Research Article Open Access
Biodegradation and Decolorization of Reactive Orange 16 by Nocardiopsis alba Soil Isolate
| Sugumar Shobana1* and Berla Thangam E2 | |
| 1Department of Bioinformatics, School of Bioengineering, SRM University, Kattankulathur 603203, India | |
| 2Department of Biotechnology, School of Bioengineering, SRM University, Kattankulathur 603203, India | |
| Corresponding Author : | Shobana Sugumar Department of Bioinformatics School of Bioengineering SRM University, Kattankulathur 603203, India E-mail: ksemaa@gmail.com |
| Received April 30, 2012; Accepted June 04, 2012; Published June 06, 2012 | |
| Citation: Shobana S, Thangam EB (2012) Biodegradation and Decolorization of Reactive Orange 16 by Nocardiopsis alba Soil Isolate. J Bioremed Biodeg 3:155. doi:10.4172/2155-6199.1000155 | |
| Copyright: © 2012 Shobana S, et al. This is an open-a ccess article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. | |
Related article at Pubmed Pubmed  Scholar Google Scholar Google |
|
Visit for more related articles at Journal of Bioremediation & Biodegradation
Abstract
A bacterium identified as Nocardiopsis alba was isolated from acclimated sludge from a dyeing wastewater treatment plant. This strain rapidly decolorized Reactive orange16 (RO16) at 0.1% (w/v) concentration of both sucrose and peptone supplemented in Mineral Salt Medium (MSM) under static conditions at a temperature of 30°C with in 24 h with an initial dye concentration of 100 mg/L. The organism exhibited a remarkable color removal capability (95%) even at a high concentration of 1000 mg/L (RO16) dye within 24 h. The biodegradation products were analyzed by, FTIR spectroscopy and LC-MS analysis. The LC-MS analysis indicated the presence of 1-amino- 1-napthalene sulphonic acid in degraded product of the dye. The degraded product is less toxic to the growth of Vigna mungo seeds when compared to the non degraded dye.
| Keywords |
| Nocardiopsis alba; Biodegradation; Reactive orange 16; Decolorization |
| Introduction |
| Azo dyes constitute the largest and most versatile class of dyes, used in textile industry. They are characterized by the presence of one or more azo groups -N=N- and the bond between two aromatic ring [1]. Approximately 1,00,000 different dyes are used industrially and over 0.7 million tonnes of synthetic dyes are produced worldwide annually [2]. During processing about 10-15% of used dye stuff is lost in industrial effluent. The presence of dye pollutants in the water sources makes it undesirable due to its appearance and toxicity [3]. These dyes are regarded as relatively persistent pollutants because they are extremely stable when exposed to light and aerobic conditions [4]. Various chemical and physical methods to remove color have been well used [5]. But, all those methods have suffered from disadvantages due to high operation cost, incomplete removal of color and high energy requirement. None-the-less, the ability of micro-organisms to carry out dye decolorization has received much attention. Microbial decolorization and degradation of dyes is seen as a cost-effective method for removing these pollutants from the environment [2,6]. Many bacterial, fungal and algal species have the ability to adsorb and/or degrade azo dyes [7,8]. Moreover, bacterial decolorization is normally faster compared to fungal systems with regard to the decolorization and mineralization of azo dyes. It has been observed that mixed cultures are particularly useful in this area, as some microbial consortia can collectively carry out biodegradation tasks that no individual pure strain can undertake successfully [2]. However, mixed cultures only provide an average macroscopic view of what is happening in the system, and the results