Research Article Open Access
Biodegradation of Acid Blue 113 Containing Textile Effluent by Constructed Aerobic Bacterial Consortia: Optimization and Mechanism
| C Valli Nachiyar1*, Swetha Sunkar1, G Narendra Kumar1, A Karunya1, PB Ananth1, P Prakash1 and S Anuradha Jabasingh2 | |
| 1Department of Biotechnology, Sathyabama University , Jeppiaar Nagar, Old Mamallapuram Road, Sholinganallur, Chennai 600 119, Tamil Nadu, India | |
| 2Department of Chemical Engineering, Sathyabama University, Jeppiaar Nagar, Old Mamallapuram Road, Sholinganallur, Chennai 600 119, Tamil Nadu, India | |
| Corresponding Author : | C Valli Nachiyar Department of Biotechnology Sathyabama University Jeppiaar Nagar, Old Mamallapuram Road Sholinganallur, Chennai 600 119 Tamil Nadu, India Fax: 91-44-24501270 Tel: 91-44-2450 3145 E-mail: vnachiyar@gmail.com |
| Received July 20, 2012; Accepted August 14, 2012; Published August 17, 2012 | |
| Citation: Nachiyar CV, Sunkar S, Kumar GN, Karunya A, Ananth PB, et al. (2012) Biodegradation of Acid Blue 113 Containing Textile Effluent by Constructed Aerobic Bacterial Consortia: Optimization and Mechanism. J Bioremed Biodeg 3:162. doi:10.4172/2155-6199.1000162 | |
| Copyright: © 2012 Nachiyar CV, et al. This is an open-a ccess article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. | |
Related article at Pubmed Pubmed  Scholar Google Scholar Google |
|
Visit for more related articles at Journal of Bioremediation & Biodegradation
Abstract
A bacterial consortium was constructed using five different bacterial strains isolated from the effluent with the ability to degrade Acid Blue 113, a diazo dye. These organisms were identified as Citrobacter freundii (2 strains), Moraxella osloensis, Pseudomonas aeruginosa using 16S rRNA analysis and Pseudomonas aeruginosa CLRI BL22. The consortium was found to degrade 90% of the dye by 22 h in 80% diluted textile effluent supplemented with glucose and ammonium nitrate. Optimization studies using Response Surface Methodology have confirmed that the degradation process was predominantly influenced by agitation and pH where as glucose was found to have negative effect. TLC analyses indicated the presence of metanilic acid and peri acid in 24 h sample which disappeared by 48 h. The GC-MS analysis has confirmed the presence of methyl salicylic acid, catechol and β-ketoadipic acid with the RT values of 7.71, 10.88 and 15.04 respectively confirming the complete degradation of Acid Blue 113.
| Keywords |
| Biodegradation; Acid blue 113; Citrobacter freundii; Moraxella osloensis; Pseudomonas aeruginosa; Pathway |
| Abbreviations |
| rRNA: Ribosomal Ribonucleic Acid; RT: Retention Time; rDNA: Ribosomal Deoxyribonucleic Acid; EMBL: European Molecular Biology Laboratory; DDBJ: DNA Databank of Japan; BLASTN: Basic Local Alignment Search Tool; FTIR: Fourier Transform Infrared; TLC: Thin Layer Chromatography; GC-MS: Gas Chromatography-Mass Spectroscopy; RSM: Response SuRface Methodology; CCD: Central Composite Design; Rf : Relative Front; CN: Consortia Number; TCA: Tricarboxylic Acid Cycle |
| Introduction |
| An estimated 120 crore people in the world today do not have access to safe drinking water. Water pollution due to industrial effluents and sewage mixing is a major cause of this malady. Colored effluent discharge from tanneries, textile and dye stuff industries to neighboring water bodies and waste water treatment systems is a major environmental hazard in this regard causing significant health concerns to environmental regulatory agencies. While colored organic compounds per se generally impart only a minor fraction of the organic load to waste water, their colour renders it aesthetically unacceptable. The colour of the industrial effluents is mainly due to the presence of azo dyes which are aromatics substituted with azo groups (-N=N-). Therefore, colour from the industrial effluents has become a subject of great scientific interest as evidenced from the flurry of research papers published in recent years in this niche segment [1-3]. Despite the existence of a variety of chemical and physical treatment processes, bioremediation of the effluents is seen as an attractive solution due to its reputation as low-cost, environment friendly and publicly acceptable treatment technology. |
