Review Article Open Access
Low Serum Alpha-fetoprotein Level an Important Predictor for Therapeutic Outcome in Egyptian Patients with Chronic Hepatitis C: A Data-Mining Analysis
Naglaa Zayed1*, AbuBakr Awad2, Wafaa El-Akel1, Wahid Doss1,3, Maissa El-Raziky1 and Mahasen Mabrouk1
1Endemic Medicine and Hepatology Department, Faculty of Medicine, Cairo University, Egypt
2Computer Science Department, Faculty of Computers and Information, Cairo University, Egypt
3National Hepatology and Tropical Medicine Institute, Ministry of Health and Population, Cairo, Egypt
- *Corresponding Author:
- Naglaa Zayed
Kasr Al-Aini Street, Faculty of Medicine
Cairo University, Cairo, Egypt
Tel: +201001460366
E-mail: naglaazayed@yahoo.com
Received date: October 09, 2014; Accepted date: November 12, 2014; Published date: November 18, 2014
Citation: Zayed N, Awad A, El-Akel W, Doss W, El-Raziky M, et al. (2014) Low Serum Alpha-fetoprotein Level an Important Predictor for Therapeutic Outcome in Egyptian Patients with Chronic Hepatitis C: A Data-Mining Analysis . J Gastrointest Dig Syst 4:240. doi:10.4172/2161-069X.1000240
Copyright: ©2014 Zayed N, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Journal of Gastrointestinal & Digestive System
Abstract
Background Data mining can build predictive models for the response to antiviral therapy in chronic HCV patients. Objective To develop a prediction model for therapeutic outcome in chronic HCV genotype-4 patients using different decision-trees learning algorithms. Study Design Data of 3719 chronic HCV patients who had received PEG-IFN/RBV therapy at Cairo-Fatemia Hospital, Egypt was retrieved. Factors predictive of SVR were explored using data mining analysis. Weka implementations C4.5, classification and Reduced Error Pruning tree were constructed using 22 attributes from initial patients’ data. Results End of treatment response and estimated SVR were 61.6%, 52.5% respectively. Low median AFP; 2.9 ng/ml was significantly associated with SVR; compared to relapse group 5.06 ng/ml; p value<0.01. AFP was identified as the most decisive variable of initial split by both decision-tree models. Various cutoff levels were related to different probability of SVR. Baseline AFP ≤2.48 ng/ml was associated with 72%SVR while levels ≥ 7.8 ng/ml demonstrated 32%. Other attributes such as age, BMI, ALT, hepatic fibrosis and activity were less decisive in prediction of response. This was further confirmed by univariate logistic regression analysis; p value<0.01. Conclusion Low AFP Levels were significantly related to SVR in an HCV population presumably genotype-4 as demonstrated by data mining.
Keywords
AFP; Egypt; Fast decision tree learner (FDTL); HCV; Pegylated interferon; Reduced Error Pruning (REP); Ribavirin; Sustained virological response (SVR); Weka implementation C4.5
Abbreviations
AFP: Alpha Feto-Protein; ALP: Alkaline Phosphatase; ALT: Alanine Aminotransferase; ANA: Anti-Nuclear Antibody; ANC: Absolute Neutrophil Count; AST: Aspartate aminotransferase; BMI: Body Mass Index; FDTL: Fast Decision Tree Learner; Hb: Hemoglobin; n: number; PEG-IFN: pegylated interferon; PLT: Platelet Count; RBV: Ribavirin; REP: Reduced Error Pruning; TSH: Thyroid Stimulating Hormone; SVR: Sustained Virologic Response; WBC: While Blood Cell Count
Background
A 48-week course of antiviral therapy with pegylated interferon (PEG-IFN) and ribavirin (RBV) therapy is the cornerstone of treatment for chronic HCV patients. However, its cost, adverse events in addition to the sub-optimal sustained virologic response (SVR) in patients with genotype1 and 4 of 42%–52% [1-3] had considered these patients as difficult to treat. In Egypt, data regarding predictive factors of HCV genotype-4 are relatively scarce. Thus the possible prediction of response will have a great value on the response rate and the national economic burden as well. Clinical data have provided new insight into HCV-4 infection and have resulted in a refinement of the treatment strategies and eventual optimization of treatment regimens [4]. Baseline viremia, early viral kinetics, treatment duration, and stage of liver disease were considered and thus can determine the therapeutic outcomes for chronic HCV patients [5].
