Review Article Open Access
How Influenza A Virus Causes ‘‘Epidemics’’ and ‘‘Pandemics’’ among the Populations?
Sleman SS*Department of Microbiology, College of Veterinary Medicine, University of Sulaimani, Sulaimanyah, Iraq
- *Corresponding Author:
- Sleman SS
Department of Microbiology, College of Veterinary Medicine
University of Sulaimani, Sulaimanyah, Iraq
E-mail: sirwan.sleman@univsul.edu.iq
Received date: April 26, 2017; Accepted date: May 5, 2017; Published date: May 11, 2017
Citation: Sleman SS (2017) How Influenza A Virus Causes ‘‘Epidemics’’ and ‘‘Pandemics’’ among the Populations?. J Infect Dis Ther 5:319. doi: 10.4172/2332-0877.1000319
Copyright: © 2017 Sleman SS. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Journal of Infectious Diseases & Therapy
Abstract
Influenza is a major cause of sickness and death around the world and is one of the most important infectious diseases confronting the world today. Influenza A virus is the leading cause of seasonal epidemic and pandemic flu globally. Both epidemic and pandemic flu are found to be associated with the high morbidity and mortality rates worldwide, but that pandemic is thought to have accounted for about 50 million or more deaths and is obviously a matter of great concern, can cause a severe, easily transmissible disease among the community and everyone at risk of infection. The purpose of this study is to describe the mechanisms by which the influenza A viruses can cause epidemic and pandemic flu worldwide. In conclusion the extensive infection is mainly because the influenza A virus, like other RNA viruses, has capability to undergo a continuous antigenic variation either via a point mutation causing gradual change in the shape of the HA surface antigens known as ''Antigenic drift'' and it is found to be responsible for seasonal epidemics or through a genetic reassortment that leads to a major change in the surface antigens known as ''Antigenic shift'' and generation of entirely novel strains among the population with no pre-existing immunity as a result sporadic pandemics will occur.
Keywords
Pandemic flu; Seasonal flu; Human flu; Antigenic shift; Antigenic drift
Introduction
Influenza or flu is a contagious respiratory disease of birds, human and many other mammals and it is caused by influenza viruses. Influenza viruses are of three types A, B and C but only type A is found to cause severe epidemics and pandemics among human population. This is because the influenza A virus is serologically has several different subtypes based on combination between their surface antigens known as Haemagglutinin (HA) and Neuraminidase (NA) [1], which are 18 and 11 in number respectively [2]. This serological variation is thought to enable the virus toundergo a constant antigenic variation to evade host immune system by using different mechanisms and as a result new strains with new surface antigens will develop which cannot be recognized by the pre-existing immunity arouse against the older virus. Thus the new variants cause infection and spread rapidly among the previously resistant population causing an epidemic or my spread more extensively and globally causing a rare pandemic [1].
Influenza A virus
Influenza A virus is an enveloped RNA virus that has two major surface proteins known as Haemagglutinin (HA) and neuraminidase (NA) [3]. The HA is a molecule that responsible for binding to specific receptors on the host cell surface where by which the virus picks up and produces infection whilst neuraminidase molecule is an important enzyme that releases new virus particles are produced by the host cell and also cleaves sialic acid from the host cell surface and virion envelope to prevent aggregation of emerging viral particles [4,5].
Influenza A viruses are reported to have 18 different HA molecules (H1-18) and 11 different NA molecules (NA1-11) on their cell surfaces till now (Table 1) [2]. Based on the combination of these two proteins, many different subtypes can be produced and each with a specific immunity [1].
| H Subtypes | Human | Bird | Swine | Bat | N Subtypes | Human | Bird | Swine | Bat |
|---|---|---|---|---|---|---|---|---|---|
| H1 | 1934 | 1976 | 1930 | N1 | 1934 | 1959 | 1930 | ||
| H2 | 1957 | 1973 | N2 | 1957 | 1965 | 1970 | |||
| H3 | 1968 | 1963 | 1970 | N3 | 1963 | ||||
| H4 | 1956 | N4 | 1968 | ||||||
| H5 | 1997 | 1961 | N5 | 1972 | |||||
| H6 | 1965 | N6 | 1956 | ||||||
| H7 | 1996 | 1927 | N7 | 2003 | 1927 | ||||
| H8 | 1968 | N8 | 1963 | ||||||
| H9 | 1999 | 1966 | N9 | 2013 | 1974 | ||||
| H10 | 1949 | N10 | 2009 | 2009 | |||||
| H11 | 1956 | N11 | 2010 | 2010 | |||||
| H12 | 1976 | ||||||||
| H13 | 1977 | ||||||||
| H14 | 1982 | ||||||||
| H15 | 1983 | ||||||||
| H16 | 2005 | ||||||||
| H17 | 2009 | 2009 | |||||||
| H18 | 2010 | 2010 |
Table 1: This table illustrate Influenza A virus subtypes reported in human and animals [2].
