Nucleic Acid Based Biosensors for Clinical Applications
Received: 12-Sep-2013 / Accepted Date: 25-Oct-2013 / Published Date: 04-Nov-2013 DOI: 10.4172/2090-4967.1000104
Abstract
Biosensors are analytical tools which have applications in diverse fields like diagnostics, disease monitoring etc. Since last few decades, nucleic acid based analytics have gained much interest in detection and monitoring of various clinical manifestations. Nucleic acid based biosensors (NABs) are preferred choice due to their high sensitivity and selectivity. Novel synthetic probes (PNA, Aptamer) have been exploited to fabricate point-of-care NABs. Due to flexible and low-cost fabrication, they are evolving as rapid and reliable tools in clinical diagnostics market. In this review, we focus on different transducer platform (Optical, Electrochemical, Piezoelectrical) based NABs which have been implemented to detect broad spectrum of metabolic and infectious diseases. A few commercialized NAB products are also summarized.
Keywords: Biosensors, Nucleic Acid Biosensors (NABs, Clinical applications
Introduction
The timely monitoring and regulation of various parameters is important to many applications in food industry, clinical diagnoses, hygiene, environmental protection, drug development, forensics etc. [1]. Therefore, there is a requirement to have reliable analytical devices, which can perform rapid and accurate analyses. Biosensors have been developed using the specificity of a biological molecule integrated to a physicochemical transducer to convert a perceived signal (usually very weak) into a comprehensible electrical signal [2]. The use of nucleic acid as a recognition layer in biosensor design is relatively a new and exciting area in analytics [3]. Nucleic acid based biosensors (NABs) have applications in
(i) clinical diagnostics of inherited diseases and pathogenic infections
(ii) laboratory analysis like probe development for different hybridization techniques and microarray systems.
NABs primarily use deoxyribonucleic acid (DNA), ribonucucleic acid (RNA), peptide nucleic acid (PNA), and aptamers (both DNA and RNA) as oligonucleotide probes. The fundamental principle behind NABs depend on sequence complementarity as per Chargaff's rules of base pairing (for DNA, A=T, G≡C) except in the case of aptamers. Principle of aptamer based detection is more akin to antigen-antibody or receptor-ligand interactions.
Some of the DNA-based sensors that can detect the presence of pathogenic micro-organisms, other infectious agents like virus, genetic polymorphisms and detection of point mutations (SNP) have reached the market recently [4].
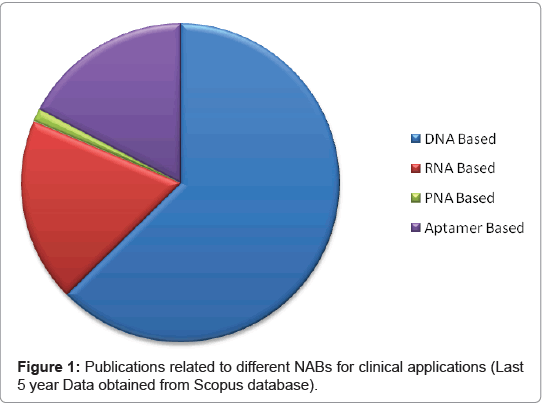
The major advantage of DNA based probes are the ability to amplify a desired target DNA from the host pathogen using PCR and consequently augment the signal generated by the biosensor. Unlike other sensing molecules like enzymes or antibodies, DNA forms easily synthesizable biological recognition layers which are highly stable and reusable after simple thermal melting of DNA duplex [5]. DNA as a sensing tool is preferred among other nucleic acid based probes (Figure 1).
Still much advancement is awaited in development of highly precise and sensitive NABs which can detect multi-analytes reducing the time and cost.
Design And Fabrication Of NABs
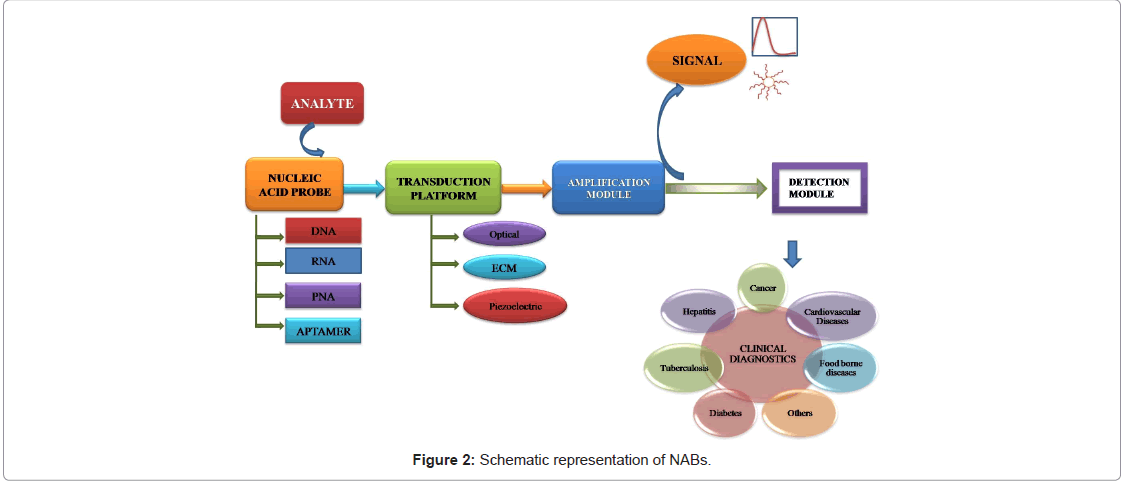
Hybridization based biosensors rely on the duplex formation between nucleic acids. Hybridization normally takes place between a known DNA sequence i.e., probe and an unknown counterpart i.e. target DNA, but DNA–RNA and RNA–RNA hybridizations can also occur [6]. The basic schema of NAB is depicted in the Figure 2. DNA probes can be synthesized by available chemical methods or PCR. But for RNA, a probe may be obtained by reverse-transcription (RT) of a previously isolated or specific messenger RNA (mRNA), or predicting its nucleotide sequence based on the amino acid sequence of the protein expressed by that DNA. Despite validated proof of the later strategy is limited due to codon degeneracy. Traditional nucleic acid hybridization methods involve gel electrophoresis and Southern and Northern blotting. These are usually lengthy and labor-intensive compared to which hybridization process occurs directly on the surface of a physical-transducer [7].
