enzyme activity was determined according to Mandal et al. [19] using an extinction coefficient of O-dianisidine (26.2 mM-1 cm-1). One unit (U) of enzyme is defined, as the time required for changing the absorbance by 0.1 per unit gram of protein. For in gel studies of isozymes of GPX, the protein was run in a non-denaturing 10% polyacrylamide gel at 10 V/lane under cold condition [20]. The detection of specific band of polypeptide was resolved in an incubation mixture (50 mM Potassium phosphate buffer, 0.5 mM O-dianisidine and 0.5% H2O2).
For isolation of APX (EC 1.11.1.11), leaf tissue was homogenized in 100 mM Tris-Cl (pH 7.8) buffer containing 10 mM MgCl2, 1 mM PMSF, 100 mM EDTA, 10 mM DTT and 2% PVP, 0.1% protease inhibitor and centrifuged at 17,000 × g for 25 min at 4°C. For in vitro assay of APX, the reaction mixture containing 100 mM phosphate buffer (pH 7.5), 0.5 mM ascorbate and 0.2 mM H2O2, then equivalent amount of protein from enzyme source was added and absorbance was recorded at 290 nm [21]. For in gel staining of APX, 50 μg protein was loaded in a native PAGE which was pre run in a 2 mM ascorbate for 30 min at 4°C and gel was run for 3-4 hours at 4°C. Thereafter, the gel was equilibrated with 50 mM sodium phosphate buffer (pH 7.0) containing 2 mM ascorbate and changed in a buffer of 4 mM ascorbate and 2 mM H2O2 for 20 min. Finally, the gel was stained in a solution containing 50 mM sodium phosphate buffer and 2.45 mM NBT. The reaction was stopped by adding 10 mM TEMED in the same solution.
For assay of CAT (E.C. 1.11.1.6), 50 μg protein was incubated in a reaction mixture containing 0.5 mM potassium phosphate buffer (pH 7) and 10 mM H2O2. The activity was determined by reading the decreasing absorbance at 240 nm and activity was detected using the extinction coefficient of H2O2 as suggested by Aebi [22]. For isoenzymic studies, 50 μg of protein was loaded in a non-denaturing 10% polyacrylamide gel at 10 V/lane under cold condition. Then, incubated the gel in 0.05% H2O2 and the bands were developed in a staining solution (1% (w/v) potassium ferricyanide and 1% (w/v) ferric chloride sequentially) for 5 min and fixed with 1% hydrochloric acid.
Glutathione reductase (GR) (EC 1.8.1.7) was assayed in a reaction mixture containing 100 mM HEPESKOH pH 7.5, 0.5 mM DTT, 10 mM magnesium chloride (MgCl2),0.5 mM methionine, 0.2 mM NADH, 0.5 mM oxidized glutathione (GSSG) and 0.1% protease inhibitor according to Schaedle [23]. The mixture was incubated at 37°C for 15 min and then immediately added with 50 μg of enzyme protein. The activity was recorded from oxidation of NAD(P)H with its extinction coefficient 6.22 mM-1 cm-1. One unit of enzyme was regarded as μM NADPH oxidized min-1 μg protein-1 under assay condition. The isoforms of GR was detected by electrophoresis of protein (50 μg lane-1) in a non-denaturing polyacrylamide (10%) native gel at 4°C under 6 V/lane and 10 V/lane in stacking and resolving gel respectively. The bands were developed on thegel in an assay mixture containing 0.5 mM NAD(P)H, 3.4 mM GSSG, 0.5 mM EDTA, 25 mM Tris-Cl (pH 7.6), dichlorophenol indophenol (DCPIP), 3-(4,5-dimethyl-2 thiazolyl 2,5 diphenyl-H tetrazolium bromine (MTT), under dark condition for 10 min with repeated washing by de-ionized water.
For the extraction of SOD (EC 1.15.1.1), fresh sample was homogenized in ice cold 50 mM phosphate buffer (pH 7.0) followed by centrifugation at 15000 × g for 30 min at 4°C [24]. An aliquot of supernatant equivalent to 50 μg protein was incubated in an assay mixture containing 50 mM sodium phosphate buffer pH 7.0, 15 mM methionine, 75 μM NBT and 100 mM EDTA and kept under fluorescent light for 10 min followed by reading the absorbance at 560 nm. For in gel staining of isozymes, 100 μg of protein was loaded in a 10% native PAGE which was incubated in two successive buffers: 50 mM sodium phosphate (pH 7.5) with 2.45 mM NBT, then transferred to 50 mM sodium phosphate (pH 7.5) buffer with 26.5 mM TEMED and 26.5 μM NBT in dark for 40 min, after that the gel was exposed to fluorescent light for the development of bands.
Statistical Analysis
All the observations were recorded with three replications (n=3) and data were expressed as mean ± SE. The statistical analysis was performed by one-way (ANOVA) followed by least significance difference (LSD) test taking p ≤ 0.05 levels of significance [25]. Duncan’s multiple range test was performed as post hoc on parameters subjected to ANOVA (only if the ANOVA was significant). All the statistical tests were performed using SPSS software (SPSS Inc., version 16.0). Windows Microsoft Excel 2003 software was employed for computation, data analysis and graphics.