Data mining analysis using decision-tree is one of the most effective machine learning approach for extracting practical knowledge and exploration of previously unknown and potentially valuable information from large amounts of from real world datasets [6]. Decision-tree can be basic such as C4.5 algorithm [7] and scalable that can classify data sets with hundreds of attributes with reasonable speed; Reduced Error Pruning (REP) Tree as a fast decision tree learner (FDTL) [8]. These decision-tree algorithms have high accuracy in classifying large medical data such as clinical and laboratory parameters, which are sometimes difficult to interpret by clinicians, can handle both categorical and continuous attributes, and can handle missing values to build a valid decision-tree and predictive models. Thus the linearity of the relationship between covariates and outcome have been always a subject of debate [9] specially like the situation of HCV in Egypt where clinicians are provided with an enormous amount of datasets which may be difficult to interpret and relate with patients’ outcome. Previous studies have built a prediction model for the early virological response as well as SVR among HCV patients using simple and noninvasive as parameters in addition to specialized tests such as viral mutations and host genetics [10,11]. However these studies were performed on HCV genotype-1 and included small number of patients.
Objective
In this study, we compiled a database of clinical data from 3719 Egyptian patients with chronic HCVgenotype-4. We used Weka implementation C4.5 decision-tree algorithm and Reduced Error Pruning (REP) tree as one of the Fast decision tree learner (FDTL) algorithms to build a model for the prediction of SVR in this subset of patients.
Study Design
Study population: This cross sectional observational study was conducted on baseline data belonging to 3719 adult Egyptian patients with chronic HCV infection of both sexes who were diagnosed by reactive anti-HCV antibodies, positive HCV-RNA, histological evidence of chronic hepatitis with negative HBsAg and no other causes of chronic liver disease. In Egypt, >95% of infections are HCV genotype 4 (HCV-G4) HCV so genotypes was not performed [12,13]. Patients had been treated with Peg-IFN alpha 2a or 2b plus weight-based RBV [14,15] from 2008-2011 at Cairo-Fatemic Hospital, MOHP-Cairo-Egypt.
This study is based on the retrospective analysis of clinical and laboratory data collected as part of the routine management of chronic HCV. No additional testing ws ordered other than that performed as part of routine management of patients according to the guidelines National Committee for the Control of Viral Hepatitis, MOHP. A specific study code number was created for each patient so that no name or identifier appears in the database.
The study was done in the context of the project: Bioinformatics in predicting the response to interferon-ribavirin combination therapy in patients with HCV genotype-4 (Bio-IN-therapy) which was financially supported by the Science and Technology Development Fund (STDF), Egypt, Grant No.1708.The project was approved by the ethics committee of the MOHP and all patients were previously consented for the blood sampling and possible data application in future research.
Data Collection
Enrollment and follow up data: A standardized enrollment questionnaire which was previously completed by patients’ physicians was retrieved from patients’ medical records. The initial questionnaire included medical number, full name and contact details for each patient in addition to the demographic data such as age, gender, body mass index (BMI). Laboratory parameters in the form of hematological tests, liver biochemical profile, ANA(anti-nuclear antibody), serum creatinine, alpha feto-protein (AFP), TSH(thyroid stimulating hormone), anti-schistosomal antibody, fasting or radom glucose level in addition to the HCV viral load, grade of hepatic necro-inflammation and stage of fibrosis according to Metavir score [12] were also included. Follow up visits included clinical and laboratory assessment to report possible adverse side effects and treatment response.
Data cleansing: Data cleansing was applied due to the fact that most patient records have suffered from missing data, typos, and the usage of different measuring standards. High quality data was characterized by accuracy, integrity, completeness, validity, consistency, uniformity and uniqueness.
