XB130 Expression in Human Osteosarcoma: A Clinical and Experimental Study
Received: 26-Feb-2014 / Accepted Date: 17-Apr-2014 / Published Date: 19-Apr-2014 DOI: 10.4172/2161-0681.1000169
Abstract
Identifying prognostic factors for osteosarcoma (OS) aids in the selection of patients who require more aggressive management. XB130 is a newly characterized adaptor protein that was reported to be to be a prognostic factor of certain tumor types. However, the association between XB130 expression and the prognosis of OS remains unknown. In the present study, we investigated the association between XB130 expression and clinicopathologic features and prognosis in patients suffering osteosarcoma, and further investigated its potential role on OS cells in vitro and vivo. A retrospective immunohistochemical study of XB130 was performed on archival formalin-fixed paraffin-embedded specimens from 60 pairs of osteosarcoma and noncancerous bone tissues, and compared the expression of XB130 with clinicopathological parameters. We then investigate the effect of XB130 silencing on invasion in vitro and lung metastasis in vivo of the human osteosarcoma cell line. Immunohistochemical assays revealed that XB130 expression in osteosarcoma tissues was significantly higher than that in corresponding noncancerous bone tissues (P=0.001). In addition, high XB130 expression more frequently occurred in osteosarcoma tissues with advanced clinical stage (P=0.002) and positive distant metastasis (P=0.001). Moreover, osteosarcoma patients with high XB130 expression had significantly shorter overall survival and disease-free survival (both P<0.001) when compared with patients with the low expression of XB130. The univariate analysis and multivariate analysis shown that high XB130 expression and distant metastasis were the independent poor prognostic factor. We showed that XB130 depletion by RNA interference inhibited invasion of XB130-rich U2OS cells in vitro and lung metastasis in vivo. This is the first study to reveal that XB130 overexpression may be related to the prediction of metastasis potency and poor prognosis for osteosarcoma patients, suggesting that XB130 may serve as a prognostic marker for the optimization of clinical treatments. Furthermore, XB130 is the potential molecular target for osteosarcoma therapy.
Keywords: Osteosarcoma; XB130; Immunohistochemistry; Prognosis; RNA interference
313828Introduction
Osteosarcoma is a type of aggressive bone cancer of mesenchymal origin generally found in youths aged between 10 and 25 years. Any bone of the human body can be affected by this neoplasia and the general survival at five years is approximately 65 to 75% [1]. The main causes of death are pulmonary metastases diagnosed by computed tomography (CT) in 35 to 45% of the patients [2]. Even though effective chemotherapy for patients with OS has indeed led to a significant improvement of clinical outcome, between 30 and 50% of patients with non-metastatic disease of extremities still die from this neoplasia, in spite of having had a complete surgical removal of the tumor and intensive chemotherapy [3,4]. Currently, many immunohistochemical markers and genetic proteins have been studied, but with prognostic and therapeutic relevance doubtful [5-9]. Thus, it is crucial to identify new and efficient biomarkers which can provide insight into tumor progression and outcome, especially in screening poor prognosis patients who should be offered more aggressive therapy at an early time point in the clinical treatment.
XB130 is a newly discovered adaptor protein for intracellular signal transduction; it is involved in gene regulation, cell proliferation, cell survival, cell migration, and tumorigenesis [10]. XB130 has been detected in human esophageal squamous cell carcinoma (ESCC) [11], follicular and papillary thyroid carcinoma, as well as in human lung carcinoma cell lines [12]. In ESCC cells, expression of XB130 may affect cell cycle progression and impact prognosis of patients with ESCC [11]. In thyroid and lung cancer cells, XB130 has been implicated as a substrate and regulator of tyrosine kinase-mediated signaling and in controlling cell proliferation and apoptosis [12]. In the gastric cancer, reduced XB130 protein expression is a prognostic biomarker for shorter survival and a higher recurrence rate in patients with GC, as well as for the response to chemotherapy [13]. However, in patients with hepatocellular carcinoma (HCC), protein expression of XB130 is not associated with the postoperative prognosis of patients with HCC [14]. However, clinical evidence is not yet well established between expression of XB130 and clinical significance in osteosarcoma patients.
To date, the association between XB130 expression and prognosis of OS remains unknown. In this study, we analyzed the association of XB130 expression in OS with clinical factors and overall survival, and further investigated its potential role on cell proliferation, invasion and lung metastasis in vitro and vivo.