| Microorganisms are frequently the sole means, biological or nonbiological, in detoxifying these synthetic chemicals by converting them into inorganic products. Microorganisms play a significant role in soils, water and sediments because of their ability to mineralize anthropogenic compounds [4,5]. Natural communities of microorganisms harbour an amazing physiological versatility and catabolic potential for the breakdown of an enormous number of organic molecules [6]. |
| Biodegradation using microorganisms is gaining importance as it is cost effective, environment friendly and produce less sludge [7]. The bioremediation process is mediated by suitable microbes (bacteria and fungi) which completely decolourise and mineralize the pollutants on site from the dye laden effluent. The advantages of such a decolourization and degradation (of azo dyes) process stems from the fact that it is an aerobic treatment system where toxic intermediates like aromatic amines produced in the effluent by abiotic and biotic means are completely mineralized by the bacteria themselves [8]. Several investigators [2,9-11] have investigated microbial biodegradation of azo dyes used in different industries and characterized the degradation products. While number of reports are available on dye degradation using single microorganism like bacteria [12,13], fungi [14-16], actinomycetes [17,18], very few reports are available that reveal the use of microbial consortia or with more than one organism [19,20] for degradation as they are found to be more effective for the treatment of effluent containing diverse group of dyes [21]. |
| Acid Blue 113 is a commercially important diazo dye used extensively in textile industry and in tannery. Degradation of this dye by chemical and biological means is gaining momentum which is evident from the number of reports available. Chemical methods of degradation concentrate mainly on the photocatalytic degradation using TiO2 [22,23] as well as nano TiO2 by Advanced Oxidation Processes [24,25] while microbial degradation of this dye was studied using Pseudomonas aeruginosa [26], Bacillus subtilis [27] and Kulyvera cryocrescens ATCC 33435 [28]. Even though these organisms were found to be highly efficient in brining about complete degradation of Acid Blue 113, we cannot solely depend on them for the treatment of dye containing effluent which warrants the construction of consortia. Therefore, an attempt has been undertaken to construct an aerobic bacterial consortium capable of degrading Acid Blue 113 in textile effluent. Further, optimization of culture conditions for the maximum degradation of this dye and identification of intermediate products have been carried out. |
| Materials and Methods |
| Chemicals |
| All the solvents used were of analytical grade, purchased from Merck Pvt. Ltd. The chemicals used for the preparation reagent, solutions and microbiological growth media were purchased from SISCO Research and Hi-Media Laboratories Pvt. Ltd, Mumbai, India. The dye Acid Blue 113 was purchased from Saujanya dye chemicals, Ahmedabad, India. |
| Effluent collection |
| Effluent samples were collected from Suntex processing mills, Gummidipoondi; Professional fabrics, Tirupur and Kafer Textile mills, Tirupur. |
| Microorganisms |
| The bacteria present in the effluent were isolated by serial dilution method and their dye degrading ability was checked by pour plate method on nutrient agar containing Navitan Fast blue S5R. All the bacterial cultures were maintained on nutrient agar slants and were used for biodegradation studies after preculturing in nutrient broth for 12 h. The bacterial cells were cultivated in nutrient broth for 24 h and the cells after centrifugation were resuspended in 20% glycerol and stored at –20°C as stock cultures. The purity of the glycerol stocks were checked on nutrient agar plates before sub culturing on nutrient agar slants for inoculum preparation. |
| Consortia development |
| The consortia was developed by aseptically transferring the 2% inoculum containing approximately 2 x 108 cells per ml of 12 h grown culture of each individual strains in different combinations in 250 ml Erlenmeyer flasks containing 50 ml of medium and 50 mg/l dye solution at optimized conditions [26]. Individual strains were inoculated with 2% (v/v) aliquots of 12 h grown culture, respectively, to maintain the same cell count in the pure culture and in the consortium. The bacterial strains in the consortia are identified by the standard 16S rRNA analysis by isolating and sequencing the 16S rDNA sequence followed by comparison with the public databases (Genbank, EMBL and DDBJ) and BLASTN sequence match routines. |