NABs are developed by immobilization of nucleic acids (DNA, RNA, PNA, oligonucleotides) to a solid support by adsorption, covalent bonding or ionic interaction and coupled to a physico-chemical transducer [8]. There are various methods available for immobilization like adsorption to inert carriers, chemical cross-linking to macroscopic beads or magnetic particles, physical entrapment in gel-like lattice, microencapsulation in nanospheres etc. The immobilization process also aids in probe orientation and ready accessibility to target element [9].
In case of physico-chemical method, various fabrication strategies are available to attach DNA probe to transducer surface. These include use of thiolated DNA for self-assembly onto gold plated transducers(electrodes, piezoelectric crystals), covalent linkages to gold surface by stabilizer molecules like 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide HCl-mediated reaction, use of biotinylated DNA to form conjugate with surface immobilized avidin or streptavidin, and carbodiimide coupling to functional groups on carbon nanotubes (SWCNTs, MWCNTs) [10]. But for solution-based hybridization assays, different conditions (ionic strength, temperature, solubility, catalyst concentration) are taken into considerations which have to be optimized in the later stage.
Smart and advanced nucleic acid recognition process involving Peptide Nucleic Acid (PNA), aptamers can augment the efficiency of DNA based biosensors. PNA is a DNA analogue in which sugar phosphate backbone is replaced with pseudo peptide. PNA is composed of repeating N-(2-aminoethyl)-glycine units linked by peptide bonds [11]. The unique structural hybridization and sensing feature of solution phase PNA can be readily extrapolated onto transducer surface to fabricate highly sensitive biosensor [12].
Aptamers are short oligonucleotides (40-60 nucleotides) which can be chemically synthesized, modified and easily integrated with a variety of nanomaterials, such as gold nanoparticles, quantum dots, carbon nanotubes, and iron oxide nanoparticles, with specific optical, magnetic, and electrochemical properties. The application of these systems as fluorescent, colorimetric, and electrochemical sensors in recent medical diagnostics is quite promising [13].
Types Of Transduction Platform
Transducer is a critical component of biosensors. It transforms resultant bio-chemical response to easily measurable output signal. NABs can be fabricated with various types of transducers like optical, electrochemical, piezoelectric, microfluidics based systems.
Optical NABs
Optical sensing for the detection of analytes is the most commonly used method in NABs. Optical biosensor is based on optical transduction of a signal and comprises ultraviolet, visible and infrared spectrophotometry in transmission or reflectance modes. Lambert- Beer principle easily correlates the relationship between incident light intensity and the transmitted radiation. Optical phenomenon like absorption, refractive indices, fluorescence, phosphorescence, and chemiluminescence are exploited to monitor the biological recognition in biosensors. One great advantage is that optical biosensors can be miniaturized by using fiber optic, semiconductor photo devices, quantum dot based detection modules.
Optical biosensors are based on fiber optics which convert the emission signal to a detectable fluorescent signal. DNA probe and target hybridization event was detected by fluorescence marker ethidium bromide. Total internal reflection in the optical fiber was measured which is proportional to the total amount of intercalated ethidium bromide [14]. Optical transmission through minuscule structures gained a new dimension with the advent of the surface plasmon resonance. SPR is the common platform to address the various types of nucleotide probes like DNA, RNA and aptamers. Using SPR sensogram analysis real time monitoring can also be done.
Molecular beacons (MB) are oligonucleotides having a stem and loop structure labeled with a fluorophore at one end and a quencher at the other. After hybridization process, amount of oligonucleotides involved can be estimated by changes in fluorescent signal. Flexibility in designing MBs and various attachment platforms make them potential candidate for biosensor application. Biotinylated ssDNA molecular beacons can be surface-immobilized at a liquid-solid interface of optical fiber probes through avidin-biotin binding to detect SNP in sub-micro molar level.
Another important nanostructure generated by bottomup approach is the quantum dot, a type of nanoparticle used for fluorescence tagging of probe biomolecules. Due to smaller band gap in quantum dots, quantum confinement occurs and they become photo stable. The sensitivity enhancement suggests a potential utility for detection of trace amounts of DNA [15]. In the presence of DNA-target, quantum dots serve as FRET energy donors. The result is fluorescence emission from the acceptor fluorophores by means of illumination of the quantum dot donor, thus indicating the presence of target DNA.
A common single molecule based detection system uses SiO2 surface-attached DNA polymerase, with real-time monitoring of the incorporation of fluorophore-labeled nucleotides into a growing DNA chain, by using a different fluorophore for each nucleotide [16].
Recently developed ‘lab-on-a-chip’ microsystem photolithographically integrated with specifically patterned polymeric layers and precise detection was performed for real-time and label-free detection of DNA hybridization [17]. By detecting single-molecule fluorescence with reduction in overall hybridization assay cost, the detection limit was 14 zmol. A microfluidic sensor array was developed for specific detection of ribosomal RNA-targets of several bacterial pathogens in human fluids [18]. After extraction process, RNA was detected by immunofluorescence with labeled antibody-conjugated horseradish peroxidase (HRP). It can also detect nearly 2600 cultured bacterial cells within less than five minutes with 100% sensitivity for gram-negative bacteria detection.
Electrochemical NABs
Electrochemical NAB has fast response time and as it requires simple instrumentation it can be developed at low cost. In electrochemical DNA biosensors, DNA is immobilized onto an electrode, and a change in electrical parameters (e.g., current, potential, impedance, and conductance) generated by the hybridization reaction is measured. The emergence of solid electrodes has enhanced the applicability of electrochemical methods for nucleic acid analysis. Currently, Carbon Nano Tubes (CNTs) are playing a leading role in development of electrochemical DNA based biosensors. The unique electrical, thermal, chemical, mechanical and 3D spatial properties of SWCNTs, MWCNTs have made them very popular in the biosensor community.