are not easily reproduced, making thorough, effective interpretation of the results quite difficult. Efforts to isolate pure bacterial cultures capable of degrading azo dyes started in the 1970s with reports of a Bacillus subtilis, Aeromonas hydrophila and Bacillus cereus [9]. Recently a substantial amount of research on the subject of color removal has been carried out using single bacterial cultures like Proteus mirabilis, Pseudomonas luteola, and Pseudomonas spp., and isolated Pseudomonas spp. SUK1 has shown very promising results for the azo dye degradation under anoxic conditions [10-13]. Pseudomonas aeruginosa decolorized a commercial tannery and textile dye, Navitan Fast Blue S5R, in the presence of glucose under aerobic conditions. This organism was also able to decolorize various other azo dyes [14]. The use of pure culture system ensures reproducible data, and thus interpretation of experimental observations becomes easier. It also becoming easier to determine the detailed mechanisms of biodegradation using the tools of biochemistry and molecular biology, and this information could be useful to regulate the enzyme system in order to produce modified strains with enhanced enzymatic activities [15]. To our knowledge, this was the first study on efficient azo dye decolorization using Nocardiopsis alba strains and might help development of biological processes for treatment of dye-polluted wastewaters. In the present investigation, one of the prominent azo dyes Reactive orange 16 used in textile industry was decolorized using Nocardiopsis alba a textile soil isolate. |
| Materials and Methods |
| Chemicals and media |
| All chemicals used in our experiment were of analytical grade. Reactive orange (RO16), NADH were obtained from Sigma Aldrich Company (St. Louis, MO, USA). The physical and chemical characteristics of RO16 are listed in table 1. |
| Culture media |
| Nutrient broth and nutrient agar of Himedia Lab. Pvt. Ltd. (India) were used. Nutrient broth ingredients per liter peptone 10 g, sodium chloride 5 g, meat extract 3 g, pH 7.6. Nutrient agar ingredients per liter peptone 10 g, sodium chloride 5 g, meat extract 3 g, agar powder 30 g, and pH 7.6. Mineral salt medium (MSM) per liter; (NH4)2SO4 1 g, K2HPO4 6 g, KH2PO4 1 g, MgSO4.7H2O 0.1 g, NaCl 5 g, pH 7.0. |
| Isolation and screening of dye degrading microorganism |
| The nutrient broth along with dye (Reactive orange RO16, 100 mg l-1) was inoculated with 10% (w/v) of soil sample collected from the waste disposal site of textile processing and dye manufacturing units in and around Theco silks, Kumbakonam (India). The flask was incubated at temperature 37°C under the static condition. After 48 h of incubation, 1.0 ml of culture was diluted and plated on the nutrient agar plate containing 200 mgl-1 Reactive orange RO16. Bacterial isolate showing the clear zone around the colonies were screened for their ability to decolorize dye in the culture broth. Out of 30 isolates, the one showing faster and higher decolorization under static condition was chosen for further study. The pure culture was maintained on dyecontaining nutrient agar slants at 4°C. |
| Molecular identification of the isolate |