Materials and Methods
Cell lines and culture conditions
OS cell lines (U2OS) were purchased from the American Tissue Culture Collection (ATCC). The cells were cultured in DMEM (Life Technologies, Inc., San Diego, CA) supplemented with 10% FCS, 2 mM glutamine, and 1% nonessential amino acids (complete medium).
Patients and tissue samples
This study was approved by the Research Ethics Committee of Yishui central hospital, China. All specimens were handled and made anonymous according to the ethical and legal standards. Paraffin-embedded OS sections from 60 patients who were diagnosed with primary OS and had undergone initial surgery at the Yishui central Hospital, Yishui, China, between January 2008 and December 2012 were obtained from the Department of Orthopedics for immunohistochemical staining. No patients had received blood transfusion, radiotherapy, or chemotherapy before surgery. Clinical stage of these osteosarcoma patients was classified according to the sixth edition of the tumor–node–metastases (TNM) classification of the International Union Against Cancer (UICC) [15] .Overall survival was calculated as the time from the date of diagnosis to the date of death or the date of last follow-up if the patient was still surviving. The clinicopathological information of the patients is shown in (Table 1).
| XB130 expression | ||||
|---|---|---|---|---|
| Clinicopathological features | No. of cases | Positive(+) | Negative(-) | p-value |
| Tissue | 0.0001 | |||
| Normal tissue | 60 | 0(100%) | 60(100%) | |
| Osteosarcoma tissue | 60 | 24(40%) | 36(60%) | |
| Age(year) | 0.327 | |||
| >20 | 19 | 7(36.8%) | 12(63.2%) | |
| ≤20 | 41 | 17(41.4%) | 24(58.6%) | |
| Gender | 0.486 | |||
| Female | 22 | 8(36.4%) | 14(63.6%) | |
| Male | 38 | 16(42.1%) | 22(57.9%) | |
| Tumor metastasis | 0.001 | |||
| Yes | 19 | 16(84.2%) | 3(15.8%) | |
| No | 41 | 8(19.5%) | 33(80.5%) | |
| Size of primary OS(cm) | 0.187 | |||
| <8 | 28 | 9(32.1%) | 19(67.9%) | |
| ≥8 | 32 | 15(46.9%) | 17(53.1%) | |
| Clinical stage | 0.002 | |||
| Ⅰ/ⅡA | 23 | 4(17.4%) | 19(82.6%) | |
| ⅡB/Ⅲ | 37 | 20(54%) | 17(46%) | |
| Histologic type | 0.632 | |||
| Osteoblastic | 30 | 12(40%) | 18(60%) | |
| Chondroblastic | 11 | 5(45.4%) | 6(54.6%) | |
| Fibroblastic | 12 | 5(41.7%) | 7(58.3%) | |
| Mixed | 6 | 2(33.3%) | 4(66.7%) | |
| Location | 0.894 | |||
| Femur | 25 | 11(44%) | 14(56%) | |
| Tibia | 18 | 6(33.3%) | 12(66.7%) | |
| Humurs | 9 | 4(44.4%) | 5(55.6%) | |
| Others | 8 | 3(37.5%) | 5(62.5%) | |
| Response to chemotherapy | 0.086 | |||
| Good | 21 | 7(33.3%) | 14(66.7%) | |
| Poor | 39 | 17(43.6%) | 22(56.4%) | |
Table 1: Significance of the variables compared with the expression of XB130
Immunohistochemistry
Immunohistochemical staining was carried out using the Dako Envision System (Dako, Carpinteria, CA) and rabbit polyclonal anti-XB130 antibody (1:100; Ori Gene) following the manufacturer’s recommended protocol.The staining intensity and extent scores was according to the previous report [13]. Briefly, the staining intensity was scored as 0 (negative), 1 (weak), 2 (medium), or 3(strong). The extent of staining was scored as 0 (0%), 1 (1%–25%), 2(26%–50%), 3 (51%–75%), or 4 (76%–100%), according to the percentages of positively stained areas in relation to the whole carcinoma area (or entire section for normal samples). The sum of the staining intensity and extent scores was used as the final staining scores (0-7) for XB130. Tumors having a final staining score of = 3 were considered to be positive. XB130 immunostaining was evaluated independently by two individuals blinded to the clinical parameters.