Feature selection and reduction: A subset of 22 features (categorical or numerical) was selected to reduce the dimensionality of the problem and speedup the model building process (Table 1). The database was created containing three demographic variables (age, gender, BMI), four hematological variables[hemoglobin, white blood cells(WBC), absolute neutrophil count(ANC) and platelets], nine biochemical variables [blood sugar, serum bilirubin, albumin, alanine aminotransferase(ALT), alkaline phosphatase(ALP), aspartate aminotransferase(AST) folds elevation, prothrombin concentration, creatinine and AFP] in addition to ANA, TSH, HCV-RNA levels ,type of PEG-IFN and histolopatholgic features of chronic hepatitis.
| Parameter | Odds ratio | SE | 95% CI | p value |
|---|---|---|---|---|
| AFP >10ng/ml | 1.1 | 0.01 | 1.0-1.1 | <0.01 |
| Male gender | 0.67 | 0.08 | 0.53-0.85 | 0.001 |
| Grade of necro-inflammation (A≥2) | 0.47 | 0.06 | 0.37-0.59 | <0.01 |
| Stage of fibrosis (≥ F2) | 1.5 | 0.18 | 1.2-1.9 | 0.001 |
| HCV viral load ( >600 IU/ml) | 1.7 | 0.21 | 1.3-2.2 | <0.01 |
| ALT( >40 IU/L) | 1.0 | 0.0 | 0.99-0.99 | 0.01 |
| AST (>40 IU/l) | 1.0 | 0.0 | 0.99-0.99 | 0.01 |
Table 1: Multivariate Logistic regression analysis for baseline factors associated with failure of treatment (no SVR). AST:aspartate aminotransferase; ALT: alanine aminotransferase; AFP: alpha fetoprotein; n: number. Data expressed as mean ± SD.
Data formatting: A number of data transformation techniques have been used to format and prepare the patient records to be processed by the learning algorithms
Data mining: Weka implementation of C4.5 (weka J48) [7] and Reduced Error Pruning (REP) Tree as a fast decision tree learner (FDTL) [8] have been used to build the predictive model for the likelihood of SVR. Patients who discontinued the treatment were excluded from model building non-responders regardless to other factors.
Validation of the decision tree: Internal validation was performed with Test mode: 10-fold cross-validation which is a generally applied to predict the performance of a model on a validation set using computation in place of mathematical analysis.
Performance of algorithms: The performance of the algorithms was assessed according to evaluation matrix based on values for the correctly classified instance, precision, recall, F-score, and Receiver operating characteristic (ROC) ROC curve.
Correctly classified instances: evaluate the overall accuracy.
Recall (Sensitivity): the ability of the test to correctly identify those with the disease (true positive rate).
Precision (Specificity): the ability of the test to correctly identify those without the disease (true negative rate).
F-Measure: represents the weighted average of the precision and recall.
Area under the curve (AUC) serves as an indicator of the overall performance of the algorithm.
Statistical Analysis A descriptive statistical analysis was conducted to study the distributions of most dataset features. Student’s t-test was used for the invariable comparison of quantitative variables and Fisher’s exact test was used for the comparison of qualitative variables. For the multivariable analysis for factors associated with virologic response, logistic regression models with backward selection were used to identify independent predictors of SVR. Variables that showed significant association with SVR by univariate analysis were included in the multivariate analysis.
Results
End of treatment response (ETR) at week 48 was evident in 2277 patients (61.6%), failure of response and breakthrough were evident in 1442 patients (38.7%) including 492 patients (34%) who dropped to follow up and discontinued the treatment either due to side effect or non-compliance. At week 72-estimated SVR was 54.2% with a relapse rate 7.4%.
Initial pre-treatment 22 variables both categorical and numerical data associated with the likelihood of response were used for model building dataset (n=3719). Low median baseline AFP level was significantly related to for the achievement of ETR and SVR (3 ng/ml, 2.91 ng/ml) versus non-ETR and relapse (4.42, 5.06 ng/ml) respectively; p value <0.01(Figure 1). Other baseline factors associated with failure of treatment (no SVR) were shown in the multivariate logistic regression analysis (Table 1).