Results and Discussion
Accumulation of Na+ in the tissues
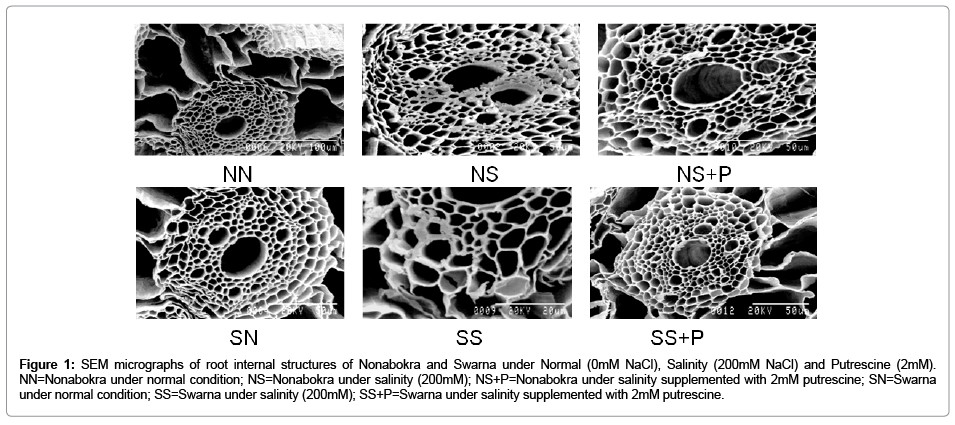
Both the varieties viz., Nonabokra and Swarna recorded a variable accumulation of Na+ and its resultant structure of their roots, when grown under 200mM NaCl (Figure 1). A critical analysis of the accumulation pattern as revealed by SEM-EDXS study depicted some interesting features. Plants recorded a significant variation in tissue Na+ content as detected by atomic absorption spectrophotometric study. This has more been confirmed with SEM-EDXS technique of the root samples where specific tissues showedsignificant variations in their conformity due to Na+ induced damages, if any. Irrespective of varieties, Nonabokra and Swarna accumulated Na+ significantly more in 200 mM NaCl and those were 1.7 fold and 2.6 fold respectively over the control. However, plants under 2mM putrescine supplementation had minimized the accumulation of Na+ by 31.8 per cent in Nonabokra and 21.1 per cent in Swarna. Therefore, it undoubtedly advocates the alleviating role of Na+ toxicity by putrescine application. Now from the SEM study, it is revealed that salinity affected tissues were cortical cells and vascular lumens where the Na+ vulnerability was pronounced (Figure 1). Due to accumulation of Na+, the cortical cells are somewhat diffused in nature and lumens are constricted under salinity as compared to control. This effect is shown more in Swarna than Nonabokra under same condition. Interestingly, SEM-EDXS study also revealed that putrescine could sustain the tissue integrity close to control. The cortical cells and lumens appeared more normal in shape and configuration w with putrescine treatment.
Figure 1: SEM micrographs of root internal structures of Nonabokra and Swarna under Normal (0mM NaCl), Salinity (200mM NaCl) and Putrescine (2mM). NN=Nonabokra under normal condition; NS=Nonabokra under salinity (200mM); NS+P=Nonabokra under salinity supplemented with 2mM putrescine; SN=Swarna under normal condition; SS=Swarna under salinity (200mM); SS+P=Swarna under salinity supplemented with 2mM putrescine.
Effects of salinity on GPX activity and its expression profile of polymorphism
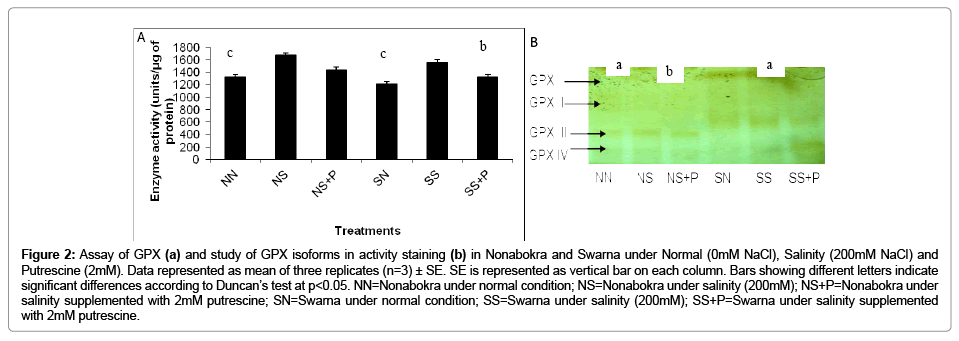
In the present experiment, plants were significantly sensitized for GPX activity in vitro with 200 mM NaCl solution supplemented with 2mM putrescine. Irrespective of the varieties, there recorded 1.27 fold increases in activity over control. Still, GPX activity was induced maximum in salt treated Nonabokra (1.23 fold) than in Swarna (1.14 fold) as compared both of those under control (Figure 2a). Relative to NaCl treatment, plants also showed variations when supplemented with 2mM putrescine in both the varieties. However, regardless of the varieties, there was significant (p ≤ 0.05) down regulation by 14.45 per cent under putrescine treatment compared to 200 mM salinity. The effects of putrescine were similar in Nonabokra also and were more responded to putrescine than Swarna. Since, there recorded 22.82 per cent and 16.56 per cent reduction in GPX activity respectively (Figure 2a). The expression profile of GPX when run in non denaturing gel followed by activity staining with orthodianisidine reveals four possible polymorphic groups (GPX I to GPX IV) differing in the intensities for both the varieties (i.e., Nonabokra and Swarna) (Figure 2b). Interesting to note that in control, expression of the four polypeptides (GPX I to GPX IV) are constitutively expressed but varying in intensities GPX III (being more in Nonabokra than Swarna). But, under 200 mM NaCl, the bands for GPX III and IV are more expressed in Swarna than Nonabokra. Similarly when putrescine was applied to both the varieties there also recorded differential sensitivity for expression of GPX, more with GPX IV. On observation, the later band appeared over-expressed by several folds under putrescine application with 200 mM NaCl for Swarna. On the contrary, no such over expression was observed regardless of GPX groups with putrescine in case of Nonabokra.
Figure 2: Assay of GPX (a) and study of GPX isoforms in activity staining (b) in Nonabokra and Swarna under Normal (0mM NaCl), Salinity (200mM NaCl) and Putrescine (2mM). Data represented as mean of three replicates (n=3) ± SE. SE is represented as vertical bar on each column. Bars showing different letters indicate significant differences according to Duncan’s test at p<0.05. NN=Nonabokra under normal condition; NS=Nonabokra under salinity (200mM); NS+P=Nonabokra under salinity supplemented with 2mM putrescine; SN=Swarna under normal condition; SS=Swarna under salinity (200mM); SS+P=Swarna under salinity supplemented with 2mM putrescine.