XB130 short hairpin RNA stable clones
The Sure Silencing shRNA plasmid kit for human XB130 was purchased from Super Array (Frederick, MD). Four shRNA plasmids for XB130 were included in the kit. Negative control shRNA plasmid was a scrambled artificial sequence that does not match any human gene.When 50% to 60% confluence, U2OS cells were transfected with the shRNA plasmids (U2OS /XB130 shRNA)using the Lipofectamine 2000 according to the manufacturer's protocol. Transfected cells were selected with 1.0 mg/mL G418 (InvivoGen) for 14 days, and antibiotic-resistant pools were collected. To screen out the best gene ablation efficiency, the expression levels of XB130 protein from antibiotic-resistant pools were determined by Western blotting. To establish single-cell colonies, cell pools were diluted to obtain 1 cell/200 uL and plated onto 96-well plates. Cell clones were expanded under continuous selection using 1 mg/ml G418. Single colonies were chosen by western blotting and used for further experiments.
Western blotting
Proteins in the total cell lysate (40 µg of protein) were separated on 10% SDS-PAGE and electro transferred to a poly vinyl idenedi fluoride membrane (Immobilon-P membrane; Millipore, Bedford, MA). After the blot was blocked in a solution of 5% skimmed milk, 0.1% Tween 20 and PBS, membrane-bound proteins were probed with primary antibodies against XB-130. The membrane was washed and then incubated with horseradish peroxidase-conjugated secondary antibodies for 30 minutes. Antibody-bound protein bands were detected with enhanced chemiluminescence reagents (Amersham Pharmacia Biotech, Piscataway, NJ) and photographed with Kodak X-Omat Blue autoradiography film (Perkin Elmer Life Sciences, Boston, MA).
Cell proliferation assay in vitro
U2OS, U2OS /XB130 shRNA and U2OS / shRNA cells were seeded in 24-well plates at a density of 1 × 104 viable cells per well. The cells were grown for up to 96 h, detached in trypsin-EDTA, and counted using a cell counter (Cedex Hi Res, Innovatis, Germany). Cell numbers were averaged and results expressed as percent of control.
Motility and invasion assays
The motile and invasive abilities of the cell lines through a filter containing 8 um pores were measured using a Boyden chamber in a 24-well plate assay system (Corning Costar, High Wycombe, Bucks, UK). Chemotactic-induced motility in response to a FCS concentration gradient was measured by adding 400 ul of Routine Medium containing 10% (v v-1) FCS to the lower compartment, and 2×105 cells (U2OS, U2OS /XB130shRNA and U2OS / shRNA)in 200 ml of Routine Medium, but with 2% (v v-1) FCS to the upper compartment of the Boyden chamber. The cells were incubated for 24 h, the upper side of the filter was wiped with a cotton swab to remove any non-motile cells, and the motile cells on the lower side of the filter were fixed and stained using the Diffquik histochemical stain (Dade Behring, Du¨dingen, Switzerland) according to the manufacturer’s instructions. The lower compartment of the Boyden chamber was checked for cells and the number of stained cells/field of 0.50mm2on the lower side of the filter was counted using a Dynascope with a ×20 objective (Vision Engineering, Working, Surrey, UK). For each cell line, four experiments were carried out, each experiment consisting of three filters and 10 fields per filter were counted. In controls, fixed numbers of cells incubated for different times up to 24 h showed a linear increase in the number of motile cells on the lower side of the filter. Thereafter, there were an appreciable number of cells present in the lower chamber. For cell invasion, the filters were coated with 50 mg Matrigel (Collaborative Biomedical Products, Becton Dickinson, Oxford, UK) and the experiments were carried out as for the chemotactic motility assays. The number ofinvasive cells/field on the lower side of the filters was determined as for the motility assays for three experiments for each cell line.
In vivo tumor model
4-6 week-old male nude mice, bred and maintained in our institutional specific pathogen-free mouse colony, were used. Briefly, 4 week-old male nude mice were injected s.c. with 2.5 × 106 U2OS cells, XB130 shRNA-transfected U2OS cells and shRNA-transfected U2OS cells, respectively, in 100 uL of PBS at a single dorsal site. Tumor size was measured every 3 days for 21 days. Tumor growth was quantified by measuring the tumors in three dimensions with calipers. The results were expressed as the mean tumor volume (n=6) with 95% confidence intervals. The statistical significance of differences in tumor growth was analyzed using Wilcoxon rank sum test.
For lung metastasis model, U2OS cells, XB130 shRNA-transfected U2OS cells and shRNA-transfected U2OS cells (1×106 cells in 300 µl of culture medium without serum) were injected intravenously through the tail vein. After 5 weeks, the mice were sacrificed, their lungs harvested, and the number of macroscopic surface tumor nodules counted. Ten mice were used in each group.