| Experimental design |
| Evaluation and optimization of culture medium was carried out using Response SuRface Methodology (RSM). RSM consists of a group of empirical techniques devoted to the evaluation of relations existing between a cluster of controlled experimental factors and the measured responses, according to one or selected criteria. A prior knowledge and understanding of the process and the process variables under investigation are necessary for achieving a more realistic model [29]. This design of experiment was done using Minitab 16. Based on the results obtained in preliminary experiments, glucose, agitation and pH were found to be major variables in the dye degradation [26]. Hence, these variables were selected to find the optimized condition for higher dye degradation using response suRface approach by central composite design. The range and the levels of the experimental variables investigated in this study are given in table 1. The central values (zero level) chosen for experiment design were agitation, pH and glucose. In developing the regression equation, the test factors were coded according to the equation. |
| Response = 0.082 + 0.011 * A - 6.194-003 * B + 4.883 - 003*C + 9.750 - 003*A*B - 0.016*A*C -0.014*B*C - 9.962 - 003* A2 - 8.725 - 003 * B2-0.013* C2 |
| A, B, C are the variables pH, glucose and agitation respectively. |
| Media and culture conditions |
| The screening studies for the dye degradation were carried out in 250 ml conical flasks containing 50 ml mineral salt media whose composition was described elsewhere [22] with 100 mg/l of dye. The medium was inoculated with 12 h old 2% inoculum containing approximately 2 X 108 cells and incubated in an incubator shaker at 200 rev min-1 at 30°C. Degradation of the Acid Blue 113 was monitored spectrophotometrically by reading the n-butanol extract of the culture medium at 585 nm [30]. |
| Degradation studies with the acid blue 113 containing effluent were carried out in 80% diluted textile effluent. 50 ml of textile effluent supplemented with 5 g/l glucose and 1 g/l ammonium nitrate was taken in 250 ml flask inoculated and incubated in orbitary shaker at 200 rev min-1. Degradation was followed as explained previously. |
| Identification of metabolites |
| 50 ml of effluent containing 100 mg l-1 of dye was inoculated with 2% inoculum and kept in the orbital shaker at 30°C. 5 ml samples were drawn at regular intervals and centrifuged at 12,000 rpm for 20 min. The culture supernatant was lyophilized and kept in a desiccator. TLC analyses for the breakdown products were done on fluorescent silica plates (Polygram Sil G/UV, 40 X 80 mm, Germany). For identification of metanilic acid, the solvent system used was Chloroform: Dichloromethane:Ethanol (5:5:5) [31] and for the identification of peri acid, the solvent system used was Chloroform: Ethanol (9:1) [32]. The compounds were identified by comparing their Rf values with that of standards. Fourier Transform Infrared analysis of the lyophilized sample was carried out in Perkin Elmer RX1 FTIR spectrophotometer. |
| The culture medium after the incubation period was centrifuged and the supernatant was extracted thrice, with equal volume of ethyl acetate, dried over anhydrous Na2SO4, and then the solvent was evaporated in a rotary evaporator. GC–MS analysis of the ethyl acetate extract was peRformed by using GC-MS-QP 2010 [SHIMADZU] MS spectrometer. The column used was VF-5 ms, 30 m x 0.250 mm dia with the film thickness of 0.25 μm and the column oven was programmed between 70 and 300°C at the rate of 10°C per minute with the injection temperature of 240°C. Mass spectra were recorded under scan mode in the range of 40–1000 m/z. Compounds were identified using WILEY8. LIB. |
| Results and Discussion |
| Dye degradation is a complex process that involves multiple steps. Eventhough, there are a number of bacteria which can bring about dye degradation efficiently, each one is specific for a particular group of dyes. But for the treatment of dye containing effluent they cannot be depended as the effluent will contain different groups of dyes. Under such condition, microbial consortia with organisms having different specificities can peRform the treatment of dye containing effluent more efficiently which is difficult or even impossible for individual strains or species [33]. |