| Genomic DNA was isolated from the organism isolated showing maximum decolorization and its presence was checked by agarose gel (0.8%) stained with ethidium bromide. Amplification of 16S rDNA sequence by polymerase chain reaction was done using thermal cycler (Gene Amp® 2720). 16S rDNA universal primers used for PCR reaction [16]. The reaction mixture of total volume of 30 μL consisted of 3 μL of 10 X Buffer, 1 μL of 10 mM dNTPs, 1 μL of 16S rDNA primer (5 picomole/μL), 3U/μL of Taq Polymerase, 5 μL of template DNA (280 ng/ml) and 19 μL of sterile distilled water. The PCR reaction was set to initial denaturation of 94°C for 5 minutes, followed by 35 cycles of denaturation at 94°C for one minute, annealing at 55°C for one minute, extension at 72°C for one minute and final extension at 72°C for 10 minutes. The amplified products were stained with 0.5 μg/ ml ethidium bromide and loaded on 0.8% agarose gel, and the DNA fragments were separated at 100V and documented. The amplified product was subjected to cycle sequencing using ABI 3130 XL (Genetic Analyser, Applied Biosystems, USA). The sequence was deposited to Genbank (http://www.ncbi.nlm.nih.gov/Genbank). The nucleotide sequence analysis of the sequence was performed at BlastN site at NCBI server (http://www.ncbi.nlm.nih.gov/BLAST). The alignment of the sequences was performed by using Clustalw program V1.82 at European bioinformatics site (http://www.ebi.ac.uk/clustalw). The Phylogenetic tree was constructed by the neighbour joining method using mega 5.1 software [17]. |
| Decolorization at different dye concentrations |
| All decolorization experiments were performed in triplicates. 100 ml MSM broth was inoculated at 1% v/v of microbial culture. The dye was added at concentrations of 100, 250, 500, 750 and 1000 mg l-1. After 24 h aliquots (3 ml) of the culture media was withdrawn at different time intervals, centrifuged at 5000 rpm for 15 min to separate the bacterial cell mass. The clear supernatant was used to measure the decolorization at the absorbance maxima of the dye 600 nm. Uninoculated medium used as control. Percentage of decolorization was calculated as mentioned by Dave and Dave [18]. |
| Decolorization (%) = (I – F /I)* 100 |
| Where, I = Initial absorbance |
| F = Absorbance of decolorized sample. |
| Optimization of parameters for decolorization |
| The isolate was cultivated for 24 h in conical flasks containing 100 ml mineral salt medium (MSM) and was amended separately with 100 mg l-1 of RO16. Decolorization was studied with various carbon sources (1%) (sucrose, glucose, lactose, maltose and starch) and nitrogen sources (1%) (yeast extract, urea, peptone, ammonium sulphate and ammonium chloride) and at different dye concentrations (100, 250, 500, 750, 1000 mg/l), pH values (5-9) and temperature (20°C-50°C). Decolorization experiments were also carried out under shaking and static conditions. Growth was monitored spectrophotometrically at 600 nm. At periodic time intervals an aliquot of 5 ml of culture media was withdrawn, centrifuged at 5000 X g for 5 min in a refrigerated centrifuge (Dupont Sorvall RC-5B) to separate the bacterial cell mass. The supernatant was used for analysis of decolorization percentage and all the experiments were repeated in triplicates and mean value was taken for analysis. |
| FTIR analysis |
| The controls and samples obtained at varied time intervals were extracted with ethyl acetate were dried and mixed with KBr (1:20; 0.02 g of sample with KBr at a final weight of 0.4 g). The samples were then ground, desorbed at 60°C for 24 h and pressed to obtain IRtransparent pellets. The absorbance FT-IR spectra of the samples were recorded using an FT-IR Spectrum 2000 Perkin–Elmer spectrometer. The spectra were collected within a scanning range of 400-4000 cm-1. The FT-IR was first calibrated for background signal scanning with a control sample of pure KBr and then the experimental sample was scanned. The FT-IR spectrum of the control was finally subtracted from the spectra of the non-degraded and degraded dyes. |
| Liquid chromatography and mass spectroscopy analysis (LCMS analysis) |