Statistical Analysis
The results were expressed as mean ± SD. The SP11.0 software was used to analyze the related data with a x2 test (chi-square test) or t test. Kaplan–Meier method and log-rank test were used for survival analysis. Survival correlation with the prognostic factors was further investigated by multivariate analysis using the Cox proportional hazards model with backward stepwise likelihood ratio. Statistical analysis was performed using SPSS11.0 statistical software (Chicago, IL, USA). Statistical significance was assumed for P<0.05.
Results
Overexpression of XB130 in osteosarcoma tissues
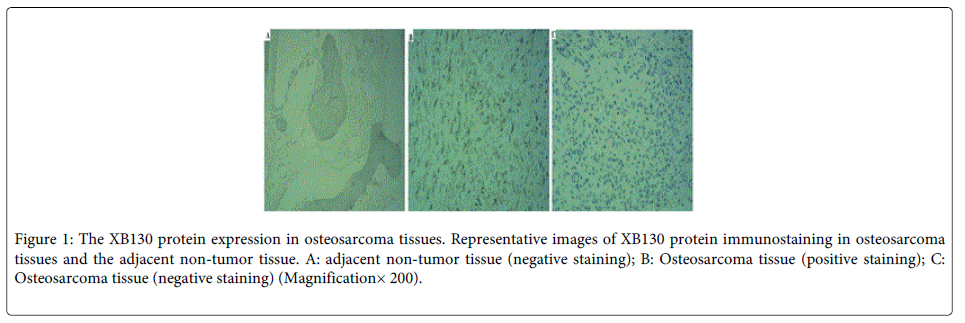
Immunohistochemistry was performed to evaluate XB130 expression in all 60 pairs of paraffin-embedded sections from OS and their adjacent non-tumor tissues. As shown in (Figure 1), XB130 did not express in the adjacent non-tumor tissue (Figure 1A). XB130 was predominantly expressed in cytoplasm in OS tissues (Figure 1B). The positive XB130 staining (Figure 1B) was 40% in OS tissues, and negative XB130 staining (Figure 1C) was 60% in OS tissues. The expression levels of XB130 in OS tissues were distinctly higher than those in adjacent non-tumor tissue (P=0.0001).
Figure 1: The XB130 protein expression in osteosarcoma tissues. Representative images of XB130 protein immunostaining in osteosarcoma tissues and the adjacent non-tumor tissue. A: adjacent non-tumor tissue (negative staining); B: Osteosarcoma tissue (positive staining); C: Osteosarcoma tissue (negative staining) (Magnification× 200).
The relationship between XB130 expression and clinicopathogic features
The patients were divided into two groups according to the staining scores: XB130 positives (high expression with score = 3) and negatives (low expression with score <3). As shown in (Table 1), XB130 was significantly up regulated in osteosarcoma patients with advanced clinical stage (P = 0.002) and positive distant metastasis (P=0.001) as compared to those with low clinical stage and without distant metastasis. No significant difference was observed between the expression of XB130 and patient’s age, gender, tumor size, location and response to chemotherapy.
The relationship between XB130 expression and cumulative survival rate
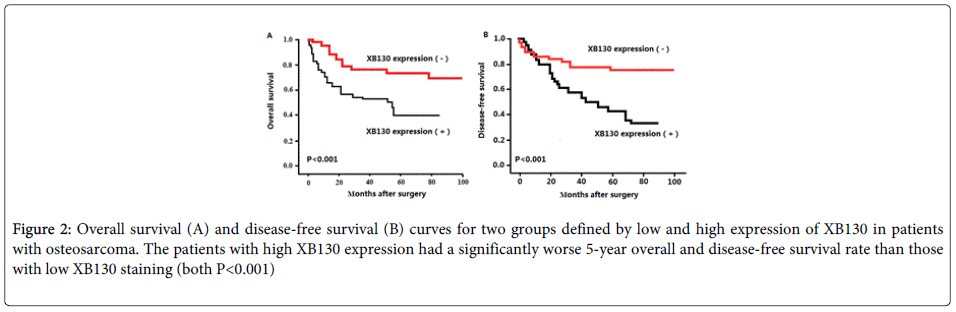
Using Kaplan–Meier method and log-rank test, the overall survival (OS, Figure 2A, P<0.001) and disease-free survival (DFS, Figure 2B, P<0.001) of osteosarcoma tissues with high XB130 expression were both significantly shorter than those with low XB130 expression (both P<0.001).
