Research Article Open Access
A Revisited Study On Phylogeography And Phylogenetic Diversity Of Myricaria (Tamaricaceae)
Youhua Chen*Department of Zoology, University of British Columbia, Vancouver, Canada
- *Corresponding Author:
- Youhua Chen
Department of Renewable Resources
University of Alberta
Edmonton, T6G 2H1, Canada
E-mail: yhchen@zoology.ubc.ca
Received date: June 29, 2013; Accepted date: July 29, 2013; Published date: July 30, 2013
Citation: Chen Y (2013) A Revisited Study on Phylogeography and Phylogenetic Diversity of Myricaria (Tamaricaceae). J Ecosys Ecograph 3:132. doi:10.4172/2157-7625.1000132
Copyright: © 2013 Chen Y. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and and source are credited.
Visit for more related articles at Journal of Ecosystem & Ecography
Abstract
As the first part of the present study, the ancestral distribution origin of Myricaria at a Eurasian perspective, a genus of Tamaricaceae, was re-evaluated by comparing three different methods, including Statistical Dispersal-Vicariance Analysis (S-DIVA), Dispersal-Extinction-Cladogenesis Analysis (DEC) and Bayesian Binary MCMC Analysis (BBM). In addition, Parsimony Analysis of Endemicity (PAE) was adopted to identify areas of endemism and test whether the endemic areas could be congruent to the origin of distribution. My results showed that two of the three ancestral distribution methods consistently identified East Asia as the historical origin of distribution, which was in line with a previous study using the distributional information of Myricaria in China only based on classical dispersal-vicariance method. PAE further supported such an observation, indicating that PAE could be an option for identifying origin of distribution. Further, it was found that the emigration of Myricaria from East Asia to the rest of Eurasian regions only occurred when the drastic uplifting of Qinghai-Tibet Plateau emerged. As another part of the study, I quantified the phylogenetic diversity patterns of Myricaria species using six phylogenetic diversity metrics, for the purpose of identifying the highest-priority species for conservation. For those species endemic to the origin of distribution, it was found that two species were assigned the highest conservation values under the phylogenetic framework, M. laxiflora and M. elegans var. tsetangensis.
Keywords
Historical biogeography; Species extinction; Biodiversity conservation; Areas of endemicity
Introduction
The small genus Myricaria is a part of the family Tamaricaceae, composed of around 11 species over the world [1]. Most Myricaria species have distributional ranges in China, especially in the Himalayan region [2]. Its ancestral origin is generally believed to be derived from Himalayan region. Due to increasing habitat fragmentation and industrialization of the native habitats, many species are believed to undergo extinction threats in China [1].
In previous studies [1,2], Himalayan region (Qinghai-Tibet Plateau) was believed to be the historical origin Myricaria by utilizing statistical dispersal-vicariance analysis. However, dispersalvicariance analysis might be misleading due to its incapability to identify extinction events [3]. Here, by utilizing different methods to reconstruct ancestral distributional ranges [4,5], the purpose of the present study is to re-evaluate the validity of the Himalayan-origin hypothesis. I am in particular focusing on the discrepancy caused by different ancestral range reconstruction methods. I also introduce the Parsimony Analysis of Endemicity (PAE) [6,7] as a new way to identify the origin of distribution of Myricaria taxa. I regard that both origin of distribution and areas of endemism are two identical quantities, although they should have different biological foundations. If the areas of endemism identified by PAE are concordant with the origin of distribution reconstructed by ancestral range inference, PAE might be a potentially promising option for identifying the origin of distribution. Otherwise, PAE would not be recommended. In a summary, the central objective of my study is to offer alternative statistical diagnoses on the distributional origin of the genus.
Besides, though conservation strategies have been advocated for the genus [1], no quantitative assessment of conservation priorities for each species has been proposed. As such, it should be of some help to utilize phylogenetic diversity methods to quantify the extinction threats of the species and evaluate the conservation values of each species [8-11]. Therefore, in the present study, another objective is to quantify conservation priorities of Myricaria species on the basis of phylogenetic perspectives.
To avoid possible repetitions of the previous studies [1,2], I am not discussing in detail the phylogenetic patterns of the Myricaria species. Instead, my study will focus on the comparative methods for identifying the ancestral distribution, the historical biogeographic scenarios and the conservation priority of Myricaria species.