All influenza A strains are basically found in birds (with exception of H17N10 and H18N11 which have only detected in bats) [2] and only H1N1, H1N2 and H3N2 has described to infect human and pass person to person (Table 1). In addition, there are some other strains of the avian influenza A viruses can human infection such as H5N1, H7N2, H7N3, H7N7, H9 [6], H7N9 [7], H17N10 H18N11 [2] but hopefully they are still not found to transmit among human populations and causing epidemics or pandemics [6].
Epidemic and pandemic flu
Epidemic flu is a seasonal outbreak that attacks many people at about the same time and may spread through one or several countries and it occurs due to re-emergence of the previous year’s virus strain with a slight change in its surface binding proteins that may not be recognizable by the body's immune system. Whereas, pandemic flu is an extensive epidemic or a worldwide outbreak that caused by the introduction of new influenza A virus strain that can pass easily from person to person with no pre-existing immunity and everybody will be at risk of gaining the infection [8,9]. By far four major influenza pandemics occur globally in the 20th and the 21st centuries which are Spanish flu in the 1918/1919, 'Asian flu' in the 1957/1958, 'Hong Kong flu' in the 1968/1969 and 'Pandemic (H1N1)' in (2009–2010) and each caused by different strains of influenza A. The most deadly pandemic was the Spanish flu outbreak, which had caused the highest morbidity and mortality rate among the population, probably more than 20 million people had died globally [5,10]. This is increasing the risk that other pandemics may occur in the future either though the reintroduction of older avian strains (especially H3, H5 and H7 that are now circulating among the population) or by the generation of other new human strains [11]. Therefore, the well understanding of influenza A strains and the process of their antigenic variation are essential and would be useful in dealing with any other possible epidemics and pandemics in the future.
Mechanism of Influenza A Virus Antigenic Variation
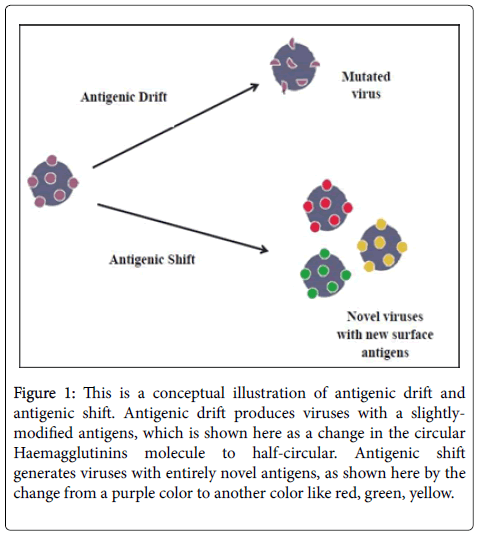
The mechanisms of surface antigenic variation by the influenza A viruses are commonly known as “antigenic drift” and “antigenic shift”. Antigenic drift is typically defined as a small change in the HA antigens caused by point mutation and immune selection pressure that directly leads to change of the HA epitopes (Figure 1). As it is well known, epitopes are binding sites of antigen that can be recognised by the components of host immune system, especially neutralizing antibodies. Therefore, any changes happen to these epitopes may not be recognition by the pre-existing antibody i.e. the new variant will replace the older ones and it may no longer or less effectively be recognized by the host immune system. As a result it rapidly spread through the same population that had developed immunity to the previous strain leading to an epidemic arise. The epidemic outbreak occurs every few years and vaccines are updated every year based on strains projected to be circulating to minimize the infection. However, some people may still normally produce immunity against newly produced viruses because they still have some similarity to the older strains [12-14].
Figure 1: This is a conceptual illustration of antigenic drift and antigenic shift. Antigenic drift produces viruses with a slightlymodified antigens, which is shown here as a change in the circular Haemagglutinins molecule to half-circular. Antigenic shift generates viruses with entirely novel antigens, as shown here by the change from a purple color to another color like red, green, yellow.
While antigenic shift is generally described as a process of a sudden, major change of influenza A surface proteins (mainly HA) resulting from the recombination of the viral genomes of two different strains which may lead to introduction of entirely new strain into the human population with little or no immunity to them (Figure 1). So that everyone are at risk of acquiring infection and extensive epidemics or pandemics will result. The source of these new strains is primarily avian influenza viruses that found naturally in migratory wild birds (the natural reservoir) [1,12-14].
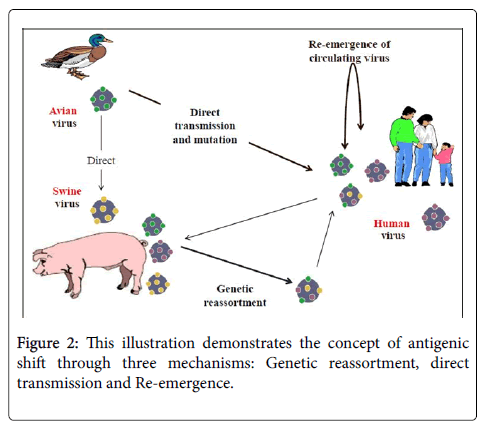
Generally, antigenic shift are described to occur through several ways: genetic reassortment; direct transmission and adaptation; reintroduction of old strains (Figure 2).