Figure 2: Overall survival (A) and disease-free survival (B) curves for two groups defined by low and high expression of XB130 in patients with osteosarcoma. The patients with high XB130 expression had a significantly worse 5-year overall and disease-free survival rate than those with low XB130 staining (both P<0.001)
Univariate analysis of prognostic factors
The univariate analysis of prognostic factors of OS is summarized in Table 2. Distant metastasis (P= 0.0024), advanced clinical stage (P=0.013), tumor size (P=0.02), response to chemotherapy (P=0.04) and high XB130 expression (P=0.001) were statistically significant risk factors affecting the outcome of patients with OS.
| 95% Confidence | |||
|---|---|---|---|
| Variable | Hazard ratio | Limits | p |
| Tumor metastasis | |||
| No (n=41) | 1 | 1.453-5.824 | 0.0024 |
| Yes (n=19) | 2.762 | ||
| Size of primary OS (cm) | |||
| <8 (n=28) | 1 | 1.620-5.219 | 0.02 |
| ≥8 (n=32) | 2.364 | ||
| Clinical stage | |||
| Ⅰ/ⅡA (n=23) | 1 | 1.372-6.837 | 0.013 |
| ⅡB/Ⅲ (n=37) | 2.765 | ||
| Response to chemotherapy | |||
| Poor (n=39) | 1 | 0.87-4.65 | 0.04 |
| Good (n=21) | 1.28 | ||
| XB130 expression | |||
| Low (n=36) | 1 | 1.47-5.36 | 0.01 |
| High (n=24) | 2.435 | ||
Table 2: Univariate analysis of prognostic factors in 60 cases of OS
Multivariate analysis of prognostic factors
The multivariate analysis of prognostic factors of OS is summarized in (Table 3). High XB130 expression (P=0.004) and distant metastasis (P=0.03) were found to be statistically significant independent poor prognostic factors.
| 95% Confidence | |||
|---|---|---|---|
| Variable | Hazard ratio | Limits | p |
| Tumor distant metastasis | |||
| No (n=41) | 1 | 1.38-5.02 | 0.03 |
| Yes (n=19) | 2.284 | ||
| XB130 expression | |||
| Low (n=36) | 1 | 1.483-8.067 | 0.004 |
| High (n=24) | 3.356 | ||
Table 3: Multivariate analysis of prognostic factors in 60 cases of OS
Effects of XB130 depletion on tumor cell growth
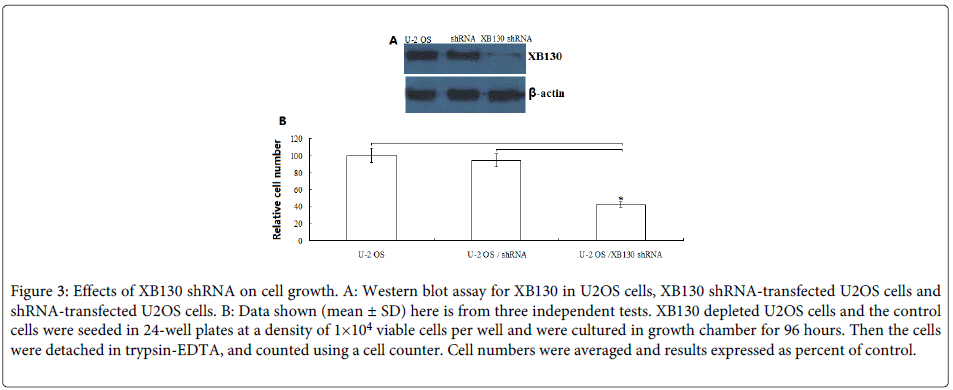
To examine the role of XB130 on the growth of human OS cells, the U2OS cells were stably transfected with human XB130 shRNA or control shRNA. Down-regulation of XB130 by shRNA transfection showed low-expression of XB130 protein as confirmed by Western blot analysis (Figure 3A).
Figure 3: Effects of XB130 shRNA on cell growth. A: Western blot assay for XB130 in U2OS cells, XB130 shRNA-transfected U2OS cells and shRNA-transfected U2OS cells. B: Data shown (mean ± SD) here is from three independent tests. XB130 depleted U2OS cells and the control cells were seeded in 24-well plates at a density of 1×104 viable cells per well and were cultured in growth chamber for 96 hours. Then the cells were detached in trypsin-EDTA, and counted using a cell counter. Cell numbers were averaged and results expressed as percent of control.
Next, we determined the proliferation of U2OS, XB130 shRNA /U2OS and control shRNA/ U2OS. As shown in (Figure 3B), compared with U2OS and control shRNA/ U2OS, the proliferation of XB130 shRNA /U2OS was significantly inhibited to 60% (P<0.01) at 96 h. There was no significant difference between U2OS and control shRNA/ U2OS (P>0.05).