Materials and Methods
Sequence data
DNA sequences for Myricaria taxa were collected for the nuclear ITS region (partial sequences of ITS1 and ITS2, complete sequence of 5.8S) and psbA-trnH intergenic spacer, which have been widely used in the molecular phylogenetic analysis. These sequences, available from GenBank database were directly obtained from previous works [1,2]. The list of species and associated GenBank sequence accession numbers are presented in Table 1.
| No. | GenBank ID | Species | ED | TD | ES | UPD | PE | BED |
|---|---|---|---|---|---|---|---|---|
| 1 | EU240605 | M wardii* | 7.766 | 0.033 | 6.276 | 5.898 | 9.775 | 6.711 |
| 2 | AY572413 | Mgermanicassp germanica | 6.721 | 0.027 | 4.907 | 4.679 | 5.681 | 2.44 |
| 3 | EU240606 | M platyphylla | 5.11 | 0.02 | 2.894 | 2.398 | 5.955 | 2.339 |
| 4 | EU240604 | M squamosa | 5.11 | 0.02 | 2.894 | 2.398 | 5.555 | 1.939 |
| 5 | EU240600 | M paniculata* | 4.392 | 0.02 | 2.139 | 1.513 | 6.426 | 2.683 |
| 6 | AF484746 | M germanica ssp alopecuroides | 3.848 | 0.019 | 1.283 | 0.427 | 5.291 | 1.412 |
| 7 | EU240596 | M bracteata | 3.848 | 0.019 | 1.283 | 0.427 | 5.291 | 1.412 |
| 8 | EU240602 | M pulcherrima | 6.273 | 0.023 | 4.269 | 4.142 | 8.397 | 5.058 |
| 9 | EU240607 | M prostrata | 8.534 | 0.042 | 7.144 | 5.366 | 5.937 | 3.114 |
| 10 | EU240608 | M rosea | 8.534 | 0.042 | 7.144 | 5.366 | 6.831 | 4.008 |
| 11 | EU240609 | M laxiflora* | 11.332 | 0.083 | 11.332 | 9.937 | 13.006 | 10.551 |
| 12 | EU240594 | M elegans | 15.384 | 0.333 | 15.384 | 5.484 | 8.428 | 6.778 |
| 13 | EU240595 | M elegans var tsetangensis* | 15.384 | 0.333 | 15.384 | 5.484 | 12.084 | 10.434 |
Table 1: Phylogenetic diversity measurements of Myricaria species based on the Bayesian MCMC maximum clade credibility tree presented in Figure 2. Endemic species only found in East Asia (China) were marked in asterisks. GenBank accession numbers for the sequences utilized for the present study are provided as well.
Phylogenetic analysis and reconstruction
Sequence alignment was done using Cluster X program [12]. The region with ambiguous alignment was manually excluded using BioEdit program [13]. Phylogenetic analysis was performed using maximum parsimony method (MP) on PAUP version 4b10 package [14]. Partition homogeneity test was performed to see the heterogeneity of the combined data for maximum parsimony analysis [1]. Most-parsimonious trees were obtained by 1000 replicates of random sequence addition using Tree Bisection-Reconnection (TBR) branch swapping under the Fitch criterion. The Bayesian tree was constructed using the program MrBayes [15]. The evolutionary model GTR+Gamma was selected using MrModeltest [16] under the Akaike information criterion. The Markov chain Monte Carlo chains were run simultaneously for 1000000 generations and trees were sampled for each 1000 generation. Two independent chains were run simultaneously. Average standard deviation of split frequencies was used as the diagnosis of stationary status of the MCMC chains. The first 25% trees were regarded as the unstable burn-in ones and discarded, the remaining trees were used to construct 50% majority rule consensus tree [4].
Molecular dating
Because of the lack of fossil records [2], I used the relaxed molecular clock Bayesian model [17] to date the tree with BEAST software by inputting it as the starting tree [18]. The general substitution rate ( u =1.0×10-9ss-1year-1 ) of the plastid sequence was used [4,19]. 1000000 generations and trees were sampled for each 1000 generation. Stationary status of MCMC chains were checked using Tracer [20]. The maximum clade credibility tree was constructed using TreeAnnotator [18] after pruning the first 400 sampled trees. The tree was viewed and edited using FigTree software [21].