Label based indirect analysis: Heterocyclic dyes (e.g., methylene blue, ethidium bromide), ferrocene derivatives and organometallic complexes are widely used indicators in case of indirect analysis. Electrically stimulated hybridization indicators bind ss DNA and ds DNA with different affinities. This causes different concentration gradient around the electrode surface and variation in electrochemical response. Electrostatic attraction (to sugar-phosphate backbone), intercalation, groove-binding (within the double helix of DNA) are the major interacting forces between the DNA target and the indicators. In order to increase affinity for human immunodeficiency virus (HIV) and Hepatitis B virus HBV DNA sequences, electrochemical adsorption was done after pretreatment of sample with carbon paste and screenprinted electrodes [19,20]. The changes in output potential was measured by potentiometer using Co (phen)3 3+ as the hybridization indicator (phen=phenanthroline).
In another study, dsDNA groove-binder ferrocene was used with a chitosan coated glassy carbon electrode (GCE) to detect dengue-related oligonucleotide sequence. [21].
Polyaniline (PANI) nanotube array-immobilized DNA sensor is considered to have an immense potential to emerge as a smart sensor system. The array fabricated onto a graphite electrode, using a thin nanoporous layer as the template exhibited an ultralow detection limit of DNA (around 1 fM) and good discrimination of single nucleotide mismatches as low as 38 fM [22]. Arrays of nanotubes bound to different DNA-probes may be designed to detect specific genes for diagnostic purposes [23].
Label-free direct analysis: In label free detection, alteration in intrinsic electrical property of DNA is monitored after the occurrence of hybridization [24]. This being simple can be done rapidly avoiding other secondary processes. The difference in intrinsic DNA redox signal and guanine oxidation detects the chemical and physical damage of nucleic acids.
The surface potential difference due to hybridization of a silicon nitride gate insulator-immobilized peptide nucleic acid (PNA) with its negatively charged DNA counterpart was exploited to specifically detect the hybridization event with a FET sensor [25].
A unique label-free DNA biosensor has been recently developed on the basis of Molecular Sentinel (MS) immobilized on a plasmonic 'Nanowave' chip and named as metal film over nanosphere (MFON). The sensing principle involves decrease of the surface-enhanced Raman scattering (SERS) intensity when Raman label tagged at one end of MS is being removed from the MFON's surface due to DNA hybridization. The MFON fabrication based on a thin shell layering of gold over tightly packed arrays of nanospheres is relatively simple and cheaper with high reproducibility. The sensing process involves a single hybridization step between the target DNA sequences and the complementary MS probes. The rapid and potential application of this technique as a biosensor for medical diagnostics was demonstrated by detection of human radical S-adenosyl methionine domain containing 2 (RSAD2) gene which is a common inflammation biomarker [26].
Piezoelectric NABs
A stretch of nucleotide sequence with a few hundred base pairs possesses a measurable amount of molecular weight. The increment of mass due to hybridization of NA probe with its complimentary counterpart immobilized on the surface of a piezoelectric quartz crystal will lead to an increase in the resonance frequency of the crystal. A system comprising of DNA piezoelectric biosensor has been developed to detect a point mutation in a human gene apolipoprotein–E. The hybridization probes with complementary, non-complementary and mismatched DNA and PCR amplified samples from human blood DNA were taken as sample and the sensor was able to distinguish polymorphism [27]. Novel piezoelectric transducer based biosensor emerged when simultaneously detection and genotyping of 16 strains of the human papilloma virus was done by designing and immobilizing a degenerate probe (based on conserved genomic region) and two specific probes (based on less-conserved regions) [28]. Recently microchip based sensing technology is being developed with the help of Surface Acoustic Wave (SAW), Bulk Acoustic Wave (BAW) based piezoelectric devices apart from Quartz Crystal Microbalance (QCM) based device [29].
Clinical Applications Of NABs
Nucleic acid testing for various diseases at the point of care is being practiced in clinical settings of developed countries. Current clinical diagnostic systems are mainly PCR-based, limited only in hospitals, clinics and are still relatively complex and expensive. Integrated sample preparation with nucleic acid amplification and detection in a robust, cheaper and user-friendly manner still remain challenging. Recent NA based point-of care diagnosis tools are being advanced by novel technologies like isothermal NA amplification methods [30]. NABs have also proved promising in the detection and diagnosis of various diseases like
a) Metabolic Disorders (Cancer, diabetes, cardiovascular diseases) and
b) Infectious Diseases (Tuberculosis, hepatitis, dengue, food borne diseases like diarrhoea, cholera, Salmonellosis etc.)
Various types of NABs against several metabolic and infectious diseases are listed in Table 1.