Down-regulation of XB130 decreased cell motility and invasion in vitro
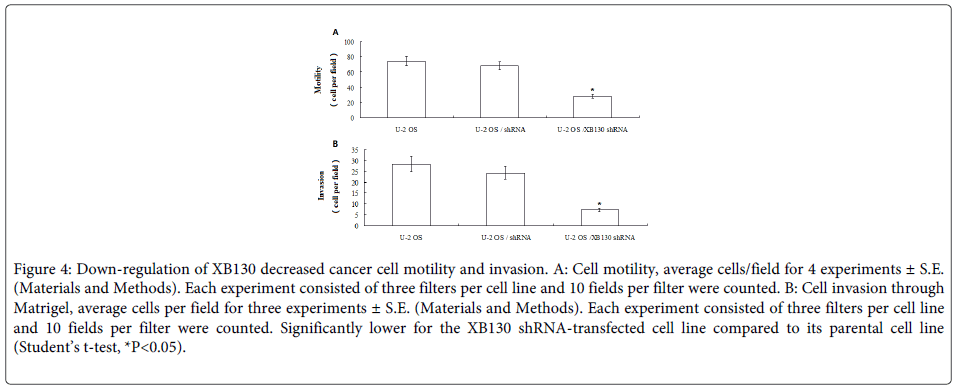
Chemotactic motility was measured as described in Materials and Methods. Motility was determined over a 24 hours period, a time period, which was shown to ensure that all motile cells remained attached to the lower side of the filter. Cell motility varied from U2OS cells, XB130 shRNA-transfected U2OS cells and shRNA-transfected U2OS cells. Motility of the XB130 shRNA-transfected U2OS was significantly lower than their respective parental cell lines U2OS and shRNA-transfected U2OS cells (Figure 4A, Student’s t-test, P<0.05, respectively).
Figure 4: Down-regulation of XB130 decreased cancer cell motility and invasion. A: Cell motility, average cells/field for 4 experiments ± S.E. (Materials and Methods). Each experiment consisted of three filters per cell line and 10 fields per filter were counted. B: Cell invasion through Matrigel, average cells per field for three experiments ± S.E. (Materials and Methods). Each experiment consisted of three filters per cell line and 10 fields per filter were counted. Significantly lower for the XB130 shRNA-transfected cell line compared to its parental cell line (Student’s t-test, *P<0.05).
To allow for direct comparison between cell invasion and cell motility, the invasive abilities of the cell lines were measured using the assay conditions optimised for the motility assay but with a coating of Matrigel on the filter separating the upper and lower compartment of the Boyden chamber. The invasive ability was significantly lower in the XB130 shRNA-transfected U2OS cells, than their corresponding parental cell lines U2OS and shRNA-transfected U2OS cells (Figure 4B, Student’s t-test, P<0.05).
Down-regulation of XB130 inhibits tumorigenicity and lung metastasis of OS cells in vivo
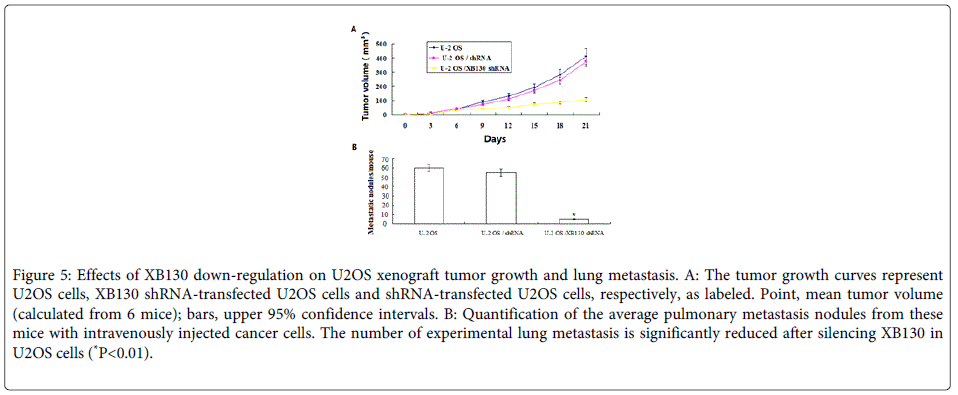
To further determine a role of XB130 in progression of OS, we did an in vivo animal experiment. We found that U2OS cells stably transfected with XB130-shRNA formed substantially smaller tumors in nude mice compared with those transfected with negative control shRNA (Figure 5). The tumor volume for mice with cells transfected with the XB130 shRNA was 98.7 ± 40.6 mm3 compared with 410.8 ± 140.3 or 390.7±128.5 mm3 for mice with U2OS cells or negative control shRNA, respectively (P=0.014 or P=0.039) respectively).