Ancestral range reconstruction
Prior to the reconstruction of historical ranges, I defined the following five bigoeographic regions by using Qinghai-Tibet Plateau as the reference center: East Asia (A), covering the ranges of south, western and eastern part of China, but excluding the northern part of China; South Asia (S), covering the countries including India, Nepal, Pakistan and others; Central Asia (C), including the countries in that region, like Afghanistan and others; North Asia (D), including northern part of China (Xinjiang, Inner Mongolia), Mongolia and Eastern part of Russia; and Europe (E).
Reconstruction of ancestral ranges were implemented using the software RASP [5,22]. Three methods available were all carried out, including S-DIVA method [22,23], Bayesian Binary MCMC method (BBM) [4,5] and dispersal-extinction-cladogenesis method (DEC) [3].
Parsimony Areas of Endemicity (PAE)
Areas of endemism [6,7,24] were defined as the areas where two or more species that have sympatric distribution. PAE is a cladistic method for identifying areas of endemicity, in which the clades were identified as endemic areas when the lineages leading to these clades were supported by two or more synapomorphies [25]. In the empirical studies of historical biogeography, PAE has been widely employed [25,26]. In the present study, I employ it to find the areas of endemism for Myricaria and compare whether the identified endemic areas could be fully congruent with the reconstructed origin of distribution from the abovementioned ancestral range reconstruction methods. The abovementioned five biogeographic regions were used as the operating areal units for the PAE analysis.
Phylogenetic diversity analysis of Myricaria species
The following species-based phylogenetic diversity metrics were calculated, including evolutionary distinctiveness (ED) [8,27], taxonomic distinctiveness (derived from ED index, but the branch lengths are ignored and all treated equally during the calculation) (TD) [28], equal splits (ES) [8,29], unshared phylogenetic diversity (i.e., the terminal branches linking to the external species directly) (UPD) [27], Phylogenetic Endemism (PE) [30,31] and Biogeographically weighted phylogenetic Distictiveness (BED) [32] indices. The definition and calculation of these indices were not present here for simplicity, and readers could refer to the original papers for further details.
Results
Phylogenetic history of Myricaria species
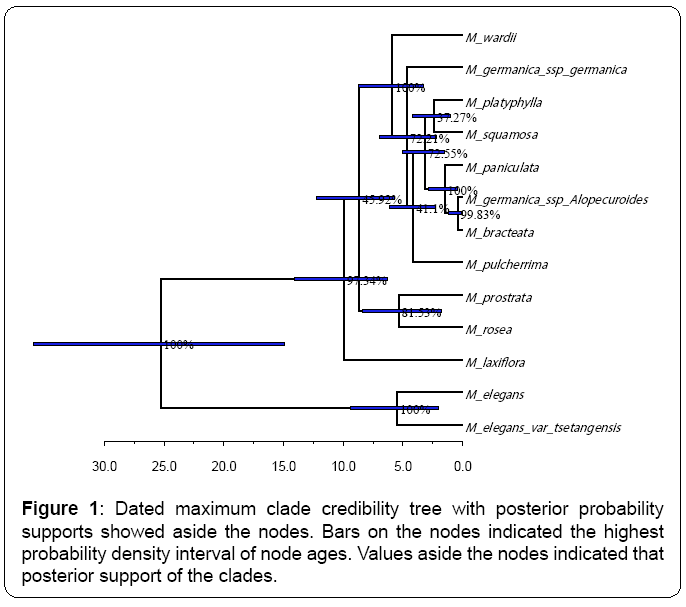
The constructed 50% major-rule consensus MP tree is highly identical to the previous studies [1,2].Thus, it is not presented here for simplicity. The dated maximum clade credibility Bayesian tree (Figure 1) based on was highly similar to the MP tree and previous studies [1,2], using GTR+G as the evolutionary model which was selected by MrModelTest. The ancestor of all Myricaria species has a clade age around 25 million years ago (Ma) (95% HPD: 14.9~35.9 Ma). For all the subsequent analyses, this dated Bayesian tree was used to infer the origin of distribution for Myricaria species and identify conservation values of species since it contained branch length information and fully dichotomous.