| The isolates for the consortium developed were selected based on three factors, their ability to degrade the dyes efficiently (>90%), rapidity and their ability to degrade a wide variety of dyes. Further, many researchers have mentioned that a higher degree of biodegradation and mineralization can be expected when co-metabolic activities within a microbial community complement each other [34,35]. |
| Hence, the present work was undertaken to investigate the dye degradation potential of the developed aerobic bacterial consortium, isolated and identified from different textile effluents from different places with optimized physiological conditions to degrade Acid Blue 113, a commercially important dye used in textile industries. |
| Isolation, identification and construction of bacterial consortia |
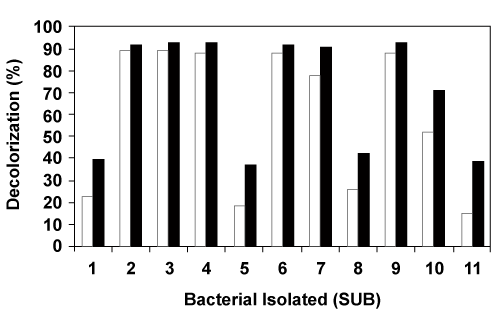
| Initial screening studies have resulted in the isolation of 10 bacterial cultures with dye degrading ability based on the zone of clearance on Acid Blue 113 containing nutrient agar plates. These 10 bacterial isolates along with Pseudomonas aeruginosa CLRI BL22 [31] designated as SUB 7 were checked for their ability to degrade Acid Blue 113 at 50 mgl-1 concentration. The results of degradation after 24 h and 48 h were given in Figure 1. All these 11 bacteria were used in different combinations in the construction of 30 consortia and their respective decolorization percentages were given in table 1. The difference in degradation individually and in combination with other bacteria may be due to the compatibility of these bacteria with one another, displaying synergistic activity in the process of degradation [34-36]. |
| Of these 30 consortia, CN 1 was found to degrade 97% of Acid Blue 113 at 100 mgl-1 in 80% diluted effluent supplemented with glucose and ammonium nitrate. These bacteria individually as well as in consortia were unable to degrade the dye in the absence of the above mentioned supplements as these are required for the production of biomass. This indicates that degradation of azo dye by these bacteria is dependent on carbohydrate metabolism and the dye degradation takes up a cometabolic pathway [26]. This finding is supported by previous reports in which the organisms bring about dye degradation in the presence of carbohydrate rather than using the dye as sole substrate [37,38]. The requirement of nitrogen source in the culture medium for maximum degradation may be explained on the basis of requirement of reduced nucleotides for the oxidative hydroxylation enzyme system [31]. Several investigations carried out earlier also reported the requirement of electron donating co-substrates such as glucose and yeast extract for the reduction of azo bonds by bacteria [36]. Safia Moosvi et al. [19] have also reported the importance of carbohydrate and nitrogen source in the form of glucose and yeast extract respectively for the maximum degradation of Reactive Violet 5 by bacterial consortium RVM11.1. The organisms constituting CN1 obtained from effluent characterized molecularly using 16S rRNA analysis were identified as Citrobacter freundii (Genbank ID: HQ324431), Moraxella osloensis (Genbank ID: HQ130446.1), Pseudomonas aeruginosa and Citrobacter freundii of different strains. |
| Citrobacter freundii was well known for its degrading ability and has also been studied specifically for its dye degrading ability [39]. The other organism constituting the consortia namely Pseudomonas aeruginosa, known as super bug, is an excellent candidate for xenobiotics biodegradation studies [26-40] and is also extensively studied for their ability to degrade a wide variety of azo dyes [41,42] including the pathways [11]. To our knowledge, no reports were available on the other constituent of the consortia Moraxella osloensis in biodegradation studies especially in dye degradation. |
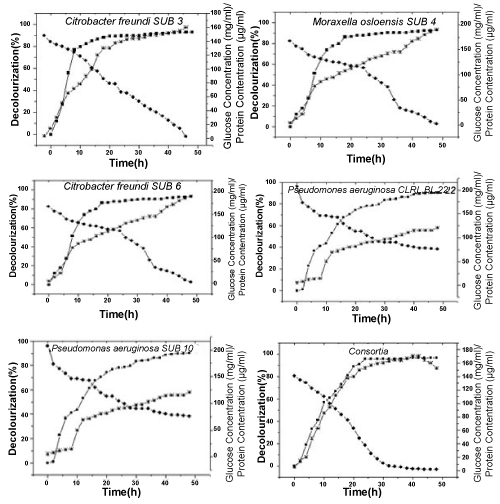