| About 100 ml of Reactive orange (200 mg/L) containing MSM Media treated with the isolate and the purified enzymes was extracted with equal volume of ethyl acetate at various time intervals (0,24 h). The extract was evaporated in a vacuum evaporator (Buchii R 124, Germany) and used for LC-MS analysis. The powdery residue was then dissolved in acetonitrile (HPLC Grade). LC-MS analysis was performed using Finnigan model Mass Spectrometer (Thermo Electron Corporation, USA) using C-18 column from Waters. The cartridges were conditioned with pure acetonitrile, washed with deionized water (0.1% Formic acid) and the elution took place with 70% acetonitrile, containing 0.1% formic acid. The flow rate was 0.8 ml/ min. The ion trap detector with atmospheric pressure electro-spray ionization (APIESI) source was used for quantification in negative ionization mode. Operating conditions were dry with temperature of 325°C, Capillary voltage 3500 V, Nebulizer 14 psi, dry gas Helium 5.0 l/min. Ion trap full scan analyses were conducted from m/z 200-1400 with an upper full time of 300 minutes. The nebulizer gas flow and the curtain gas flow (Nitrogen gas) were set at 10 and 8 psi. The ion spray, orifice and ring voltage were set at +4800, 40, +70 V respectively. Instrumentation control of data acquisitions were performed with data analysis MS (X caliber, USA). |
| Phytotoxicity studies |
| Phytotoxicity tests were performed in order to assess the toxicity of the untreated and treated dye of Reactive orange 16. The ethyl acetate extracted products of RO16 degradation were dried and dissolved in10 ml sterile distilled water to make a final concentration of 1500 mg/l for phytotoxicity studies. The phytotoxicity study was carried out (at room temperature) on one kind of seeds commonly used in the Indian agriculture [19], Vigna mungo (10 seeds) by watering separately 10 ml sample of control RO16 and its degradation products (1500 mg/l) per day. Control set was carried out using distilled water at the same time. Germination (%) as well as the length of plumule and radical was recorded after 7 days (Data not shown). |
| Results and Discussion |
| Identification and phylogenetic position of bacterial isolate |
| A bacterial strain having remarkable Reactive orange 16 decolorization capacity was isolated from dye contaminated soil sample collected in and around the Theco silks Dyeing Industry, Kumbakonam, India. The identification of the strain was done on the basis of 16S rDNA gene sequences. Bacterial strain was identified as, Nocardiopsis alba. The sequence was deposited to Genbank (http://www.ncbi.nlm.nih.gov/Genbank). The nucleotide sequence analysis of the sequence was performed at BlastN site at NCBI server (http://www.ncbi.nlm.nih.gov/BLAST). The alignment of the sequences was performed by using Clustalw program V1.82 at European bioinformatics site (http://www.ebi.ac.uk/clustalw). To analyze the phylogenetic position, the Phylogenetic tree was constructed by the neighbour joining method using mega 5.1 software [17] (Figure 1a). The 16S rDNA gene sequences available in GenBank database with the accession number JQ897970. |
| Effect of physico-chemical factors on decolorization |
| The bacterial strain srm 1 2012 showed maximum dye decolorization at pH 7 (Table 2). At this optimum pH, the strain srm 1 2012 showed 95% of decolorization of Reactive orange 16. At pH 8, the strain srm 1 2012 showed 65% decolorization of Reactive orange 16. Whereas at pH 4.0, the strain showed only 25% dye decolorization respectively. Rate of decolorization decreased at lower pH (4-6) and at higher pH (8-9). Chan and Kuo reported that the neutral pH would be more favorable for decolorization of the azo dyes and is suitable for industrial applications [15]. The temperature effect on the decolorization of Reactive orange 16 was significant for the strain Nocardiopsis alba srm 1 2012. When the decolorization of the