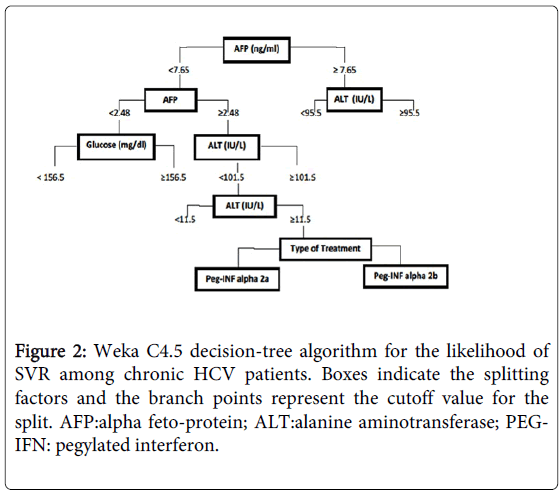
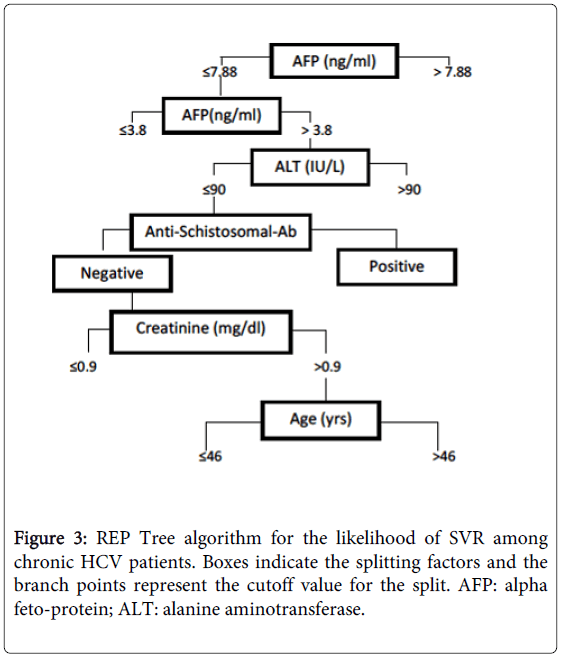
Both decision-trees models revealed that in patients who were adherent to treatment, AFP was selected as the variable of initial split (most decisive). Various cutoff levels of baseline AFP at 2.48, 3.4, 4.8 and 7.8 ng/ml were set and were able to classify patients according to SVR (Figures 2 and 3). Among patients who were adherent to treatment, patients with baseline AFP levels ≤2.48 ng/ml were classified as high probability group; SVR (72%) while in those with AFP levels >7.8 ng/ml, SVR dropped to 32%(low probability group).
On the other hand, patients with AFP levels between the cutoff values, other attributes as age, BMI, ALT, anti-schistosomal antibodies, prothrombin concentration, serum creatinine, type of pegylated IFN, HCV-RNA viral load, hepatic fibrosis and activity, had less decisive role for prediction of response. These results were confirmed statistically using univariate logistic regression analysis; p- value < 0.01.
Decision-trees performance parameters
The performance parameters for both the C4.5 and REP tree models were comparable. However, the REP tree consumed less time to build the predictive model as shown in Table 2.
| Performance Parameters | C4.5 | REP Tree |
|---|---|---|
| Time taken to build model | 0.25 seconds | 0.07 seconds |
| Correctly Classified Instances | 68.97% | 68.84% |
| Precision | 0.655 | 0.652 |
| Recall | 0.69 | 0.688 |
| F-Measure | 0.631 | 0.627 |
| AUC | 0.594 | 0.614 |
Table 2: Comparison between C4.5 and REP Tree using the standard performance parameters AUC: area under the curve.