Effects of salinity on APX activity and its expression profile of polymorphism
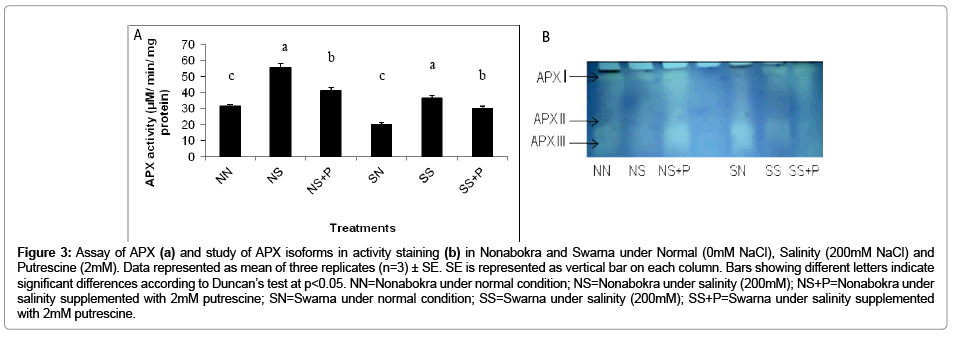
Figure 3b shows the in vitro activity of APX in rice (cv. Nonabokra and Swarna) under NaCl and NaCl supplemented with putrescine. The activity of APX recorded almost similar trend as that of GPX being significantly increased under salinity and thereafter declined with putrescine application irrespective of varieties. When the plants were exposed to salinity irrespective of varieties, the activity was increased by 1.75 fold as compared to control. Swarna recorded higher activity (1.80 fold) than Nonabokra (1.7 fold) to be induced by salinity as compared to control (Figure 3a). Still, on closer observation, the variation for induction of activity compared to control was more significant (p ≤ 0.05) in case of Swarna than Nonabokra. Therefore, Swarna appeared more responsive to salinity stress than the other variety, Nonabokra. Putrescine on the other hand, had also been similarly effective in modulating the activity of APX. Thus, the down regulation of activity was 21.44 per cent with putrescine over stress irrespective of varieties. The activity was minimized by 25.4 per cent for Nonabokra and 17.48 Per cent for Swarna (Figure 3a). The expression pattern for APX isozyme with activity staining in gel revealed three distinct peroxidase groups (APX I to APX III) (Figure 3a). Amongst those, two activity bands viz. APX I and APX II were most noticeable irrespective of varieties under the treatments. Still, for Nonabokra, APX II was significantly increased in intensity of band under NaCl treatment than the same in case of Swarna. But putrescine could hardly be able to induce any changes of band of its intensities for APX in Swarna. On the contrary, Nonabokra though recorded a significant change in activity of APX with putrescine, the band intensities for any of APX isozyme (APX I and APX II) were consistent under same condition. An additional band, APX III, showed its expression only under control which was not distinctly differentiated under salinity and even with putrescine treatment. Almost a similar expression profile for APX isoenzymes was observed in variety Swarna also.
Figure 3: Assay of APX (a) and study of APX isoforms in activity staining (b) in Nonabokra and Swarna under Normal (0mM NaCl), Salinity (200mM NaCl) and Putrescine (2mM). Data represented as mean of three replicates (n=3) ± SE. SE is represented as vertical bar on each column. Bars showing different letters indicate significant differences according to Duncan’s test at p<0.05. NN=Nonabokra under normal condition; NS=Nonabokra under salinity (200mM); NS+P=Nonabokra under salinity supplemented with 2mM putrescine; SN=Swarna under normal condition; SS=Swarna under salinity (200mM); SS+P=Swarna under salinity supplemented with 2mM putrescine.
Effects of salinity on CAT activity and its expression profile of polymorphisms
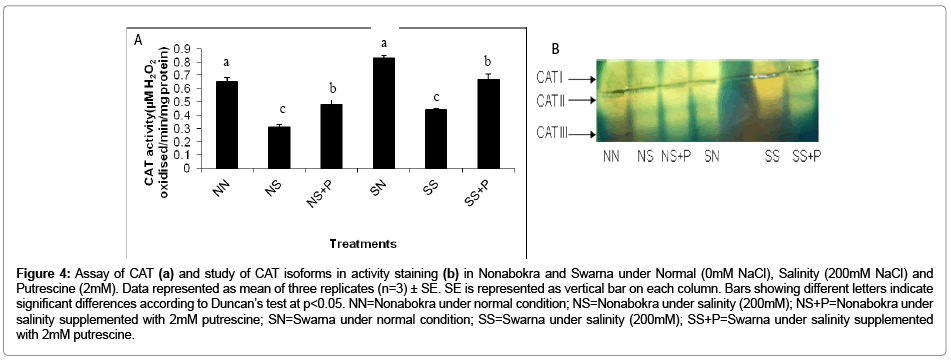
In the present experiment, this enzyme happens to be operating in some different manner under salinity. It is interesting to note that the activity was significantly (p ≤ 0.05) down regulated as a function of NaCl. The activity was reduced by 49.6 percent, irrespective of varieties being exposed to stress as compared to control condition (Figure 4b). The tolerant cv. Nonabokra had recorded the maximum subdued activity (52.30 percent) whereas for susceptible cv. Swarna, it was comparatively less (46.9 percent) in comparison to control. Putrescine had been efficient in retrieving the CAT activity by 1.5 fold irrespective of the varieties. The gain in activity by putrescine application recorded 1.5 fold in Nonabokra. For Swarna, the response with putrescine is same as that of Nonabokra 1.5 fold. On non-denatured native gel, the expression of CAT activity by staining with ferricyanide and ferric chloride appeared as three distinct negative bands. These were varied strikingly in their intensities according to varietal differences through induction of salinity. For the cvs (Swarna and Nonabokra), the CAT profile recorded three markedly distinct bands CAT I, CAT II and CAT III under salinity (Figure 4a). Moreover, salinity had differentiated the expression of CAT I and CAT II in their intensities in those rice varieties, however, higher in Nonabokra than Swarna. Still, for CAT III, there recorded no such changes in intensitiy irrespective of varieties. Therefore, for the sensitivity of CAT, only two activity specific bands were restored (CAT I and CAT II) under salinity. On the other hand, CAT expression was not greatly changed by application of putrescine in any of the varieties compared to salinity.
Figure 4: Assay of CAT (a) and study of CAT isoforms in activity staining (b) in Nonabokra and Swarna under Normal (0mM NaCl), Salinity (200mM NaCl) and Putrescine (2mM). Data represented as mean of three replicates (n=3) ± SE. SE is represented as vertical bar on each column. Bars showing different letters indicate significant differences according to Duncan’s test at p<0.05. NN=Nonabokra under normal condition; NS=Nonabokra under salinity (200mM); NS+P=Nonabokra under salinity supplemented with 2mM putrescine; SN=Swarna under normal condition; SS=Swarna under salinity (200mM); SS+P=Swarna under salinity supplemented with 2mM putrescine.