Figure 5: Effects of XB130 down-regulation on U2OS xenograft tumor growth and lung metastasis. A: The tumor growth curves represent U2OS cells, XB130 shRNA-transfected U2OS cells and shRNA-transfected U2OS cells, respectively, as labeled. Point, mean tumor volume (calculated from 6 mice); bars, upper 95% confidence intervals. B: Quantification of the average pulmonary metastasis nodules from these mice with intravenously injected cancer cells. The number of experimental lung metastasis is significantly reduced after silencing XB130 in U2OS cells (*P<0.01).
Next, we sought to determine whether XB130 silencing in U2OS cells would affect its ability to metastasize in vivo. To that end, nude mice were injected intravenously with U2OS cells, XB130 shRNA-transfected U2OS and control shRNA-transfected U2OS cells. As demonstrated in (Figure 5B), XB130 silencing resulted in a significant reduction in the median number of lung metastases in mice injected with XB130 shRNA-transfected U2OS cells (P<0.01) Overall, these results confirm that knockdown of XB130 inhibits OS tumorigenicity and metastasis.
Discussion
XB130 has been recently cloned as a molecular homologue of AFAP-110 [16]. Previous study provides evidence that XB130 is regulated by the actin cytoskeleton, and it impacts key cytoskeletal functions, such as cell migration and invasion [17]. XB130 is also involved in cell proliferation, cell survival and tumorigenesis [10]. Recently, expression of XB130 has found in a variety of cell lines derived from thyroid, lung, esophageal, pancreatic, and colon cancers. In pancreatic cancer [18], XB130 expression was significantly up-regulated in tumor tissues, however, no XB130 expression was found in the adjacent normal pancreas. Furthermore, increased XB130 expression was correlated with lymph node metastasis, distant metastasis, high tumour-node-metastasis (TNM) stage, and high tumour grade. The survival of patients with high XB130 expression was significantly worse than that of the patients with low XB130 expression. In hepatocellular carcinoma [14], positive expression rate of XB130 in HCC was 75.0%, but XB130 is not associated with the postoperative prognosis of patients with HCC. In human esophageal squamous cell carcinoma (ESCC) [11], 71.2% of the patients expressed XB130 in the nuclei and/or cytoplasm of the ESCC cells. Further, nuclear expression of XB130 was an independent prognostic factor of postoperative survival. Shi et al. [13] has reported that both XB130 mRNA and protein expression were detectable in normal gastric tissues. The reduced XB130 protein expression is a prognostic biomarker for shorter survival and a higher recurrence rate in patients with GC, as well as for the response to chemotherapy. The studies above indicated XB130 expression was related with the tumor type.
In the current study, we examined a cohort of 60 OS specimens and report the evidence that XB130 correlated with distant metastasis and high clinical stage. No significant difference was observed between the expression of XB130 and patients’ age, gender, tumor size, location and response to chemotherapy. The patients with high XB130 expression had a significantly worse 5-year overall and disease-free survival rate than those with low XB130 staining. The multivariate analysis showed that high XB130 expression was found to be statistically significant independent poor prognostic factors. Our data suggest that XB130 may serve as a marker for poor prognoses. To our knowledge, this is the first IHC study to investigate the potential utility of XB130 as a biomarker of OS among Chinese patients.
In thyroid tumor cells, knockdown of XB130 using small interfering RNA inhibited G (1)-S phase progression, induced spontaneous apoptosis, and enhanced intrinsic and extrinsic apoptotic stimulus-induced cell death. Growth of tumors in nude mice formed from XB130 shRNA stably transfected WRO cells were significantly reduced, with decreased cell proliferation and increased apoptosis [12]. In the present study, we found that knockdown of XB130 inhibited invasion and mobility in OS cells in vitro, but the molecular contribution of XB130 in lung metastasis of OS cancer is still unknown. Particularly, it is unclear whether XB130 gene is relevant to the progression of OS cancers in lung tissue. Here, we found XB130 silencing resulted in a significant reduction in the number of lung metastases in mice injected with XB130 shRNA-transfected U2OS cells.