| Target | Disease | Transducer Platform | Detection type | Detection Limit | References | |
|---|---|---|---|---|---|---|
| DNA | ||||||
| TP53 | Cancer | SPR, QCM | Label-free | 0.03–2 μM | Altintas and Tothill [39] | |
| EBNA-1 | Cancer | QCM | Label-free Amplified detection | 50 ng/ml 0.5ng/mL | Garai-Ibabe et al. [72] | |
| FGFR3, HRAS | Bladder cancer | Optical | Label free and amplified detection | 400fM | Shin et al. [41] | |
| Vibrio cholera DNA | Cholera | Glass fiber based Lateral flow | Labeled probe | 5 ng | Chua et al.[64] | |
| IS6110 gene | Tuberculosis | Piezoelectric (QCM) | Labelled probe | 10fM | Kaewphinit et al. [56] | |
| IS6110 gene | Tuberculosis | Electrochemical | Labelled probe | 0.01 ng/μl | Torres-Chavolla et al. [57] | |
| Mycobacterium tuberculosis | Tuberculosis | Electrochemical | Labeled probe | 0.065ng/μL | Das et al.[58] | |
| Hepatitis B virus DNA | Hepatitis | FRET | fluorescein (FAM)/DNA | 15 pmol/L | Lu et al. [61] | |
| invAgene of Salmonella | Salmonellosis | SPR | Label-free | 0.5 nM | Zhang et al. [65] | |
| 31-mer oligonucleotide sequence | Dengue | Electrochemical | Label-free | 2.7 × 10-12 M | Deng et al. [67] | |
| RNA | ||||||
| NS3 | Hepatitis | Optical | biotinylated RNA probes | 500pg/ml | Roh et al. [62] | |
| miRNA-21 | Cancer | Electrochemical | Direct detection | 100 aM | Hong et al. [33] | |
| miRNA-21 | Carcinoma | Electrochemical | indirect detection | 0.006 pM | Meng et al. [73] | |
| PNA | ||||||
| HER2 | Breast cancer | Optical | Direct and label-free | 0.4 to 400 fmol | Metaferia et al. [43] | |
| CF gene point mutation | CF | Optical | Label-free | 100 nM | Candiani et al. [74] | |
| M. tuberculosis DNA | Tuberculosis | SPR | Labeled probe | 1.0 ng/ml | Prabhakar et al. [55] | |
| HPV | Cervical cancer | Piezoelectric QCM | Direct detection | 1.21 pg/L | Wang et al. [45] | |
| Aptamer | ||||||
| AGR2 | Cancer | Optical | aMB probe | nanomolar | Wu et al. [36] | |
| MCF-7 cells | Breast Cancer | Electrochemical | Sandwich type | 38 cells/mL. | Yan et al. [75] | |
| Thrombin | Inflammatory disorder | Electrochemical | Labeled probe | 2.3x 10-11mol/L | Cui et al. [76] | |
| Vasopressin | Traumatic injuries | Electrochemical | label-free | 43 pM | He et al. [51] | |
| E. coli O157:H7 and Salmonella typhimurium | Diarrheal diseases | Colorimetric | Label-free | 105 cfu/ml | Wu et al. [63] | |
| RBP4 | Type 2 Diabetes | SPR | Label-free | 0.2± 0.03μM | Lee et al.[49] | |
| CRP | Cardiovascular disease | Electrochemical | Label-free | 100–500 pg/ml | Qureshi et al.[50] | |
| IFN- γ | Tuberculosis | Electrochemical, QCM | Direct | 100 Fm (RNA), 1-10 pM (DNA) depending upon buffer | Min et al. [59] | |
| Protein markers (HER2, ER, PR, ki-67) | Breast cancer | Microfluidic PDMS | Amplification based | Nanomolar level | Won et al. [37] | |
| Mucin 1 | Breast cancer | Electrochemical | Sandwich assay | As low as 100 cells | Zhu et al. [44] | |
Table 1: NABs in the diagnosis of various diseases.
Metabolic disorders
Metabolic disorders, may be inborn or inherited, are caused due to abnormality in the normal metabolic reactions in the body which may further lead to several diseases as described below-
Cancer: According to WHO, cancer is spread worldwide and its incidences are projected to climb by 50% to 15 million cases in 2020. In United States, the mortality due to cancer may mount by 580,350 cases in 2013. The early detection and diagnosis of this disease is one of the burning issues in terms of clinical practice. To address this issue, development of various sensitive, cost-effective and rapid cancer detection methods is growing. NAB technology has a great potential in the cost-effective monitoring of various types of cancer by early detection of the level of expression of various cancer biomarkers using wide array of DNA, PNA and RNA probes as the most valuable diagnostic tools [31].
The development of RNA based biosensor technology is still in its infancy. However it has gained much interest in cancer diagnosis and prognosis due to its specificity and easily accessible nature. Presently, miRNA based biosensors employing nanoparticles are emerging as potential diagnostic tools in cancer [32,33]. Biosensors exploiting aptamers as probes have also emerged as promising diagnostic candidates in various kind of cancers such as lung cancer [34], ovarian cancer [35] and several other carcinomas [36,37].
Lung cancer is the most common cancer spread worldwide. It can now also be detected using various NABs. Biosensors are developed to detect genes, biomarkers associated with lung cancer like TP53, k-ras etc. [38]. Recently a DNA biosensor has been developed to detect single nucleotide polymorphism (SNP) in TP53 gene. This system utilized synthetic DNA probe which can hybridize with its complementary mutated TP53 gene and subsequently detected by SPR and QCM analysis [39].
Several studies reveal the identification of various biomarkers for diagnosis of bladder cancer using NABs [40]. In a recent study, a highly sensitive nucleic acid (DNA) biosensor has been implemented for the early detection of biomarkers for bladder carcinoma such as FGFR3 (fibroblast growth factor receptor 3) and HRAS (Harvey RAS) in urine samples using silicon resonator as an optical sensor. This biosensor employed specific interaction between the DNA probes with their mutant FGFR3 and HRAS targets [41].
Breast cancer is one of the critical and severe cancers mostly prevalent amongst women (almost 23 % of all cancers occurring in female) with poor prognosis [42]. This type of carcinoma occurs due to malfunctioning in the expression of various hormones (HER2, ER, PR, ki-67, mucin 1 i.e. MUC1) and aberrations in the signal cascade (p53 pathway, RAS pathway etc.) [37]. Recently, PNA based biosensor has emerged as promising tool for the detection and quantification of expression of HER2 oncogene utilizing fluorescence assay.
This PNA based biosensor has immense potential to identify precisely the wild type target and the mutant with high specificity and sensitivity [43]. A novel electrochemical aptamer based biosensor has been reported to detect the breast cancer cells that over expresses MUC1 proteins. This system employed sandwich formation between MUC1 specific aptamer-cell-aptamer assays [44]. Cervical cancer, another common cancer occurring amongst women is caused by human papilloma virus (HPV). A leaky surface acoustic wave (LSAW) PNA biosensor was developed for the first time for the detection of HPV genomic DNA in the clinical samples using PNA probes with high sensitivity and selectivity [45]. The development of NABs with multimarker detection may show promising results for the rapid and cheap diagnosis of cancer.