Origin of distribution of Myricaria species
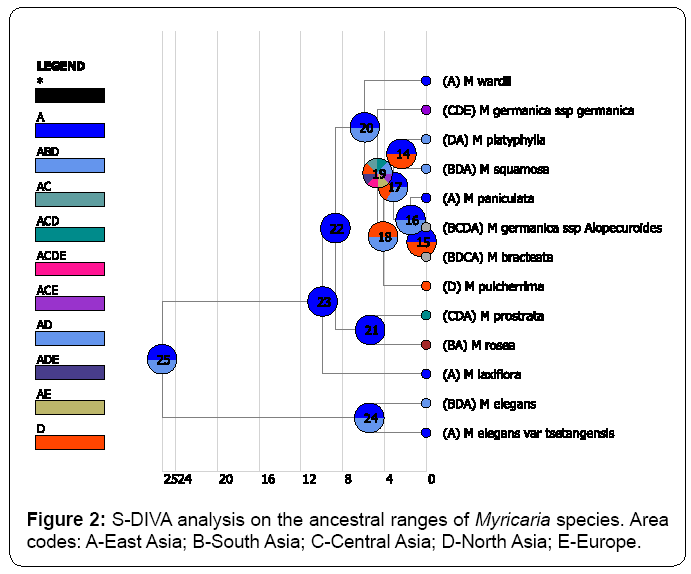
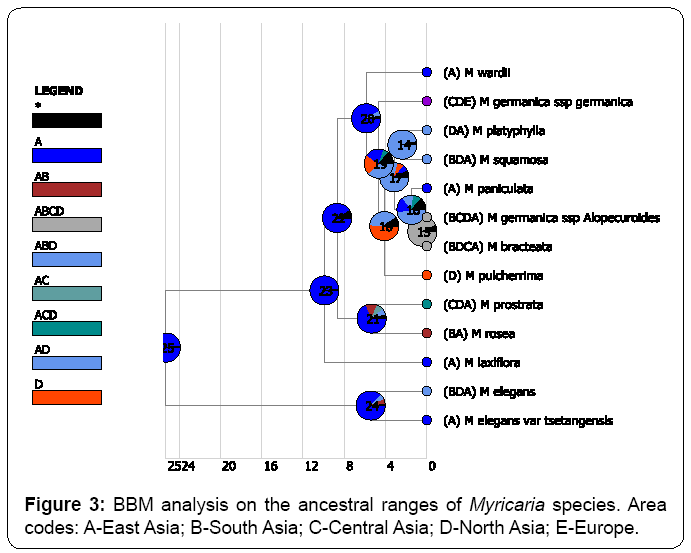
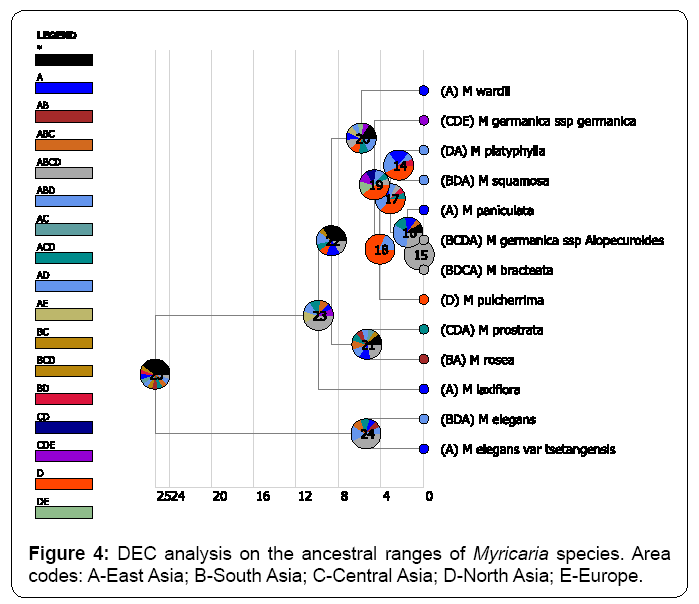
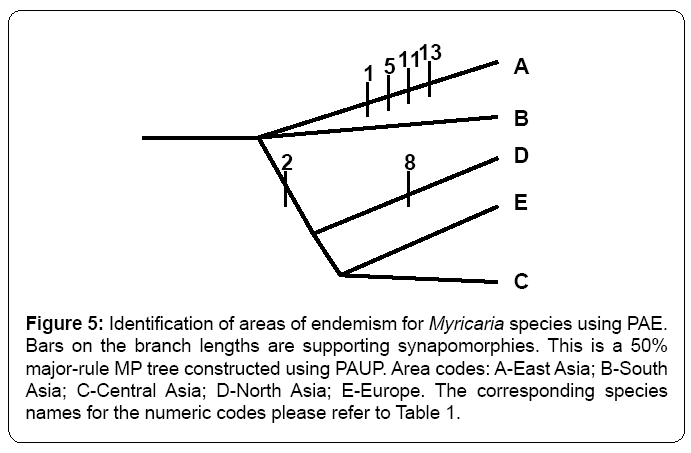
Both BBM and S-DIVA methods congruently identified East Asia is the origin of distribution of Myricaria species at the node of 25 (Figures 2, 3). In contrast, DEC method (Figure 4) returned a very confusing diagnosis on the origin of distribution of Myricaria because either single region or combined regions were supported with very low probabilities in the root. PAE method further verified that East Asia is the areas of endemism, as supported by the four endemic species (Figure 5). Thus, in my study, without specific references, the inferred origin of distribution is identical to the identified area of endemism for Myricaria species, both meaning East Asia (Qinghai-Tibet Plateau).