Discussion
This study confirms our previous findings regarding the predictive role of AFP with regard to the likelihood ETR in chronic hepatitis C presumably genotype-4 where we have applied one of the decision-tree algorithms [16]. In this study, we applied both C4.5 and REP decision-tress for treatment outcome to PEG-IFN/ RBV therapy among Egyptian patients infected with HCV and explored that various cutoff levels for baseline AFP at 2.48, 3.8, 4.8, 7.8 ng/ml were identified and patients were classified into subgroups with different probabilities of developing SVR. Based on these results, clinicians might consider serum AFP to be of the same magnitude compared to the other well-accepted predictive factors of treatment response such as HCV genotype and viral load. Other variables such as patients’ gender, serum ALT, stages of hepatic fibrosis and grades of activity had less decisive roles for prediction. Because each of the above values can be determined and easy interpretable by clinicians prior to therapy, they may be considered as a guide to establishing a treatment regimen in the potentially difficult-to-treat patients.
What’s unique in this study that the decision-trees were able to determine a very low baseline AFP levels (≤2.48 ng/ml) among chronic HCV genotype-4 that was associated with very high probability of SVR (72%) which approaches other rates in other easy treatable genotypes (2 and 3). This was further elaborated by conventional statistical analysis which demonstrated that low median baseline AFP (2.9g/ml) was significantly related to for the achievement of SVR.
In Egypt, various patients’ demographics, viral factors in addition to the extent of the disease have been used as prognostic factors for the therapeutic outcome in chronic HCV patients. Traditional well known predictors like pre-treatment viral load and extent of hepatic fibrosis have been previously studied while other non-conventional factors like AST and AFP were identified to be predictive. In a previous study of 250 Egyptian genotype-4 patients, the presence of severe fibrosis, hepatic steatosis, treatment with conventional interferon, and AFP level were found to predict SVR [14]. These findings were further supported by further studies which augmented the pool of evidence regarding the association between higher AFP level and negative treatment outcome [17,18] . Similar findings have been found in genotype 1 patients as well [19]. Moreover, serum AFP levels, might serve as surrogate markers of advanced fibrosis [20,21] and hence, the finding of the present study that advanced fibrosis is associated with failure of SVR. Serum AFP was independently associated with SVR after controlling other known factors associated with SVR, including liver fibrosis in a previous study among Egyptian chronic HCV patients [22]. The expression of hepatic progenitor cells (HPC), which are present in the peri-portal region, has been associated with response to treatment [23]. They were found to express highlevels of AFP and certain keratin markers [24,25] and their presence was related to the severity of fibrosis [23].
Baseline status of demographic and routine assessment parameters between patients achieving and not achieving SVR were compared among HCV genotype-1 patients. There was no significant difference in age of patients achieving SVR compared to those who did not (45.2 vs. 48.8 years, p=0.09). Moreover, neither sub-genotype (1a versus 1b) nor gender was associated with SVR; different stages of hepatic fibrosis were not significant as well. However, baseline HCV RNA below 5.6 log IU/mL was significantly associated with SVR [26].
An advantage of decision-tree analysis over traditional regression models is that the decision-tree model is user-intuitive and can be easily interpreted by medical professionals without the need for any specific knowledge of statistics. Clinicians often experience difficulty applying standard statistical methods to assess the interactions between clinical variables, determining the cumulative effect of these variables on response and disease progression and later translating this information into appropriate management.
In a previous study, a simple decision-tree model (CART analysis) was applied to explore the possible pre-treatment predictive factors for the response to PEG-IFN using. Hepatic steatosis (<30%) was identified as the first predictor of response followed by low-density-lipoprotein cholesterol (LDL-C) (≥100 mg/dL), age (<50 and <60 years), blood sugar (<120 mg/dL), and gamma-glutamyltransferase (GGT) (<40 IU/L). Patients were classified based on these variables to low (16%), intermediate (46%) and high (75%) probabilities of achieving rapid and early virologic response among difficult to treat chronic hepatitis C patients [10].
In clinical practice, the possible applications of these complementary approaches; logistic regression and decision-tree algorithms; can improve confidence in the results and partially protect against any intrinsic bias. Comparing the results of a standard analysis with an alternative technique may reveal the most robust and sensitive variables. On the other hand, decision-tree which are widely used in biomedical studies [27,28] may provide a simple and hierarchical format that can be used without a computer.