Effects of salinity on GR activity and its expression profile of polymorphism.
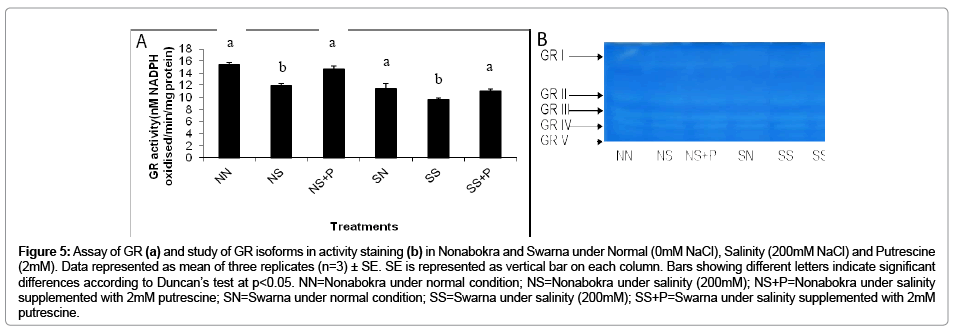
In the present experiment the rice varieties were evaluated with the activity of GR in vitro as well as activity staining. The activity of GR followed a similar trend like CAT as recorded in the present experiment. There recorded a significant (p ≤ 0.05) down regulation in activities irrespective of varieties by 19.69 per cent under salinity (Figure 5b). It also recorded a marked variation in the sensitivity of the varieties viz. Swarna and Nonabokra with salinity and those were 22.82 per cent and 16.56 per cent respectively. When plants were supplemented with 2mM putrescine, it showed retrieval of activity irrespective of varieties by 1.18 fold over salinity. As expected Nonabokra being tolerant showed its efficiency by increasing GR activity by 1.23 fold and Swarna by 1.14 fold under the application of 2mM putrescine respectively (Figure 5b). As already recorded GR is coded by multigenic gene family and therefore it might be expressed variably with different isozymic polypeptides. In the present experiment, five distinct isozymic bands of GR were resolved (GR I to GR V) when protein was run on non denaturing gel followed by activity staining (Figure 5a). It recorded that two major bands (GR I and GR II) were prominent in control condition irrespective of varieties. For both of those the bands of GR III to GR V were over expressed under salinity. Moreover, the bands IV and V in Nonabokra showed more intensities than Swarna under salinity. On the other hand, bands II and III were more expressed when supplemented with 2 mM putrescine regardless of varieties.
Figure 5: Assay of GR (a) and study of GR isoforms in activity staining (b) in Nonabokra and Swarna under Normal (0mM NaCl), Salinity (200mM NaCl) and Putrescine (2mM). Data represented as mean of three replicates (n=3) ± SE. SE is represented as vertical bar on each column. Bars showing different letters indicate significant differences according to Duncan’s test at p<0.05. NN=Nonabokra under normal condition; NS=Nonabokra under salinity (200mM); NS+P=Nonabokra under salinity supplemented with 2mM putrescine; SN=Swarna under normal condition; SS=Swarna under salinity (200mM); SS+P=Swarna under salinity supplemented with 2mM putrescine.
Effects of salinity on SOD activity and its expression profile of polymorphism
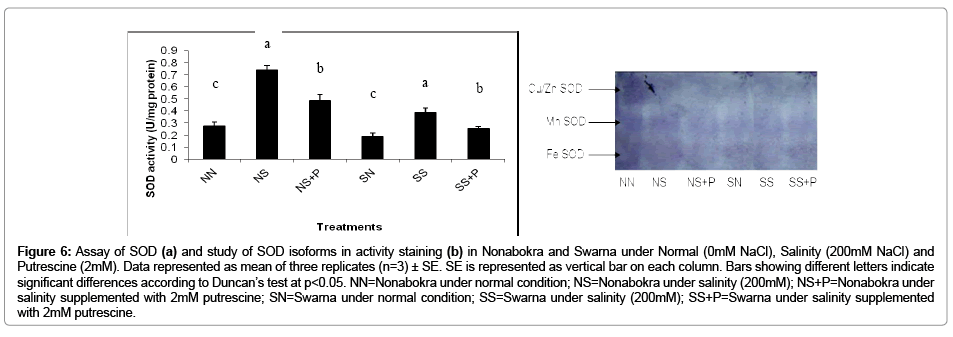
In the present experiment, the plants were evaluated with respect to NBT reduction in vitro from leaf protein extract under intense illumination. The activity of the SOD recorded a significant (p ≤ 0.05) increase irrespective of the varieties when the plants were exposed to salinity and it was 2.34 fold over control (Figure 6b). As expected, there were notable differences in SOD activity between the tolerant cv. Nonabokra and the susceptible cv. Swarna. On comparative basis, the activities were 2.65 fold and 2.04 fold in Nonabokra and Swarna respectively under that condition. Putrescine had almost played a similar role in modulating the enzyme activity like GPX and APX. There recorded a decrease of enzyme activity by 34.33 per cent as compared to salinity irrespective of varieties. Nonabokra (34.3 per cent) and Swarna (34.37 per cent) may indicate insensitivity to putrescine for salinity response. Now, the activity staining in gel for the expression of SOD showed some distinct variations in appearance of polypeptide bands in Nonabokra and Swarna (Figure 6a). SOD activity revealed four distinct polypeptide bands in control and treatments. According to the sensitivity of SOD bands with either KCN and H2O2 or KCN only there recorded viz., Mn-SOD, Fe-SOD and Cu/Zn-SOD. The intensities of the Mn-SOD and Cu/Zn-SOD were most vividly expressed, whereas for Fe-SOD, it was less in case of Nonabokra under normal condition. Almost similar pattern of expression was found under salinity for the same variety. But, for Swarna (the sensitive variety), there recorded distinct expression and variation of three band intensities under control and salinity. More so, as compared to control, Cu/Zn-SOD was less expressed. But for Mn-SOD, the reverse was observed. Interestingly, the application of putrescine had significantly modulated for different forms of SOD in both the varieties, however, differentially. The Mn- SOD and Cu/Zn-SOD which were expressed in less were retrieved with application of putrescine by several folds in cv. Nonabokra. Still, for the Fe-SOD, there recorded no such remarkable development in expression with putrescine. Therefore, Nonabokra seemed to be furnishing its tolerance to salinity by mitochondrial (Mn-SOD) and cytosolic (Cu/Zn-SOD) proteins up-regulation. On the contrary, for Swarna, there observed down regulation of SOD isoforms with putrescine application. This imparts the insensitivity of putrescine towards SOD in non tolerant cv. Swarna for conferring the susceptibility of oxidative stress. The effect of putrescine on diminishing the expression for SOD is more pronounced in Cu/Zn-SOD followed by Mn-SOD as revealed from the (Figure 6a). Thus, the differential sensitivity for putrescine is indicative for discriminating the tolerance under oxidative stress adhered to salinity in these two varieties.