Figure 5: Identification of areas of endemism for Myricaria species using PAE. Bars on the branch lengths are supporting synapomorphies. This is a 50% major-rule MP tree constructed using PAUP. Area codes: A-East Asia; B-South Asia; C-Central Asia; D-North Asia; E-Europe. The corresponding species names for the numeric codes please refer to Table 1.
Phylogenetic diversity patterns of Myricaria species
As showed in Table 1, the six phylogenetic diversity indices quantified the relative importance of priority species for conservation under different phylogenetic perspectives. In specific, M. elegans and M. elegans var. tsetangensis have equally highest ED, TD, and ES indices, while M. laxiflora has highest UPD, PD, and BED indices.
Discussion
Evolutionary history of Myricaria species
Based on my fossil-free analysis, I estimated that the most recent ancestor for all the Myricaria species could be dated back to 25 Ma. My estimation is based on the substitution rate of plastid sequence. In the previous study [2], the likely earliest occurrence time of Tamarix was used as the reference dating time of the occurrence of Myricaria ancestor at the time of 57 Ma.
The previous work [2] suggested the main divergence events occurred at the time during the Late Pliocene and Early Pleistocene (approximately 2.3~1.46 Ma). My re-analysis was a bit different from such an observation with some discrepancies on the time interval. As evidenced by Figure 1, most cladogenesis events happened around the time frame during 5~2.5 Ma, which is a bit earlier than the previous observation [2]. My estimation of the late burst of species divergence during 5~2.5 Ma is fully consistent with beginning time of the principal glaciation events happened in the Qinghai-Tibet Plateau (around 4.43~1.21 Ma) [2,33,34].
During the time, another paramount geological event is the significant uplift of Qinghai-Tibet Plateau, which was happening during the time of Late Pliocene and Early Pleistocene [2,35,36]. These series of events, the glaciation and uplifting of plateau, are fully concordant with the rapid branching of Myricaria species. Thus, historical geological events have profoundly driven the dispersal, extinction and speciation of Myricaria, causing the accelerating divergence of the genus during that time window as discussed below.
Historical biogeograhic analysis of distributional ranges of Myricaria
Because DEC method returned uninformative results on the ancestral distribution of Myricaria species (Figure 4), I excluded the discussion referring to the method. Other two methods, S-DIVA and BBM methods were the focused ones.
As showed in (Figure 2, Figure 3), it was found that the “out-of- East Asia” historic episodes only happened after the drastic uplifting of Qinghai-Tibet Plateau at 5 Ma. During the orogenic movement of the plateau, many lineages of Myricaria processed a series of dispersal events from East Asia to other Eurasian regions. Interestingly, no matter what methods were used, dispersal was always identified as the principal driver of contemporary distribution of Myricaria species. In contrast, the contribution of vicariance or extinction events is quite rare. 22 dispersal events were congruently found by S-DIVA and BBM methods. In contrast, only 2 vicariance events and 1 extinction event were found by S-DIVA method; while only 1 extinction event and no vicariance events were identified by BMM method.