Conclusion
Both prediction models decision-trees analysis classified patients with high probabilities for SVR according to AFP levels. Using this model, an estimate of SVR can be rapidly obtained before treatment, and thus may facilitate clinical decision making. Nonetheless, the growing recognition of AFP as a predictor of response, given its uniqueness, mandates further evaluation and thus the estimation of low probability should not be used to preclude patients from therapy, and the final decision should be made on a case-by-case basis.
References
- Esmat G, Abouzied A, Abdel-Aziz F. Treatment with PEG-IFN alfa-2b plus ribavirin compared to interferon alfa-2b plus ribavirin in subjects with chronic hepatitis Cinfected with HCV genotype 4. Hepatology 2002; 36: 364A(Abstract).
- Alfaleh FZ, Hadad Q, Khuroo MS, Aljumah A, Algamedi A, et al. (2004) Peginterferon alpha-2b plus ribavirin compared with interferon alpha-2b plus ribavirin for initial treatment of chronic hepatitis C in Saudi patients commonly infected with genotype 4. Liver Int 24: 568-574.
- El-Zayadi AR, Attia M, Barakat EM, Badran HM, Hamdy H, et al. (2005) Response of hepatitis C genotype-4 naïve patients to 24 weeks of Peg-interferon-alpha2b/ribavirin or induction-dose interferon-alpha2b/ribavirin/amantadine: a non-randomized controlled study. Am J Gastroenterol 100: 2447-2452.
- Kamal SM, Nasser IA (2008) Hepatitis C genotype 4: What we know and what we don't yet know. Hepatology 47: 1371-1383.
- Fontana RJ, Everson GT, Tuteja S, Vargas HE, Shiffman ML (2004) Controversies in the management of hepatitis C patients with advanced fibrosis and cirrhosis. ClinGastroenterolHepatol 2: 183-197.
- Mahmood AM, Kuppa MR, Reddi KK. A new decision tree induction using composite splitting criterion. Journal of Applied Computer Science & Mathematics 2010; 9:69–74.
- Quinlan JR. C4.5: Programs for Machine Learning, San Mateo, CA, Morgan Kaufmann (1993).
- [8].Mehta M, Agrawal R, Riassnen J. SLIQ: a fast scalable classifier for data mining. Extending database technology. Springer, 1996; Avignon 18–32.
- Lemon SC, Roy J, Clark MA, Friedmann PD, Rakowski W (2003) Classification and regression tree analysis in public health: methodological review and comparison with logistic regression. Ann Behav Med 26: 172-181.
- Kurosaki M, Matsunaga K, Hirayama I, Tanaka T, Sato M, et al. (2010) A predictive model of response to peginterferon ribavirin in chronic hepatitis C using classification and regression tree analysis. Hepatol Res 40: 251-260.
- Kurosaki M, Sakamoto N, Iwasaki M, et al. Pretreatment prediction of response to peginterferon plus ribavirin therapy in genotype 1 chronic hepatitis C using data mining analysis. J Gastroenterol. 2011; 46:401-9.
- Bedossa P, Poynard T (1996) An algorithm for the grading of activity in chronic hepatitis C. The METAVIR Cooperative Study Group. Hepatology 24: 289-293.
- Ray SC, Arthur RR, Carella A, Bukh J, Thomas DL (2000) Genetic epidemiology of hepatitis C virus throughout egypt. J Infect Dis 182: 698-707.
- Manns MP, McHutchison JG, Gordon SC, Rustgi VK, Shiffman M, et al. (2001) Peginterferon alfa-2b plus ribavirin compared with interferon alfa-2b plus ribavirin for initial treatment of chronic hepatitis C: a randomised trial. Lancet 358: 958-965.
- Hadziyannis SJ, Sette H Jr, Morgan TR, Balan V, Diago M, et al. (2004) Peginterferon-alpha2a and ribavirin combination therapy in chronic hepatitis C: a randomized study of treatment duration and ribavirin dose. Ann Intern Med 140: 346-355.