Figure 6: Assay of SOD (a) and study of SOD isoforms in activity staining (b) in Nonabokra and Swarna under Normal (0mM NaCl), Salinity (200mM NaCl) and Putrescine (2mM). Data represented as mean of three replicates (n=3) ± SE. SE is represented as vertical bar on each column. Bars showing different letters indicate significant differences according to Duncan’s test at p<0.05. NN=Nonabokra under normal condition; NS=Nonabokra under salinity (200mM); NS+P=Nonabokra under salinity supplemented with 2mM putrescine; SN=Swarna under normal condition; SS=Swarna under salinity (200mM); SS+P=Swarna under salinity supplemented with 2mM putrescine.
Conclusion
The oxidative stress out of various abiotic fluctuations happens to be the most crucial factor in maintenance of normal cellular activities in plants. Salinity is the most predominant form in establishing the oxidative stress in plants. The stress is dependent on metal concentration, duration of exposure, growth stages and finally plants’ genetic back up to react it. The responses in plants to Na+ mediated oxidative stress are manifested into gene expression through up or down regulation of the cellular functionales. In general, the spillage of electrons from electron transport chain (ETC) in some organelle and its reduction to molecular oxygen generates the ROS. There is a synchronization procedure involving generation and lysis of those ROS and thereby homeostasis of cellular redox as maintained [26]. However, plants are prone to oxidative stress when the balance for generation and lysis are perturbed under various abiotic stresses [27]. In the present investigation, salinity had significantly induced the oxidative stress by generation of ROS that directly impacts the tissue lysis irrespective of varieties. The application of polyamines for down-regulation of ROS generation and its concomitant lysis of tissues were also reported in rice under drought and other abiotic stresses [28]. In most of the cases, it is claimed that polyamine might shield the negatively charged domain of cellular membrane with its polycationic residues. The electrostatic interactions between these two oppositely charged domains are supposed to be the most possible way to facilitate the role of polyamine to oxidative damages [29]. Therefore, polyamines are evident to be a reliever molecule by the avoidance of lipid peroxidation reactions in plants under various abiotic stresses [19]. It was also reported earlier that polyamines could also act as antioxidant directly or inducing the anti-oxidation pathways in plants. Putrescine unlike other profiles of polyamines (viz spermine and spermidine) have not been explored for significant correlation with anti-oxidative responses. As for example in some tolerant rice varieties like Pokkali recorded more accumulation of putrescine in roots than shoots, while the other sensitive varieties recorded a reverse trend [30]. In other studies the putrescine was reported as lower or without any significant alleviating roles in other plant species [31] Spermine in general has been serving as well protective moieties and renders its effect irrespective of plants and stressors. In comparison spermidine was reported as an intermediate efficient in stress recovery in many plants [7]. Therefore, no clear and consistent relationship was recorded between any one of the fraction of the polyamines (spermidine, spermine or putrescine) content with salinity resistance. However, on an average basis polyamines could modulate total anti-oxidation activity under any sort of abiotic stress. For the past several years we have been investigating to elucidate the physiological, biochemical and cellular aspects of the major polyamine in crop plants, rice being the most important. Therefore, in the present experiment putrescine was considered to be a stress alleviating agent under Na+ induced oxidative stress for the two rice varieties. With the SEM study, a clear picture of tissue disintegration in the plants was found with variations in Nonabokra and Swarna. The former variety shows tolerance and recorded lesser affected tissues than the later. However, putrescine, irrespective of varieties had shown the reduced effects of tissue disintegration. It was earlier reported that polyamines might act by alleviating the antioxidant pool or by up/down regulating enzymatic anti-oxidation pathway [5,32]. Likewise the anti-oxidative enzymes like GPX, APX, SOD, CAT, and GR were modulated according to the concentrations of Na+ and its concomitant effects by application of putrescine. The results of the present investigation show that irrespective of varieties, rice plants recorded an up-regulation of enzyme activities except for CAT and GR under stress. More so, putrescine had modulated the activities in each case. These enzymatic anti-oxidation might be limiting the tolerance to salinity as recorded from sensitive cv. Swarna under that condition. Thus, SOD activity with its increased trend might be attributed for deactivation of O2- regardless of varieties, however, more in Nonabokra than Swarna. SOD is believed to be the first line of defense by dis-mutation of O2- with one electron transfer and its conversion to H2O2. SOD constitutes the group of metalloenzymes those are displayed with several isozymic forms as reported in higher plants [33]. The activity of SOD with its up regulation is also evident from its isozymic profiles as found in the present experiment. SOD is furnished with three distinct bands demarcated as SOD I, SOD II and SOD III. Classically, these three bands are identified with their inhibitor sensitivity to different metals as CU-SOD, Zn/Mn-SOD and Fe-SOD. According to the expression potential in different cellular compartments these are also regarded as chloroplastic, mitochondrial and cytosolic counterparts of total pool of SOD [34]. Interestingly, SOD II and SOD III are distinctly variable in both the varieties under salinity as compared to control. The pattern of in vitro activities for SOD and isozymic bands for trinuclear (Cu/Zn/ Mn-SOD) forms have been evident as species dependant, tissue and growth stage specific and even stressortypes. It seems, at least apparent that varied number of SOD isoforms should render the tolerance differtially to rice varieties subjected to heavy metal and heat stress [35]. The increase in SOD activity is likely to be related with an altered gene expression for Cu/Zn SOD (for cytosolic and chloroplast) and possibly is over expressed under salt in excess. In rice also, a Cu-SOD was found to be up regulated under heat stress and displayed variable forms in photosynthetic tissues (leaves/shoots) and non-photosynthetic tissues (like roots) [36]. Admittedly, in the present experiment, the lesser activities of SOD in Swarna could limit the scavenging ability for ROS and thereby becomes more prone to oxidative damages. Moreover, chances could not be overlooked for an impaired synthesis of transcripts or/and accumulation of less active enzymes in susceptible varieties as compared to tolerant ones [3].