Contribution of endemic species in the origin of distribution to the diversity of Myricaria species
Based on PAE (Figure 5), there are four endemic species characterizing the origin of distribution of Myricaria in East Asia, which are M. laxiflora, M. elegans var. tsetangensis, M. paniculata, and M. wardii. Based on the phylogenetic diversity scenario, it is predicted that the loss of these four species would lead to the loss of ED by 38%, TD by 46%, ES by 43%, UPD by 43%, PE by 42% and EBD by 52%. Not surprisingly, the loss of EBD would be the highest since it is strongly correlated to the endemism status of species.
One Myricaria species, M. laxiflora, is subjected to immediate extinction due to Three Gorges Dam Project [2]. This species is endemic to Three Gorges reservoir only (Chongqing City and Hubei Province), listed as one of endangered plant species in China. It has gained much public and scientific awareness [2,37,38]. Conservation strategies on the species have been carried out in recent years. As mentioned above, it has the highest UPD, PE and BED values; second highest ED, TD and ES values (Table 1).
Another endemic species M. elegans var. tsetangensis might be overlooked up to date because it is a subspecies and not subjected to immediate extinction due to dam construction. However, from the perspective of phylogenetic diversity indices, it has highest ED, TD and ES values, second highest PE and BED values. Finally, the contribution of another two endemic species, M. paniculata and M. wardii, to the global phylogenetic diversity pattern is not so remarkable. But still, M. wardii has second highest UPD value (Table 1).
In conclusion, conservation priority should be given to M. laxiflora and M. elegans var. tsetangensis on the basis of their unique contribution to global diversity of Myricaria genus and the diagnosis of areas of endemism.
Conflict of Interests
The author declares that there is no conflict of interests regarding the materials of the paper.
References
- Wang Y, Liu Y, Liu S, Huang H (2009) Molecular phylogeny of Myricaria (Tamaricaceae): implications for taxonomy and conservation in China. Botanical Studies 50: 343-352.
- Liu Y, Wang Y, Huang H (2009) Species-level phylogeographical history of Myricaria plants in the mountain ranges of western China and the origin of M. laxiflora in the Three Gorges mountain region. Mol Ecol 18: 2700-2712.
- Ree R, Smith S (2008) Maximum likelihood inference of geographic range evolution by dispersal, local extinction and cladogenesis. Syst Biol 57: 4-14.
- Ali S, Yan Y, Pfosser M, Wetschnig W (2012) Inferences of biogeographical histories within subfamily Hyacinthoideae using S-DIVA and Bayesian binary MCMC analysis implemented in RASP (Reconstruct Ancestral State in Phylogenies). Annals of Botany 109: 95-107.
- Yu Y, Harris A, He XJ (2013) A Rough Guide to RASP 2.1 (Beta). University of Sao Paulo, Brazil.
- Morrone JJ, Crisci J V (1995) Historical Biogeography: Introduction to Methods. Annu Rev Ecol Syst 26: 373-401.
- Morrone J (1994) On the Identification of Areas of Endemism. Syst Biol 43: 438-441.
- Redding D, Hartmann K, Mimoto A, Bokal D, Devos M, et al. (2008) Evolutionarily distinctive species often capture more phylogenetic diversity than expected. J Theor Biol 251: 606-615.
- Faith D (1992) Conservation evaluation and phylogenetic diversity. Biological Conservation 61: 1-10.
- Posadas P, Esquivel D, Crisci J (2001) Using phylogenetic diversity measures to set priorities in conservation: an example from southern South America. Conservation Biology 15: 1325-1334.
- Strecker A, Olden J, Whittier J, Paukert C (2011) Defining conservation priorities for freshwater fishes according to taxonomic, functional and phylogenetic diversity. Ecological Applications 21: 3002-3013.
- Thompson J, Gibson T, Plewniak F, Jeanmougin F, Higgins D (1997) The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 24: 4876-4882.
- Hall T (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symposium Series 41: 95-98.
- Swofford D (2002) PAUP* Phylogenetic Analysis Using Parsimony, version 4.0b10. Sinauer Associates, Sunderland, MA.
- Huelsenbeck J, Ronquist F (2001) MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17: 754-755.
- Nylander J (2004) MrModelTest, version 2..
- Drummond A, Ho S, Phillips M, Rambaut A (2006) Relaxed phylogenetics and dating with confidence. PLoS biology 4: e88.