dye was tested for a wide range of temperatures from 20 to 50°C, it was observed that the increase in decolorization of Reactive orange 16 with increase in temperature and was optimum at 30°C. Further increase in the temperature increases the decolorization of dye up to 40°C and above this temperature a decreased dye decolorization was noticed (Table 2). To understand the effect of low temperature, room temperature and high temperature on the decolorization of dye, the assay was carried out at different temperature range from 20 to 50°C. The decrease in dye decolorization at high temperature can be attributed to the decline in microbial activity that led to the inactivation of the enzyme and eventually the loss of cells viability [6]. These results further showed that there is no thermal deactivation of decolorization activity under operational temperatures. Therefore, the strain srm 1 2012 could acclimatize to broad range of temperature of practical dyeing wastewater. The isolate is capable of decolorizing the dye up to 1000 mg/L. Lee et al. [20] stated that the increase in the concentration of the dye, the ability of the organism to decolorize decreased. In contrast to this, decolorization was not affected by increasing the dye concentration (100 mg-1000 mg/L). As indicated in previous studies [21], the chemical structure characteristics (e.g., resonance and inductive effects) and reactivity of dyes strongly determined color removal efficiency of bacterial decolorizers. At 100 mg of dye concentration 92% dye removal was observed; 83-85% decolorization was observed at concentration of 250 mg and 80% of dye was removed at concentration 500 mg. The time required for decolorization of 100 mg dye was 24 h, whereas it was 48 h for 250mg dye concentration. As concentration was increased (750 mg), the time required for decolorization varied from 48 to 72 h. When the dye concentration was as high as 1000 mg, almost 80% of the dye was removed after 72 h. This means that an acceptable high color removal can be achieved by the Nocardiopsis alba srm 1 2012 strain in an extensive range of azo dye concentrations. This decrease in decolorization efficiency at high dye concentrations above 1000mg may be due to the toxicity of the dye to bacteria and/or inadequate biomass concentration for the uptake of higher concentrations of dye [5]. The observed gradual decrease in decolorization might be due to the culture entering into the stationary phase and subsequently into the death phase, resulting in the inhibition of enzyme systems gradually. Nocardiopsis alba srm 1 2012 exhibited dye-decolorizing activity when incubated under stationary conditions, whereas agitated cultures grew well but showed negligible decolorization. To conform whether this decolorization was due to the microbial action or change in pH, the change in pH was recorded, which was in the range of 6-7 at static condition. This confirms the decolorization of dye was due to microbial action. Therefore, static conditions were adopted to investigate bacterial decolorization in the following experiments. |
| Earlier studies revealed that the dye could not be used as the sole carbon and nitrogen source by the organism and the organism required additional carbon and nitrogen sources to co-metabolize the dye. Hence, different carbon sources and nitrogen sources were evaluated for dye decolorization at 100 mg/ l dye concentration. The reduction of azo dyes depends on the presence and availability of cosubstrate because it acts as electron donor for the azo dyes reduction. It was found that isolate Nocardiopsis alba srm 1 2012 gave maximum decolorization with the combination of sucrose and peptone followed by glucose and yeast extract decolorization (above 70%).and with sucrose gave the highest decolorization (92%) (table 1). The organic nitrogen sources are considered essential medium supplements for the generation of NADH that act as electron donor for the reduction of azo dyes by microorganisms [22]. The results showed that there is a moderate increase in cell growth and dye decolorization, when both carbon and nitrogen sources were added together each at 0.1% concentration. |