Peroxidases, the enzymes that scavenge the H2O2 using a variety of phenolic moieties as electron donors are distributed in different cellular compartments. Likewise, APX, a form of peroxidase can lyse the H2O2 with ascorbate as reductant. APX is typically characterized with higher affinity for H2O2 than other peroxidases like guaiacol peroxidase, glutathione peroxidase or even catalase (CAT) [37]. In the present experiment there is a clear cut indication of ascorbateglutathione pathway through an up regulation of APX activity as observed irrespective of rice varieties. However, cv. Swarna recorded higher activity than cv. Nonabokra. Putrescine had relieved the stress for both of those. The electrophoretic separation of APX isoforms (APX I, APX II and APX III) are distinctively variable in Nonabokra and Swarna. Interesting to note that APX I and APX III are more expressed in Nonabokra under salinity and could have major contribution to the total activities also reported in other crops [38]. Thus, over expression in different APX gene has successfully enhanced the activity of the enzyme in response to drought, salinity, metal toxicity and UV irradiation [39]. GPX is another class of peroxidase that essentially requires guaiacol as reductant for lysis of H2O2. GPX is independent to ascorbateglutathione pathway. Still, it is associated with many biosynthetic pathways (lignifications of phenolics, IAA catabolism, wound healing, ethylene biosynthesis and resistance against many abiotic and biotic stresses). GPX imparts its activity as effective quencher of ROS, peroxy radicals as deduced under different conditions of stresses. In reference to oxidative stress under metal toxicity GPX is suggested as potential biomarker for sub lethal metal toxicity in plants. GPX has also been reported in many isoforms with varying kinetic properties [40]. In the present experiment, four distinct isoforms (GPX I, GPX II, GPX III and GPX IV) were recorded when separated through electrophoretic field in native gel. Amongst those, GPX III and GPX IV appeared with variable intensities induced by NaCl and with putrescine also as found in the present experiment. Peroxidase with their high number of isoforms and heterogeneous regulation are involved for both metabolic and catabolic activities for plant’s development. GPX, in fact belongs to class III peroxidases and are mostly soluble, cytosolic and appoplastic or cell wall bound in nature (Mika and Luthje). The various isoformic structure of GPX is significant in plants, particularly, under stressful conditions. With two possible roles, (i.e., peroxidative and hydroxylic), however, mutually exclusive in nature, peroxidase could detoxify and/ or generate ROS (OH-, HOO- etc.). The later role is also frequently used in polymerization of cell wall residues with regulation of cellular H2O2 [41]. Therefore, the distinct isoforms of GPX as also elucidated in the present investigation might be employed for any of those activities as necessitates under oxidative stress. On the contrary, CAT recorded a down regulated trend under induction of salinity in both the varieties in the present investigation. Moreover, application of putrescine could be able to retrieve the activity, though not significant. The diminishing trend for CAT activity in context to other enzymatic anti-oxidationis exclusively species specific and depends on kinds of stressors [31]. The down regulation of CAT by salinity or other abiotic stresses inducing oxidative damages could be assumed due to impairment of new enzyme synthesis at the transcriptional level. CAT is characterized by a very rapid turnover rate. However, with its higher affinity towards H2O2 it is regarded as most efficient quencher than APX and other peroxidases. In the present experiment CAT in native gel was separated into three distinct isoforms: CAT I, CAT II and CAT III. All angiospermic crop species where CAT is expressed as antioxidizing enzyme under stress condition are represented in both photosynthetic and nonphotosynthetic tissues. In most of the cases CAT I is photosynthetically light regulated and required to lyse the H2O2 generated in sub cellular fractions [42]. Moreover, crops like C3 categories (rice being most common) can exercise photo-respiratory carbon metabolism and generation of H2O2 that happens to be a surplus in the tissues. This over-accumulation of H2O2 could be effective for denaturing and deactivation of CAT as an enzymatic protein [42]. A significant decline in activities of CAT was similarly recorded in drought [43], water stress [3], heavy metal stress [12], under toxic chemicals [44]. Therefore, in plants, catalase alike other anti-oxidative enzymes play an important role to both environmental as well as physiological oxidative stress with its varying expression potentiality in different tissues.
In enzymatic and non-enzymatic pathways glutathione (GSH) is alternated in two forms: oxidized (GSSG) and its reduced (GSH) forms. Glutathione reductase (GR), a flavoenzyme can retrieve the GSH by NAD(P)H- dependant reduction. Thus, it maintains a cellular homeostasis for GSH to GSSG ratio. This enzyme was studied in the present investigation with a declining trend irrespective of varieties under salinity and thereafter, was recovered with putrescine application. The activity of GR remains diminished with other heavy metals like Hg, Pb, Cu etc. and thereafter, steadily decreased [45]. GR predominantly located in cytosol, however, few variants are also found in sub-cellular compartments. In addition the major GR activity in photosynthetic tissues as expressed in chloroplastic isoforms to replenish the reduced glutathione (GSH) [46]. In that organelle GSH and GR are also involved in detoxification of H2O2 generated by many biosynthetic byproducts. So, GR exists in variable polypeptide forms as revealed from its separation of isozymic bands as resolved in non denaturing gel in the present experiment (GR I to GR V). Those forms are specifically required for complexion with ROS with reduced glutathione and thus detoxification is ensured. On the other hand, activities of GR are also facilitated for more acquisition of GSH through ascorbate glutathione cycles [47]. In turn, GSH could replenish the conversion of dehydroascorbate into ascorbate. The later is the prime non- antioxidant moiety in plant system effective under oxidative exposure out of abiotic and biotic stresses [19]. In the present case the varietal differences in GR isoforms might be involved in a discriminatory manner according to the sensitivity to salinity of those varieties. The specific expression of the polypeptide bands and their varying intensities are indicative of the fact of the induced tolerance through GSH-GSSG mediated anti-oxidation. In this experiment we also found over expression of GR in plants leading to higher foliar glutathione and ascorbate content and thereby imparting an improved tolerance in such cases [16]. The sensitivity of the band intensity to putrescine is also more implicative for modulation of glutathione to maintain adequate redox in plants under oxidative stress. It also highlights the behavior of plants to impart their sensitivity in sustenance of lesser oxidative damages by polyamines [48].