Diabetes mellitus: Diabetes mellitus is the most common metabolic disorder due to abnormal levels of blood glucose. The detection and maintenance of the blood glucose level is therefore necessary for the proper diagnosis and control of diabetes. The development of biosensor technology for glucose detection has contributed almost about 85% of the world’s biosensor market [46]. Till date, few NABs have been reported for detection of biomarkers and miRNA level associated with diabetes. DNA based personal glucose meters (PGMs) have been reported for monitoring the blood glucose level. Instead of PCR, PGM can detect upto 40 pM DNA based on enzymatic turnovers. The quantification is based on target-dependent binding of cDNA-invertase conjugated with the analyte DNA [47]. The aberration in the level of miRNA is also associated with diabetes. Recently, a PNA based biosensor has been developed to quantify the level of miRNA expression. This system utilized nano-sized graphene-oxide which acts as a quencher and quenches the fluorescence of PNA probes. The fluorescence is recovered by subsequent binding of PNA and target miRNA [48]. Type 2 Diabetes is a widespread metabolic disorder which causes insulin resistance due to over-expression of its biomarkers (RBP4). An aptamer based biosensor was developed for the rapid detection of level of Retinol binding protein 4 (RBP4) in serum samples. The developed biosensor utilized the immobilization of RBP4-specific aptamers on the Au chip and subsequent detection by SPR detection method [49]. There is an increasing effort made in the development of simple, rapid, cheap and highly sensitive nucleic acid based biosensors for the diagnosis of diabetes.
Cardiovascular diseases (CVD): Cardiovascular disease is another commonly occurring disease with growing incidences and therefore detection of the cardiovascular biomarkers is necessary in clinical perspectives. In a study, RNA based aptasensor employed the charge distribution phenomenon exhibited by synthetic aptamer-CRP complex on the gold interdigitated (GID) capacitor under electric field. It can detect C-reactive protein (CRP), the most common biomarker for CVD with a detection limit of 100–500 pg/ml [50]. Recently, small and easily portable aptamer based electrochemical biosensor was developed for the detection of vasopressin, a biomarker for traumatic injuries. The aptamers were immobilized on carbon nanotubes (CNTs) and the conductivity change was measured upon specific aptamer-vasopression conjugation [51].
Systemic lupus erythematosus (SLE): SLE is an auto-immune disorder which affects different parts of human body. The patients suffering from SLE develop a wide variety of serologic manifestations, including double-stranded DNA autoantibodies (anti-dsDNA). SPR based biosensor chip was developed to detect pathogenic dsDNA in case of autoimmune disorder SLE [52].
Infectious diseases
Globally infectious diseases are the most prevalent and wide spread diseases caused by a wide array of infectious agents. Several NA based biosensing system to combat against infectious diseases are described below.
Tuberculosis: Tuberculosis, one of the world’s lethal diseases is caused by pathogenic bacterium M. tuberculosis [53]. Rapid diagnosis and management of this infectious agent is therefore crucial for clinical perspectives. A wide variety of DNA [54], RNA and PNA [55] biosensors were developed against tuberculosis biomarkers based on optical, piezoelectric, electrochemical transduction systems.
The QCM based piezo-electric biosensor was developed to detect tuberculosis biomarker, IS6110 [56]. Dextrin coated GNP conjugated electrochemical DNA biosensor was developed for diagnosis of M. tuberculosis. This biosensor utilizes the hybridization between AuNPs, magnetic particles (MPs) each functionalized with DNA probes and tuberculosis specific DNA fragment within IS6110 gene [57]. In another study, ZrO2-based electrochemical DNA biosensor was demonstrated for the rapid, early and sensitive detection of M. tuberculosis [58].
SPR technique was employed for tuberculosis detection using thiolated DNA and PNA probes immobilized on the gold electrode surface. PNA probes showed enhanced sensitivity and improved detection limit in comparison to DNA probes in M. tuberculosis infected clinical samples [55]. Apart from DNA, PNA and RNA based biosensors, aptamer based biosensors are also evolving as faster, highly sensitive and selective diagnostic tools for tuberculosis detection [59]. Recently, aptamer based biosensor chip has been developed which utilizes the aptamer integration with CNTs for early diagnosis of MTB [60].
Hepatitis: Hepatitis is one of the infectious inflammatory health disorders caused by various strains of hepatitis virus (HAV, HBV, HCV, HDV, HEV, HFV and HGV). Recently, the utility of gold nanoparticle based DNA biosensor has been implemented to monitor hepatitis B virus DNA with a detection limit of 15 pmol/L [61]. In another study, non-structural viral protein 3 (NS3) has been detected by biotinylated RNA probe based biosensor with a detection limit of 500 pg/ml [62].
Food-borne diseases: Food-borne diseases are the most common infectious diseases caused by various pathogenic bacteria such as E. coli and S. typhimurium, Vibrio cholera, Listeria monocytogenes, Vibrio parahaemolyticus etc. Diarrhoea is one such disease caused by pathogenic bacteria such as E. coli O157:H7, S. typhimurium and S. paratyphimurium. Aptamer based biosensor was reported for the detection of these bacteria using unmodified gold nanoparticles by colorimetric assay [63]. Cholera caused by Vibrio cholerae is another food borne disease. A DNA biosensor was developed for the detection of PCR amplicons of Vibrio cholera [64].
Salmonellosis is a globally widespread disease caused by Salmonella sp. Recently use of DNA biosensors have been reported for the detection of the invA gene of Salmonella using SPR detection method [65].
Dengue: Dengue fever is one of the vector-borne infectious and endemic diseases caused by Dengue viruses (DENV1-4). Due to high surface area nanoporous alumina membrane-based nucleic acid has been employed for diagnosis of Dengue fever. A nanoporous alumina membrane based electrochemical-DNA biosensor has been developed which can detect the cDNA sequence in Dengue virus [66]. Recently, detection of 31-mer oligonucleotide sequence of Dengue virus has been reported using electrochemical DNA sensing technology. It employed cross linking of DNA probes on the surface of nanoporous alumina membrane and subsequent measurement of impedance changes that occur during probe-target DNA hybridization [67].