| Identification of metabolic intermediates |
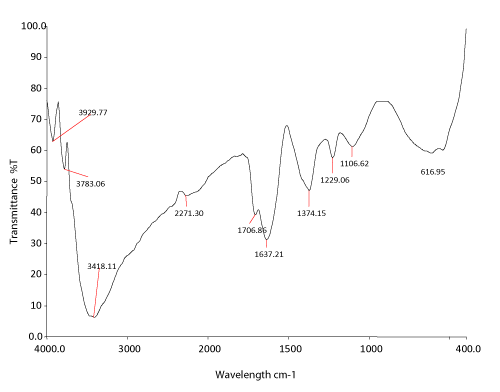
| FTIR analyses: The FTIR spectrum of a control dye and extracted metabolites (24 hours) by Nocardiopsis alba was compared. The spectrum of the control dye (Figure 2a) displayed a peak 3418.11 cm-1 for –NH stretching. The stretching between C N was reported at 2271.30 cm-1 and amide, 5-membered ring peak at 1706.86 cm-1. The peak at 1637.21 cm-1 showed carbonyl stretching vibration. Peak at 1374.15 cm-1 showed unsaturated nitrogen compounds. Peak at 1229.06 cm-1 showed S=O stretching vibrations. The peak at 1106.62 cm-1 indicates the aromatic nature. The peak at 616.95 cm-1 showed hydrocarbon chromophore-C-H bending. |
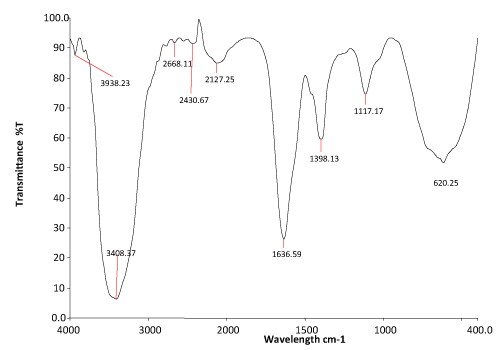
| The FTIR spectrum of 24 hours extracted metabolites of Nocardiopsis alba (Figure 2b) showed a significant change in positions of peak, when compared to the control dye spectrum. A new peak at 1636.59 cm-1 represented –N=N- stretching vibration. The C-H deformation showed at 1398.13 cm-1. The peak at 3408.37 cm-1 showed N-H stretching vibration. |
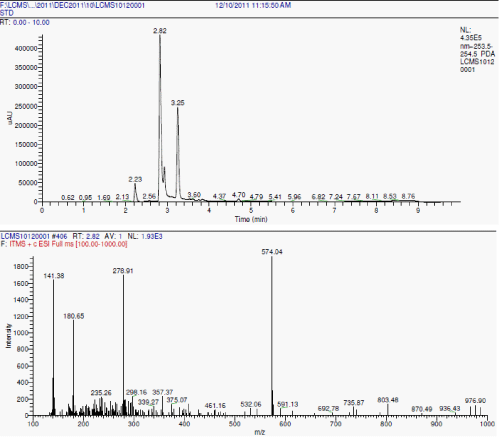
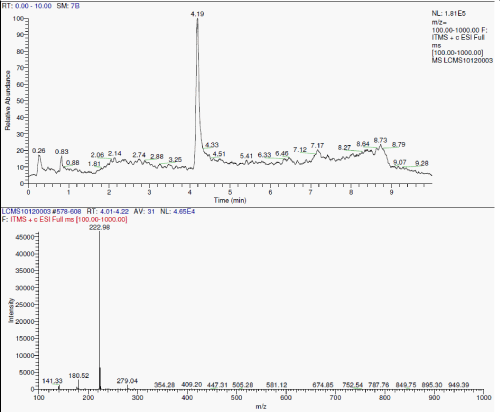
| LC-MS analyses: During the degradation there is asymmetric cleavage of azo bond in RO16 (Figure 3a and Figure 3b) resulting in formation of 1-amino-1-napthalene sulphonic acid, which was confirmed by the standard NIST library data, this is further, converted to aniline. While the naphthalene part of the dye was further biodegraded with opening of one ring, the formation of aldehyde as one of the intermediate is confirmed from the IR data. |
| Phytotoxicity studies |
| The results of the phytotoxicity studies showed no significant effect on the % germination of the Reactive orange 16 solution (1000 mg/l) wetted seeds; but it showed significant effect on the length of plumule and radicals. The germination percentage of Vigna mungo seeds were less with RO16 treatment when compared to treatment with its degradation products and water. The RO16 treatments affected the length of plumule and radical significantly, while there were no significant effects with the degradation products. The non-inhibition of germination with viable plumule and radical formation in the toxicity experiments of Reactive orange 16 (at 1000 mg) suggested that the dye was not only decolorized but also detoxified. This may be due to removal of aromatic amines by the organism. |
References
|
Tables and Figures at a glance
| Table 1 | Table 2 |
Figures at a glance
 |
 |
 |
 |
 |
| Figure 1 | Figure 2a | Figure 2b | Figure 3a | Figure 3b |
Relevant Topics
- Anaerobic Biodegradation
- Biodegradable Balloons
- Biodegradable Confetti
- Biodegradable Diapers
- Biodegradable Plastics
- Biodegradable Sunscreen
- Biodegradation
- Bioremediation Bacteria
- Bioremediation Oil Spills
- Bioremediation Plants
- Bioremediation Products
- Ex Situ Bioremediation
- Heavy Metal Bioremediation
- In Situ Bioremediation
- Mycoremediation
- Non Biodegradable
- Phytoremediation
- Sewage Water Treatment
- Soil Bioremediation
- Types of Upwelling
- Waste Degredation
- Xenobiotics
Recommended Journals
Article Tools
Article Usage
- Total views: 16147
- [From(publication date):
June-2012 - Dec 21, 2025] - Breakdown by view type
- HTML page views : 11234
- PDF downloads : 4913