Based on the facts and figures in the present experiment, it could be inferred that rice varieties are preliminarily adapted to salinity by exercising their anti-oxidation pathways. In addition, polyamine can be considered as a successful elicitor to improve or moderate the plant’s responses being induced with ROS under NaCl toxicity. Thus, Nonabokra undoubtedly proved its potential to tolerate salinity induced oxidative stress with its up regulation of enzymatic anti-oxidationcascade On the contrary Swarna was evident as more sensitive under same condition and thus showed less improved strategies. Regardless of the varieties, plants recorded discriminatory responses with putrescine those were verified with constitutive and inducible expression of anti-oxidative enzymes and also their isozymic forms. As a whole, the anti-oxidation system is facilitated co-operatively by entire enzyme cascade and by some non-enzymatic antioxidant moieties. Salinity stress in the form of NaCl in vitro system could be able to promote differential expression of those enzymes where SOD, GPX, APX, CAT and GR become integral component and were also modulated with polyamine (putrescine as in the present experiment). The expression profiles of those enzymes are clear indicative of the fact for not only varietal discrimination but also Na+ toxicity and its probable modulation with polyamines. On the other hand, the present work is significant, since it is employed with 200mM NaCl which corresponds to 18 ds/m that mimic the average range of sodicity in the saline belt soil. Admitted well that short term effects of salinity, particularly, under controlled laboratory conditions does not necessarily reflects the behavior of plants as in field. More insights in cellular responses like metallothionine, phytochelatin and other differential gene expressions are required for decisive attributes in tolerance. Still, the comparative studies of physiological responses as investigated in the present case might help in unraveling the basic mechanisms of salinity tolerance in rice. With expectation, these could be employed for selection programme of rice varieties under the field conditions of salinity or other factors inducing oxidative stress.
Acknowledgements
The fellowship for 1st author was provided by University Grants Commission, New Delhi, India through UGC-RFSMS scheme and DST-PURSE Programme, DST, Govt. of INDIA in support to University of Kalyani, Kalyani. Financial support to conduct the experiment was also provided through DST-PURSE programme, Department of Science and Technology, Govt. of India.
References
- Alcazar R, Marco F, Cuevas JC, Patron M, Ferrando A, et al. (2006) Involvement of polyamines in plant response to abiotic stress. Biotechnol Lett 28: 1867-1876.
- Zhu JK (2002) Salt and drought stress signal transduction in plants.Annu Rev Plant Biol 53: 247-273.
- Basu S, Roychoudhury A, Saha PP, Sengupta DN (2010) Differential anti-oxidative responses of indica rice cultivars to drought stress. Plant Growth Regul 60: 51-59.
- Tang W, Newton RJ (2005) Polyamines reduce salt-induced oxidative damage by increasing the activities of anti-oxidative enzymes and decreasing lipid peroxidation in Virginia pine. Plant Growth Regul 46: 31-43.
- Sanchez DH, Cuevas JC, Chiesa MA, Ruiz OA (2005) Free spermidine and spermine content in Lotus glaber under long-term salt stress. Plant Sci 168: 541-546.
- Roy P, Niyogi K, Sengupta DN, Ghosh B (2005) Spermidine treatment to rice seedlings recovers salinity stress induced damage of plasma membrane and PM-bound H+-ATPase in salt-tolerant and salt-sensitive rice cultivars. Plant Sci 168: 583-591.
- Ndayiragije A, Lutts S (2006) Do exogenous polyamines have an impact on the response of a salt-sensitive rice cultivar to NaCl?J Plant Physiol 163: 506-516.
- Shen W, Nada K, Tachibana S (2000) Involvement of polyamines in the chilling tolerance of cucumber cultivars.Plant Physiol 124: 431-439.
- Kubis J (2008) Exogenous spermidine differentially alters activities of some scavenging system enzymes, H2O2 and superoxide radical levels in water-stressed cucumber leaves. J Plant Physiol 165: 397-406.
- Yamaguchi K, Takahashi Y, Berberich T, Imai A, Miyazaki A, et al. (2006) The polyamine spermine protects against high salt stress in Arabidopsis thaliana.FEBS Lett 580: 6783-6788.
- Liu JH, Inoue H, Moriguchi T (2008) Salt-stress mediated changes in free polyamine titers and expression of genes responsible for polyamine biosynthesis of apple in vitro shoots. Environ Exp Bot 62: 28-35.
- Roychoudhury A, Basu S, Sarkar SN, Sengupta DN (2008) Comparative physiological and molecular responses of a common aromatic indica rice cultivar to high salinity with non-aromatic indica rice cultivars.Plant Cell Rep 27: 1395-1410.
- Ghosh N, Adak MK, Ghosh PD, Gupta S, Sengupta DN, et al. (2011) Differential responses of two rice varieties to salt stress. Plant Biotechnol Rep 5: 89-103.
- Hong TD, Ellis RH (2004) the survival and germinating orthodox seeds after dessication and hermatic storage. J Exp Bot 43: 239-247.
- Hoagland DR, Arnon DI (1950) The water culture method for growing plants without soil. California Agricultural Experiment Station Circular 347: 1-32.
- Lee SC, Kwon SY, Kim SR (2009) Ectopic expression of a cold-responsive CuZn superoxide dismutase gene, SodCc1, in transgenic rice (Oryza sativa L.). J Plant Biol 52: 154-160.
- Hu Y, Ge Y, Zang C, Zu T, Cheng W (2009) Cd toxicity and translocation in rice seedlings are reduced by hydrogen peroxide treatments. Plant Growth Regul 5: 51-61.
- Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding.Anal Biochem 72: 248-254.
- Mandal C, Ghosh N, Maiti S, Das K, Gupta S, et al. (2013) Antioxidative responses of Salvinia (Salvinia natans Linn.) to aluminium stress and it's modulation by polyamine.Physiol Mol Biol Plants 19: 91-103.