Commercialization Of NABs
In the past few decades, progress has been made in the development of biosensor technology in clinical perspectives. According to Global Strategic Business Report on Biosensors in Medical Diagnostics, glucose biosensors contribute largest share of the total biosensors market. However, only few biosensors have reached the level of commercialization propelling the biosensor technology towards the diagnostic market. According to Global Industry Analysts, Inc. (GIA) report, the biosensor market may rise to US$12 billion by the year 2015. Due to high sensitivity and selectivity, NABs are gaining commercially much attention as powerful and inexpensive clinical diagnostics [68].
Affymetrix (Santa Clara, CA) has developed GeneChip™, a commercial biosensor microarray for fluorescent detection of single nucleotide polymorphisms (SNPs) for diagnosis of diabetes and various infectious diseases [69]. Genelyzer™, is another electrochemical DNA chip first developed by Toshiba and then commercialized by Antara BioSciences Inc. (USA). The developed product utilizes Hoechst 33258 dye as a minor groove binder to detect viral and bacterial strains in clinical samples [70].
With the advent of nucleic acid-based biosensor technology, various commercialized products have been developed for detection of infectious pathogens in clinical samples. [71]. Dupont had developed a product named BAX® for detection of numerous infectious agents like E. coli O157:H7, Litseria, Enterobacter, Campylobacter, Vibrio and Salmonella based on DNA hybridization. Other DNA based biosensor products `ACCUPROBE´ and ANSR™ were developed by Biomėrieux and Neogen, respectively for monitoring of Campylobacter, Salmonella and Listeria in clinical samples.
Various RNA based commercial products have also been developed for the detection of food-borne pathogens. Some species of Mycobacterium including Mycobacterium avium, Mycobacterium intracellulare, Mycobacterium gordonae, Mycobacterium kansasii can be detected by a product Gen probe® developed by Accuprobe. APTIMA®, APTIMA®GC, APTIMA®CT are some other products developed by Gen-probe for the detection of Trichomonas vaginalis, Neisseria gonorrhoeae and Chlamydia trachomatis, respectively. DNA hybridization based product BD Affirm™ APIII has been developed for the detection of Gardnerella, Trichomonas vaginalis, Candida sp by Beckton Dickinson, Inc.
The high stability and specificity of nucleic acid based biosensors can be promising candidates in the future for clinical diagnostic market. However, the major bottleneck is the requirement of strategic investments for commercialization of biosensor technology in clinical field.
To conclude, in near future, the advanced level of medical diagnosis will be essentially dependent on the state-of-the-art biosensors. Huge progress is noticed in electrochemical DNA based biosensors and arrays. Various types of electrodes immobilized with specific probes are at various stages of development, fabrication and testing for the diagnosis of fatal diseases. Carbon nanotubes based on electrochemical biosensor, SPR, Quantum Dot and Piezoelectric biosensors are the promising candidates in molecular diagnosis. The use of DNA biochip technology would eliminate the role of PCR in future. New fabrication methods are emerging to develop fiber optic DNA biosensors.
Ongoing efforts with DNA chips and other smart nanomaterials however are directed toward developing PCR free DNA detection systems which are yet to be fully commercialized.
Acknowledgements
The authors thank the Department of Biotechnology, Govt. of India for supporting the research and development activities through the Institutional Biotech Hub project (Project No BT/04/NE/2009). AS and DS thank IITG and MHRD for the financial support in the form of fellowship.
References
- Barthelmebs L, Calas-Blanchard C, Istamboulie G, Marty JL, Noguer T (2010) Biosensors as analytical tools in food fermentation industry. Adv Exp Med Biol 698: 293-307.
- Turner APF, KaubeI, Wilson GS (1987) Biosensors Fundamentals and Applications. Oxford University Press 86–92.
- Wang J, Rivas G, Cai X, Palecek E, Nielsen PE et al (1997a) DNA electrochemical biosensors for environmental monitoring: A review. Anal Chim Acta 347: 1–8.
- Dell’Atti D, Tombelli S, Minunni M, Mascini M (2006) Detection of clinically relevant point mutations by a novel piezoelectric biosensor. Biosensors and Bioelectronics 21: 1876-1879.
- Mikkelsen SR (1996) Electrochemical biosensors for DNA sequence detection. Electroanalysis 8: 15.
- Junhui Z, Hong C, Ruifu Y (1997) DNA based biosensors. Biotechnol Adv 15: 43-58.
- Wang J (2002) Electrochemical nucleic acid biosensors. Anal Chim Acta 469: 63-71.
- Wang J (2000) From DNA biosensors to gene chips. Nucleic Acid research 28(16): 3011-16.
- Herne TM, Tarlov MJ (1997) Characterization of DNA probes immobilized on gold surfaces. J Am Chem Soc 119: 8916–8920.
- Sperling RA, PellegrinoT, Li JK , Chang WH, Parak WJ (2006) Electrophoretic separation of nanoparticles with a discrete number of functional groups. Adv Func Mater 16: 943–948.
- Ray A, Norden B (2000) Peptide nucleic acid (PNA): its medical and biotechnical applications and promise for the future. FASEB J 14: 1041–1060.
- Hamidi-Asl E, Raoof JB, Ojani R, Hejazi MS (2013) Indigo Carmine as New Label in PNA Biosensor for Detection of Short Sequence of p53 Tumor Suppressor Gene. Electroanalysis.
- Sett A, Das S, Sharma P, Bora U (2012) Aptasensors in Health, Environment and Food Safety Monitoring. Open Journal of Applied Biosensor 1: 9-19.
- Piunno PAE, Krull UJ, Hudson RHE, Damha MJ, Cohen H (1995) Fiber-optic DNA sensor for fluorometric nucleic acid determination. Anal Chem 67: 2635-43.
- Alivisatos P (2004) The use of nanocrystals in biological detection. Nature Biotechnology 22 (1): 47-52.
- Korlach J, Levene M, Turner SW, Craighead HG, Webb WW (2002) Single molecule analysis of DNA polymerase activity using zero-mode waveguides. Biophysical J. 82:507A–507A.
- Sepúlveda JS, Del RÃo M, Moreno FJ, Blanco K, Mayora C, et al (2006) Optical biosensor micro-systems based on the integration of highly sensitive Mach-Zehnder interferometer devices. Opt A: Pure Appl Opt 8 : S561-66.