- Zayed N, Awad AB, El-Akel W, Doss W, Awad T, et al. (2013) The assessment of data mining for the prediction of therapeutic outcome in 3719 Egyptian patients with chronic hepatitis C. Clin Res HepatolGastroenterol 37: 254-261.
- Gad RR, Males S, El Makhzangy H, Shouman S, Hasan A, et al. (2008) Predictors of a sustained virological response in patients with genotype 4 chronic hepatitis C. Liver Int 28: 1112-1119.
- Males S, Gad RR, Esmat G, Abobakr H, Anwar M, et al. (2007) Serum alpha-foetoprotein level predicts treatment outcome in chronic hepatitis C. AntivirTher 12: 797-803.
- Akuta N, Suzuki F, Kawamura Y, Yatsuji H, Sezaki H, et al. (2007) Predictors of viral kinetics to peginterferon plus ribavirin combination therapy in Japanese patients infected with hepatitis C virus genotype 1b. J Med Virol 79: 1686-1695.
- Murashima S, Tanaka M, Haramaki M, Yutani S, Nakashima Y, et al. (2006) A decrease in AFP level related to administration of interferon in patients with chronic hepatitis C and a high level of AFP. Dig Dis Sci 51: 808-812.
- Chen TM, Huang PT, Tsai MH, et al. Predictors of alpha-fetoprotein elevation in patients with chronic hepatitis C, but not hepatocellular carcinoma, and its normalization after pegylated interferon alfa 2a-ribavirin combination therapy. J GastroenterolHepatol. 2007; 22:669–75.
- Abdoul H, Mallet V, Pol S, Fontanet A (2008) Serum alpha-fetoprotein predicts treatment outcome in chronic hepatitis C patients regardless of HCV genotype. PLoS One 3: e2391.
- Tsamandas AC, Syrokosta I, Thomopoulos K, Zolota V, Dimitropoulou D, et al. (2006) Potential role of hepatic progenitor cells expression in cases of chronic hepatitis C and their relation to response to therapy: a clinicopathologic study. Liver Int 26: 817-826.
- Germain L, Noël M, Gourdeau H, Marceau N (1988) Promotion of growth and differentiation of rat ductular oval cells in primary culture. Cancer Res 48: 368-378.
- Dabeva MD, Shafritz DA (1993) Activation, proliferation, and differentiation of progenitor cells into hepatocytes in the D-galactosamine model of liver regeneration. Am J Pathol 143: 1606-1620.
- Alestig E, Arnholm B, Eilard A, Lagging M, Nilsson S, et al. (2011) Core mutations, IL28B polymorphisms and response to peginterferon/ribavirin treatment in Swedish patients with hepatitis C virus genotype 1 infection. BMC Infect Dis 11: 124.
- Augustin S, Muntaner L, Altamirano JT, et al. Predicting early mortality after acute variceal hemorrhage based on classification and regression tree analysis. Clinical Gastroenterology and Hepatology 2009; 7:1347-54.
- El Malki HO, El Mejdoubi Y, Souadka A, Mohsine R, Ifrine L, et al. (2010) Predictive model of biliocystic communication in liver hydatid cysts using classification and regression tree analysis. BMC Surg 10: 16.
Relevant Topics
- Constipation
- Digestive Enzymes
- Endoscopy
- Epigastric Pain
- Gall Bladder
- Gastric Cancer
- Gastrointestinal Bleeding
- Gastrointestinal Hormones
- Gastrointestinal Infections
- Gastrointestinal Inflammation
- Gastrointestinal Pathology
- Gastrointestinal Pharmacology
- Gastrointestinal Radiology
- Gastrointestinal Surgery
- Gastrointestinal Tuberculosis
- GIST Sarcoma
- Intestinal Blockage
- Pancreas
- Salivary Glands
- Stomach Bloating
- Stomach Cramps
- Stomach Disorders
- Stomach Ulcer
Recommended Journals
Article Tools
Article Usage
- Total views: 15475
- [From(publication date):
November-2014 - Aug 28, 2025] - Breakdown by view type
- HTML page views : 10863
- PDF downloads : 4612