- Ammar WB, Nouairi I, Zarrouk M, Ghorbel HM, Jemal F (2008) Anti-oxidative response to cadmium in roots and leaves of tomato plants. Biol Plantarum 52: 727-731.
- Davletova S, Rizhsky L, Liang H, Shengqiang Z, Oliver DJ, et al. (2005) Cytosolic ascorbate peroxidase 1 is a central component of the reactive oxygen gene network of Arabidopsis.Plant Cell 17: 268-281.
- Schaedle M (1977) Chloroplast glutathione reductase.Plant Physiol 59: 1011-1012.
- Nakano Y, Asada K (1981) Hydrogen peroxide is scavenged by ascorbate specific peroxidase in spinach chloroplasts. Plant Cell Physiol 22: 867-888.
- Gomez KA, Gomez AA (1984) Statistical procedures for agricultural research. John Wiley, New York, USA.
- Farooq M, Wahid A, Kobayashi N, Fujita D, Basra SMA (2009) Plant drought stress: effects, mechanisms and management. Agron Sustain Dev 29: 185-212.
- Sharma P, Jha AB, Dubey RS, Pessarakli Md (2012) Reactive oxygen species, oxidative damage and anti-oxidative defense mechanism in plants under stressful conditions. J Bot.
- Peremarti A, Bassie L, Christou P, Capell T (2009) Spermidine facilitates recovery from drought but does not confer drought tolerance in transgenic rice plants expressing Datura stramonium S-adenosylmethionine decarboxylase. Plant Mol Biol 70: 253-264.
- Piterková J, Luhová L, Zajoncová L, Šebela M, Petrivalský M (2012) Modulation of polyamine catabolism in pea seedlings by calcium during salinity stress. Plant Prot Sci 48: 53-64.
- Liu JH, Kitashiba H, Wang J, Ban Y, Moriguchi T (2007) Polyamines and their ability to provide environmental stress tolerance to plants. Plant Biotechnol 24: 117-126.
- Mansour MMF, Al-Mutawa MM (1999) Stabilization of plasma membrane by polyamines against salt stress. Cytobios 100: 7-17.
- Roychoudhury A, Basu S, Sengupta DN (2012) Antioxidants and stress-related metabolites in the seedlings of two indica rice varieties exposed to cadmium chloride toxicity. Acta Physiol Plant 34: 835-847.
- Tewari RK, Kumar P, Sharma PN (2006) Antioxidant responses to enhanced generation of superoxide anion radical and hydrogen peroxide in the copper-stressed mulberry plants.Planta 223: 1145-1153.
- Nahakpam S, Shah K (2011) Expression of key antioxidant enzymes under combined effect of heat and cadmium toxicity in growing rice seedlings. Plant Growth Regul 63: 23-35.
- Shah K, Nahakpam S (2012) Heat exposure alters the expression of SOD, POD, APX and CAT isozymes and mitigates low cadmium toxicity in seedlings of sensitive and tolerant rice cultivars.Plant Physiol Biochem 57: 106-113.
- Zhang M, Liu X, Yuan L, Wu K, Duan J, et al. (2012) Transcriptional profiling in cadmium-treated rice seedlings roots using suppressive subtractive hybridization. Plant Physiol Bioch 50: 79-86.
- Wang J, Zhang H, Allen RD (1999) Overexpression of an Arabidopsis peroxisomal ascorbate peroxidase gene in tobacco increases protection against oxidative stress.Plant Cell Physiol 40: 725-732.
- Zhao Z, Chen G, Zhang C (2001) Interaction between reactive oxygen species and nitric oxide in drought-induced abscissic acid synthesis in root tips of wheat seedlings. Aust J Plant Physiol 28: 1055-1061.
- Xiong L, Schumaker KS, Zhu JK (2002) Cell signaling during cold, drought, and salt stress.Plant Cell 14 Suppl: S165-183.
- Radotic K, Ducic T, Mutavdzic D (2000) Changes in peroxidase activity and isoenzymes in spruce needles after exposure to different concentrations of cadmium.Environ Exp Bot 44: 105-113.
- Minibayeva F, Kolesnikov O, Chasov A, Beckett RP, Lüthje S, et al. (2009) Wound-induced apoplastic peroxidase activities: their roles in the production and detoxification of reactive oxygen species.Plant Cell Environ 32: 497-508.
- Zhao H, Yang Z (2008) exogenous polyamines alleviate the lipid peroxidation induced by cadmium chloride stress in Malus hupehenis Rehd. Sci Hort 116: 442-447.
- Rout GR, Samantaray S, Das P (2001) Aluminium toxicity in plants: a review. Agronomie 21: 3-21.
- Morsy MR, Jouve L, Hausman JF, Hoffmann L, Stewart JM (2007) Alteration of oxidative and carbohydrate metabolism under abiotic stress in two rice (Oryza sativa L.) genotypes contrasting in chilling tolerance.J Plant Physiol 164: 157-167.
- Foyer CH, Noctor G (2005) Oxidant and antioxidant signalling in plants: a re-evaluation of the concept of oxidative stress in a physiological context. Plant Cell Environ 28: 1056-1071.
- Yoshida S, Tamaoki M, Shikano T, Nakajima N, Ogawa D, et al. (2006) Cytosolic dehydroascorbate reductase is important for ozone tolerance in Arabidopsis thaliana.Plant Cell Physiol 47: 304-308.
- Tseng MJ, Liu CW, Yiu JC (2008) Tolerance to sulphur dioxide in transgenic Chinese cabbage transformed with both the superoxide dismutase containing manganese and catalase genes of Escherichia Coli. Sci Hort 115: 101-110.
- Gill SS, Tuteja N (2010) Polyamines and abiotic stress tolerance in plants.Plant Signal Behav 5: 26-33.
Citation: Ghosh N, Adak MK (2016) Effects of Putrescine on Anti-Oxidative Enzymes in Two Rice Cultivars Subjected To Salinity. Adv Crop Sci Tech 4:210. DOI: 10.4172/2329-8863.1000210
Copyright: © 2016 Ghosh N, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Select your language of interest to view the total content in your interested language
Share This Article
Recommended Journals
Open Access Journals
Article Tools
Article Usage
- Total views: 13752
- [From(publication date): 4-2016 - Aug 29, 2025]
- Breakdown by view type
- HTML page views: 12189
- PDF downloads: 1563