- Liao JC, Mastali M, Gau V, Suchard MA and Moller AK, et al. (2006) J Clin Microbiol 44: 561.
- Wang J, Cai X, Rivas, Shiraishi H, Farias PAM, et al (1996) Anal. Chem. 68: 2629-2634.
- Erdem A, Kerman K, Meric B, Akarca US, Ozsoz M (1999) DNA electrochemical biosensor for the detection of short DNA sequences related to the Hepatitis B virus. Electroanalysis 11: 586-587.
- Chang H, Yuan Y, Shi N, Guan Y (2007) Electrochemical DNA biosensor based on conducting polyaniline nanotube array. Anal Chem 79(13): 5111-5.
- Ngo HT, Wang HN, Fales AM, Vo-Dinh T (2013) Label-free DNA biosensor based on SERS Molecular Sentinel on Nano wavechip. Anal Chem 85(13): 6378-6383.
- Tombelli S, Mascini M, Braccini L, Anichini M and Turner APF (2000) Coupling of a DNA piezoelectric biosensor and polymerase chain reaction to detect apolipoprotein-E polymorphisms. Biosensor and Bioelectronics 15: 363-370.
- Dell’Atti D, Zavaglia M, Tombelli S, BertaccaG and Cavazzana AO, et al. (2007) Development of combined DNA-based piezoelectric biosensors for the simultaneous detection and genotyping of high risk Human Papilloma Virus strains. Clinica Chimica Acta 383: 140–146.
- Sakong J, Roh H, Roh Y (2007) Surface Acoustic Wave DNA Sensor with Micro-Fluidic Channels. Jpn J Appl Phys 46: 4729-4733.
- Niemz A, Ferguson TM, Boyle DS.(2011) Point-of-care nucleic acid testing for infectious diseases.Trends Biotechnol29(5):240-50.
- Bohunicky B, Mousa SA (2011) Biosensors: the new wave in cancer diagnosis. Nanotechnology, Science and Applications 4:1-10.
- Catuogno S, Esposito CL, Quintavalle C, Cerchia L, Condorelli G, et al (2011) Recent Advance in Biosensors for microRNAs Detection in Cancer. Cancers3: 1877-1898.
- Hong CY, Chen X, Liu T, JuanLi andYangHH, et al. (2013) Ultrasensitive electrochemical detection of cancer-associated circulating microRNA in serum samples based on DNA concatamers. Biosensors andBioelectronics50:132–136.
- Zhang Y, Yang D, Weng L, Wang L (2013) Early lung cancer diagnosis by biosensors. Int J MolSci 14(8): 15479-509.
- Lin M-Y, Lu Y-P, Grumezescu AM, Ho FH, Kao Y-H, et al (2013). Tumor Marker Detection by Aptamer-Functionalized Graphene Oxide.Current Organic Chemistry 17:132-136.
- Wu J, Wang C, Li X, Song Y and Wang W, et al. (2012) Identification, characterization and application of a G-quadruplex structured DNA aptamer against cancer biomarker protein anterior gradient homolog 2. PLoS One 7(9):e46393.
- Won JY, ChoiJW, MinJ (2013) Micro-fluidic chip platform for the characterization of breast cancer cells using aptamer-assisted immunohistochemistry. Biosensors and Bioelectronics 40(1): 161-166.
- Altintas Z, Tothill I (2013) Biomarkers and biosensors for the early diagnosis of lung cancer. Sensors and Actuators B: Chemical 188: 988-998.
- Altintas Z, Tothill IE (2012) DNA-based biosensor platforms for the detection of TP53 mutation. Sensors and Actuators B: Chemical 169: 188-194.
- Proctor I, Stoeber K, Williams GH (2010) Biomarkers in bladder cancer. Histopathology57: 1–13.
- Shin Y, PereraAP, ParkMK (2013) Label-free DNA sensor for detection of bladder cancer biomarkers in urine. Sensors and Actuators B: Chemical 178: 200-206.
- Boyle P, Levin B (2008) World Cancer Report, International Agency for Research on Cancer.
- Metaferia B, Wei JS, Song YK, Evangelista J and Aschenbach K, et al. (2013) Development of Peptide Nucleic Acid Probes for Detection of the HER2 Oncogene. PLoS ONE 8(4): e58870.
- Zhu X, Yang J, Liu M, Wu Y, Shen Z, et al. (2013) Sensitive detection of human breast cancer cells based on aptamer–cell–aptamer sandwich architecture.AnalyticaChimicaActa764: 59-63.
- Wang Y, Chen M, Zhang L, Ding Y and Luo Y, et al. (2009) Rapid detection of human papilloma virus using a novel leaky surface acoustic wave peptide nucleic acid biosensor. Biosensors and Bioelectronics 24(12): 3455-3460.
- Yoo E-H, Lee S-Y (2010) Glucose Biosensors: An Overview of Use in Clinical Practice.Sensors (Basel) 10(5): 4558–4576.
- Xiang Y, Lu Y (2012) Using commercially available personal glucose meters for portable quantification of DNA. Anal Chem 84(4): 1975–1980.
- Ryoo S-R, Lee J, Yeo J, Na H-K, Kim Y-K, et al (2013) Quantitative and Multiplexed MicroRNA Sensing in Living Cells Based on Peptide Nucleic Acid and Nano Graphene Oxide (PANGO)7(7): 5882–5891.
- Lee SJ, Youn BS, Park JW, Niazi JH, Kim YS, et al (2008) ssDNA aptamer-based surface plasmon resonance biosensor for the detection of retinol binding protein 4 for the early diagnosis of type 2 diabetes. Anal Chem 80(8): 2867-2873.
- Qureshi A, Gurbuz Y, Kallempudi S, Niazi JH (2010) Label-free RNA aptamer-based capacitive biosensor for the detection of C-reactive protein. Phys Chem 12(32): 9176-82.
- He P, Oncescu V, Lee S, Choi I, Erickson D (2013) Label-free electrochemical monitoring of vasopressin in aptamer-based microfluidic biosensors. Anal Chim Acta 759:74-80.