Genetic reassortment
It occurs through cross linking between two different strains of influenza A viruses in a single host cell (Figure 1), where the RNA segments from the two strains are mixed together during the new virus particles reassembly. As a result, a third viral strain will arise which contains the combined genes of both strains [12,15,16]. Pigs and humans possess both receptors of human and avian influenza virus strains so that they can be infected by both strains at the same time leading to the mixed gene strains to emerge. 'Asian flu' in the 1957/1958, 'Hong Kong flu' in the 1968/1969 and Pandemic (H1N1) in (2009-2010) are good examples that are thought to be caused by genetic reassortment [10].
Direct transmission and adaptation
The direct transmission of avian influenza A, especially with a highly pathogenic strain, has found to occur from birds to humans and then by adaptation of the virus in humans. This adaptation has occurred through changing the receptor binding site of HA from α 2,3 linked sialic acid receptors in avian cells into α 2,6 linked sialic acid receptors in human cells by mutation. As a result, a novel human strain might generate and be capable to transmit among the population and causing a pandemic. The ''Spanish flu'' in 1918/1919 is an example of pandemics that was thought to be transmitted by direct contact from avian to human beings based on the characteristic features of genetic material isolated from victims [10].
Re-emergence of old strains
Once the population’s immunity becomes weakened to a specific stain, then the virus can re-emerge and causing a pandemic outbreak among that population. For example, the Russian flu in 1950 and 1977 is found that it was probably caused by the re-emergence of the same H1N1 virus subtype [10].
Conclusion
It is clear that influenza A virus is a real threat to human health, mostly due to the presence of non-eradicable reservoirs among migratory birds and the continuous capability of influenza A virus to undergo antigenic variation to evade the host immune system either through a slight gradual change in the surface glycoprotein's of the previous subtype (antigenic drift) causing seasonal epidemics or by a sudden big change in surface antigens (antigenic shift) with the emergence of entirely new strain that can cause serious illness and spread quickly from human-to-human resulting a rare pandemics.
Acknowledgement
Special thanks to the staff of Department of Microbiology and Virology, University of Manchester, UK.
Conflict of Interest
No conflict of interest
References
- Khan FH (2009) The Elements of Immunology. Dorling Kindersley, Pearson education.
- CDC (2014) Influenza (flu): Transmission of Influenza Viruses from Animals to People.
- Noda T (2012) Native morphology of influenza virions. Front Microbiol 2: 136-137.
- Stanley J (2002) Essentials of Immunology & Serology. Influenza Virus 138.
- Taubenberger JK and Kash JC (2010) Influenza Virus Evolution, Host Adaptation and 139 Pandemic Formation. Cell host & microbe 7: 440-451.
- WHO (2013) Influenza.
- WHO (2013) Overview of the emergence and characteristics of the avian influenza 143 A(H7N9) virus 1-38
- Nordqvist C (2014) "What is a pandemic? What is an epidemic?" Medical News Today.
- Morens DM, Folkers GK and Fauci AS (2009) What is a pandemic? J Infect Dis 200: 1018-1021.
- Knowledge Centre (2014) Pandemic Flu: How do pandemics Occur. ESWI.
- Influenza Pandemics–history (2012) Health Protection Agency.
- Parija SC (2009) Textbook Of Microbiology And Immunology. Mosby: Elsevier.
- CDC (2015) Influenza: Epidemiology and Prevention of Vaccine-Preventable Diseases 187-208.
- Glezen WP, Couch RB (1997) Influenza viruses. In: Evans AS, and Kaslow RA, Viral Infections of Humans. Epidemiology and Control 473–505
- Microblog (2006) Antigen Shift vs Antigenic Drift.
- Racaniello V (2013) Influenza A viruses in bats. Virology blog.
Relevant Topics
- Advanced Therapies
- Chicken Pox
- Ciprofloxacin
- Colon Infection
- Conjunctivitis
- Herpes Virus
- HIV and AIDS Research
- Human Papilloma Virus
- Infection
- Infection in Blood
- Infections Prevention
- Infectious Diseases in Children
- Influenza
- Liver Diseases
- Respiratory Tract Infections
- T Cell Lymphomatic Virus
- Treatment for Infectious Diseases
- Viral Encephalitis
- Yeast Infection
Recommended Journals
Article Tools
Article Usage
- Total views: 8537
- [From(publication date):
June-2017 - Aug 16, 2025] - Breakdown by view type
- HTML page views : 7512
- PDF downloads : 1025