- Drummond A, Rambaut A (2007) BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evolutionary Biology 7: 214.
- Zurawski G, Clegg M, Brown A (1984) The nature of nucleotide sequence divergence between barley and maize chloroplast DNA. Genetics 106: 735-749.
- Rambaut A, Drummond A (2007) Tracer v14.
- Rambaut A (2008) FigTree v1.1.
- Yu Y, Harris A, He X (2010) S-DIVA (statistical dispersal-vicariance analysis): a tool for inferring biogeographic histories. Mol Phylogenet Evol 56: 848-850.
- Ronquist F (1997) Dispersal-Vicariance Analysis: A New Approach to the Quantification of Historical Biogeography. Syst Biol 46: 195-203.
- Rosen B (1988) From fossils to earth history: Applied historical biogeography. Analytical biogeography 437-481.
- Contreras Medina R, Vega I, Morrone J (2007) Application of parsimony analysis of endemicity to Mexican gymnosperm distributions: grid-cells, biogeographical provinces and track analysis. Biological Journal of the Linnean Society 92: 405-417.
- Chen Y, Bi J (2007) Biogeography and hotspots of amphibian species of China: Implications to reserve selection and conservation. Current Science 92: 480-489.
- Cadotte M, Davies T (2010) Rarest of the rare: advances in combining evolutionary distinctiveness and scarity to inform conservation at biogeographical scales. Diversity and Distributions 16: 376-385.
- Freitag S, Jaarsveld A (1997) Relative occupancy, endemism, taxonomic distinctiveness and vulnerability: prioritizing regional conservation actions. Biodiversity and Conservation 6: 211-232.
- Redding D, Mooers A (2006) Incorporating evolutionary measures into conservation prioritization. Conservation Biology 20: 1670-1678.
- Rosauer D, Laffan S, Crisp M, Donnellan S, Cook L (2009) Phylogenetic endemism: a new approach for identifying geographical concentrations of evolutionary history. Mol Ecol 18: 4061-4072.
- Gudde R, Joy J, Mooers A (2012) Imperiled phylogenetic endemism of Malagasy lemuriformes. Diversity and Distributions 19: 664-675.
- Tucker C, Cadotte M, Davies T, Rebelo T (2012) Incorporating geographical and evolutionary rarity into conservation prioritization. Conserv Biol 26: 593-601.
- Shi Y (2002) Characteristics of late Quaternary monsoonal glaciations on the Tibetan plateau and in East Asia. Quaternary International 97-98: 79-91.
- Zheng B, Rutter N (1998) On the problem of Quaternary glaciations, and the extent and patterns of Pleistocene cover in the Qinghai-Xizang (Tibet) Plateau. Quaternary International 45-46: 109-122.
- Harrison T, Copeland P, Kidd W, Yin A (1992) Raising Tibet. Science 255: 1663-1670.
- Shi Y, Li J, Li B (1998) Uplift and environmental changes of Qinghai-Tibetan Plateau in the Late Genozoic. Guangdong Science and Technology Press, Guangzhou, China
- Chen F, Xie Z (2009) Survival and growth responses of Myricaria laxiflora seedlings to summer flooding. Aquatic Botany 90: 333-338.
- Chen F, Xie Z (2007) Reproductive allocation, seed dispersal and germination of Myricaria laxiflora, an endangered species in the Three Gorges Reservoir aera. Plant Ecology 191: 67-75.
Relevant Topics
- Aquatic Ecosystems
- Biodiversity
- Conservation Biology
- Coral Reef Ecology
- Distribution Aggregation
- Ecology and Migration of Animal
- Ecosystem Service
- Ecosystem-Level Measuring
- Endangered Species
- Environmental Tourism
- Forest Biome
- Lake Circulation
- Leaf Morphology
- Marine Conservation
- Marine Ecosystems
- Phytoplankton Abundance
- Population Dyanamics
- Semiarid Ecosystem Soil Properties
- Spatial Distribution
- Species Composition
- Species Rarity
- Sustainability Dynamics
- Sustainable Forest Management
- Tropical Aquaculture
- Tropical Ecosystems
Recommended Journals
Article Tools
Article Usage
- Total views: 15672
- [From(publication date):
September-2013 - Nov 17, 2025] - Breakdown by view type
- HTML page views : 10853
- PDF downloads : 4819