- Buhl A, Metzger JH, Heegaard NHH, Landenberg von P and Fleck M, et al. (2007) Novel Biosensor based analytic device for the detection of anti–double-stranded DNA antibodies. Clin Chem 53: 334-41.
- WHO Global Tuberculosis Control, 2009.Epidemiology, Strategy, and Financing.
- Zhou L, HeX, HeD, Wang K, Qin D (2011) Biosensing Technologies for Mycobacterium tuberculosis Detection: Status and New Developments. Clinical and Developmental Immunology 2011: 8.
- Prabhakar N, Arora K, Arya SK, Solanki PR and Iwamoto M, et al. (2008)Nucleic acid sensor for M. tuberculosis detection based on surface plasmon resonance. Analyst 133(11): 1587-92.
- Kaewphinit T, Santiwatanakul S, ChansiriCPK (2010) Detection of Non-Amplified Mycobacterium tuberculosis Genomic DNA Using Piezoelectric DNA-Based Biosensors. Sensors 10: 1846-1858.
- Torres-Chavolla E, Alocilja EC (2011) Nanoparticle based DNA biosensor for tuberculosis detection using thermophilic helicase-dependent isothermal amplification. BiosensBioelectron 26(11):4614-8.
- Das M, Sumana G, Nagarajan R, Malhotra BD (2010) Zirconia based nucleic acid sensor for Mycobacterium tuberculosis detection. ApplPhysLett 96: 133703.
- Min K, Cho M, Han SY, Shim YB and Ku J, et al. (2008) A simple and direct electrochemical detection of interferon-γ using its RNA and DNA aptamers. Biosensors and Bioelectronics 23(12): 1819-1824.
- Wang H-W, Huang J-T, Lin C-C (2013) Real-Time Detecting Concentration of Mycobacterium tuberculosis by CNTFET Biosensor. World Academy of Science, Engineering and Technology 79.
- Lu X, Dong X, Zhang K, Han X and Fang X, et al. (2013) A gold nanorods-based fluorescent biosensor for the detection of hepatitis B virus DNA based on fluorescence resonance energy transfer. Analyst 138(2):642-50.
- Roh C, Kim SE, Jo SK (2012) A Simple and Rapid Detection of Viral Protein Using RNA Oligonucleotide in a Biosensor. Journal of Analytical Chemistry 67(11): 925–929.
- Wu WH, Li M, Wang Y, Ouyang HX and Wang L, et al. (2012) Aptasensors for rapid detection of Escherichia coli O157:H7 and Salmonella typhimurium. Nanoscale Res Lett 7(1):658.
- Chua A, Yean CY, Ravichandran M, Lim B and Lalitha P (2011) A rapid DNA biosensor for the molecular diagnosis of infectious disease. Biosens Bioelectronics 26: 3825–3831.
- Zhang D, Yan Y, Li Q, Yu T and Cheng W, et al. (2012) Label-free and high-sensitive detection of Salmonella using a surface plasmon resonance DNA-based biosensor. Journal of Biotechnology 160(3–4): 123-128.
- Rai V, Hapuarachchi HC, Ng LC, Soh SH and Leo YS, et al. (2012) Ultrasensitive cDNA Detection of Dengue Virus RNA Using Electrochemical Nanoporous Membrane-Based Biosensor. PLoS ONE 7(8): e42346.
- Deng J, Toh CS (2013) Impedimetric DNA Biosensor Based on a Nanoporous Alumina Membrane for the Detection of the Specific Oligonucleotide Sequence of Dengue Virus. Sensors 13: 7774-7785.
- Teles FRR, Fonseca LP (2008) Trends in DNA biosensors.Talanta 77: 606-623.
- D'Orazio P (2011) Biosensors in clinical chemistry- 2011 update. Clin Chim Acta 412(19-20): 1749–1761.
- Lucarelli F, Tombellia S, Minunnia M, Marrazzaa G and Mascinia M (2008) Electrochemical and piezoelectric DNA biosensors for hybridisation detection.Anal ChimActa 609(2):139-59.
- Singh A, Poshtiban S, Evoy S (2013)Recent Advances in Bacteriophage Based Biosensors for Food-Borne Pathogen Detection. Sensors 13: 1763-1786.
- Garai-Ibabe G, Grinyte R, Golub EI, Canaan A and de la Chapelle ML, et al. (2011) Label free and amplified detection of cancer marker EBNA-1 by DNA probe based biosensors. Biosensors and Bioelectronics 30: 272–275.
- Meng X, Zhou Y, Liang Q, Qu X, Yang Q, et al (2013)Electrochemical determination of microRNA-21 based on bio bar code and hemin/G-quadruplet DNAenzyme. Analyst 138: 3409-3415.
- Candiani A, Bertucci A, Giannetti S, Konstantaki M, Manicardi A, et al (2013) Label-free DNA biosensor based on a peptide nucleic acid-functionalized microstructured optical fiber-Bragg grating. J Biomed Opt 18(5):57004.
- Yan M, Sun G, Liu F, Lu J andYu J, et al. (2013) An aptasensor for sensitive detection of human breast cancer cells byusing porous GO/Au composites and porous PtFe alloy as effectivesensing platform and signal amplification labels. Analytica Chimica Acta 798: 33-39.
- Cui HF, Fan H, Lin Y (2013) A New Electrochemical Aptasensor for Protein Detection Based on Target Protein-Induced Strand Displacement. Advanced Materials Research 753-755: 2113-2116.
Citation: Bora U, Sett A, Singh D (2013) Nucleic Acid Based Biosensors for Clinical Applications. Biosens J 2:104. DOI: 10.4172/2090-4967.1000104
Copyright: © 2013 Bora U, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Select your language of interest to view the total content in your interested language
Share This Article
Recommended Journals
Open Access Journals
Article Tools
Article Usage
- Total views: 27497
- [From(publication date): 12-2013 - Dec 05, 2025]
- Breakdown by view type
- HTML page views: 22317
- PDF downloads: 5180