In summary, our present study has provided first evidence that XB130 overexpression may be related to the prediction of metastasis potency and poor prognosis for osteosarcoma patients, suggesting that XB130 may serve as a prognostic marker for the optimization of clinical treatments. Furthermore, XB130 is the potential molecular target for osteosarcoma therapy.
References
- Miiji LN, Petrilli AS, Di Cesare S, Odashiro AN, Burnier MN Jr, et al. (2011) C-kit expression in human osteosarcoma and in vitro assays. Int J Clin Exp Pathol 4: 775-781.
- Unni KK (2009) General Aspects and Data on 10,165 Cases. Dahlin´s bone tumors 6:122-157.
- Bacci G, Ferrari S, Longhi A, Forni C, Zavatta M, et al. (2002) High-grade osteosarcoma of the extremity: differences between localized and metastatic tumors at presentation. J Pediatr Hematol Oncol 24: 27-30.
- Petrilli AS, de Camargo B, Filho VO, Bruniera P, Brunetto AL, et al. (2006) Results of the Brazilian Osteosarcoma Treatment Group Studies III and IV: prognostic factors and impact on survival. J ClinOncol 24: 1161-1168.
- Meyers PA, Heller G, Healey JH, Huvos A, Applewhite A, et al. (1993) Osteogenic sarcoma with clinically detectable metastasis at initial presentation. J Clin Oncol 11: 449-453.
- Baldini N, Scotlandi K, Serra M, Picci P, Bacci G, et al. (1999) P-glycoprotein expression in osteosarcoma: a basis for risk-adapted adjuvant chemotherapy. J Orthop Res 17: 629-632.
- Benassi MS, Molendini L, Gamberi G, Sollazzo MR, Ragazzini P, et al. (1997) Altered G1 phase regulation in osteosarcoma. Int J Cancer 74: 518-522.
- Bakhshi S, Radhakrishnan V (2010) Prognostic markers in osteosarcoma. Expert Rev Anticancer Ther 10: 271-287.
- Magnan H, Chou AJ, Chou JF, Yeung HW, Healey JH, et al. (2010) Noninvasive imaging with thallium-201 scintigraphy may not correlate with survival in patients with osteosarcoma. Cancer 116: 4147-4151.
- Shiozaki A, Liu M (2011) Roles of XB130, a novel adaptor protein, in cancer. J Clin Bioinforma 1: 10.
- Shiozaki A, Kosuga T, Ichikawa D, Komatsu S, Fujiwara H, et al. (2013) XB130 as an independent prognostic factor in human esophageal squamous cell carcinoma. Ann Surg Oncol 20: 3140-3150.
- Shiozaki A, Lodyga M, Bai XH, Nadesalingam J, Oyaizu T, et al. (2011) XB130, a novel adaptor protein, promotes thyroid tumor growth. Am J Pathol 178: 391-401.
- Shi M, Huang W, Lin L, Zheng D, Zuo Q, et al. (2012) Silencing of XB130 is associated with both the prognosis and chemosensitivity of gastric cancer. PLoS One 7: e41660.
- Zuo Q, Huang H, Shi M, Zhang F, Sun J, et al. (2012) Multivariate analysis of several molecular markers and clinicopathological features in postoperative prognosis of hepatocellular carcinoma. Anat Rec (Hoboken) 295: 423-431.
- Sobin LH, Wittekind C (2002) UICC-TNM Classificaton of Malignant Tumors. New York: Wiley.
- Xu J, Bai XH, Lodyga M, Han B, Xiao H, et al. (2007) XB130, a novel adaptor protein for signal transduction. J Biol Chem 282: 16401-16412.
- Lodyga M, Bai XH, Kapus A, Liu M (2010) Adaptor protein XB130 is a Rac-controlled component of lamellipodia that regulates cell motility and invasion. J Cell Sci 123: 4156-4169.
- Zhang J, Jiang X, Zhang J (2014) Prognostic significance of XB130 expression in surgically resected pancreatic ductal adenocarcinoma. World J Surg Oncol 12: 49.
Citation: Wang X, Huang H, Hao F, Li Y, Zhang L (2014) XB130 Expression in Human Osteosarcoma: A Clinical and Experimental Study. J Clin Exp Pathol 4:169. DOI: 10.4172/2161-0681.1000169
Copyright: © 2014 Wang X, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Select your language of interest to view the total content in your interested language
Share This Article
Recommended Journals
Open Access Journals
Article Tools
Article Usage
- Total views: 15357
- [From(publication date): 6-2014 - Aug 29, 2025]
- Breakdown by view type
- HTML page views: 10729
- PDF downloads: 4628