| The time course of degradation was followed in terms of degradation percentage, protein concentration (as an index of growth) and glucose utilization for individual as well as for consortia (Figure 2). All the individual organisms and also consortia started to degrade the dye in their late lag phase and 90% degradation was achieved after 26 h (Citrobacter freundii), 30 h (Moraxella oslensis), 28 h (Citrobacter freundii), 42 h (Pseudomonas aeruginosa) but the consortia could degrade 90% of the dye by 22 h reaching a maximum of 96% by 24 h. Biomass production was found to be proportional to glucose utilization. The reducing equivalents from glucose metabolism might have been involved in the dye degradation [43]. With reference to growth, the consortia reached its decline phase by 44 h when compared to the individual organisms which remained in their late stationary phase by that time. Khehra et al. [44] have reported a similar result for the decolorization of a group of azo dyes by consortia HM-4 containing Bacillus cereus, Pseudomonas putida, Pseudomonas fluorescens and Stenotrophomonas acidaminiphila. They have attributed the higher decolorization efficiency of consortium to the concerted activities of the constituent strains. The individual cultures might have transformed the dye to intermediates which acted on the redox mediators for efficient transfer of reducing equivalents from the strains, leading to an enhanced decolorization potential of the consortium. |
| Response suRface approach by central composite design |
| This is a very useful tool to determine the optimal level of factors and their interaction with each other. Our earlier study [11] reported the factors glucose, agitation and pH to be the significant components in the degradation of dye. Hence, CCD was used for further optimization using the above variables. Table 2 gives the variation levels of the factors which were subjected for the dye degradation, along with the average values of dye degradation showing the observed response and the predicted value calculated by using the mathematical model derived from the coefficients of the model. |
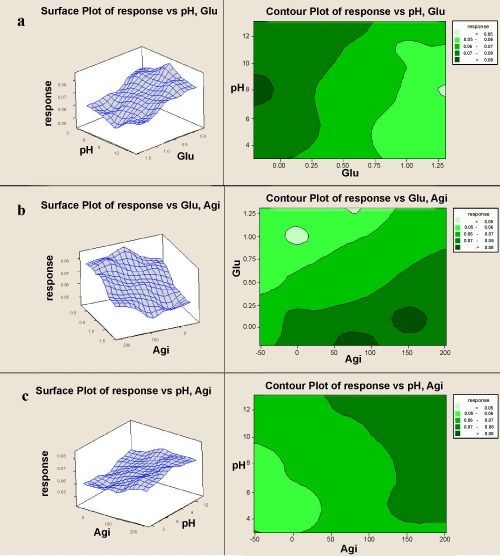
| The Fishers F-test with a very low probability value demonstrates a very high significance for the regression model. The goodness of fit of the model was checked by the determination coefficient (R). In this case, the value of the determination coefficient (R=0.983) indicates that only 5.90% of the total variations are not explained by the model. The value of the adjusted determination coefficient (Adj. R=0.980) is also very high, which indicates a high significance of the model. A higher value of the correlation coefficient (R=0.980) signifies an excellent correlation between the independent variables (Table 3). The fitted response for the above regression model was plotted and 3D graphs were generated for the pairwise combination of these factors while keeping the other one at its optimal levels for degradation process (Figure 3). The dye degradation was predominantly influenced by agitation and pH variation. Hence, these two factors were therefore the key factors which influence the degradation process. At higher concentrations of both factors, changes in the effect were observed. Glucose was also an important factor which had a negative approach towards the degradation. |
| Identification of intermediate compounds |
| TLC plates developed by chloroform: dichloromethane: ethanol has revealed a spot with Rf value 0.94 corresponding to metanilic acid. This spot was present up to 24 h of incubation and disappeared completely after 48 h. When chloroform: ethanol were used for the identification of peri acid, TLC plates have shown three distinct spots with Rf values of 0.57 (corresponding to peri acid), 0.25 and 0.11 in the sample incubated for 12 h. While the spots corresponding to Rf values of 0.25 and 0.11 disappeared completely after 24 h of incubation, there is a slight change in the Rf value of peri acid (0.47) indicating that the compound is undergoing certain chemical changes. This spot has also disappeared by 48 h. |
| The FTIR spectrum of the dye containing effluent taken at different incubation periods clearly showed a characteristic peak at 1635 cm-1 in sample taken at 0 h and 24 h corresponding to -N=Ngroup and this peak shifted to 11635 cm-1 at 72 h indicating that azo bond was undergoing certain chemical changes. The presence of new broad region between 2400 and 2500 cm-1 indicated the presence of carboxylic acid and ammonium ions suggesting partial mineralization. The presence of new peaks at 880 and 690 cm-1 associated without of plain bending vibrations of meta substituted benzenes implies that the products were undergoing irreversible chemical changes probably due to concomitant biodegradation and auto oxidation reaction of the products formed during the reductive dye degradation [45]. |
| The ethylacetate extract containing the degradation products of AB 113 was subjected to GC-MS analysis. The chromatogram and spectra are given in (Figure 4) in which three prominent peaks with RT values of 7.71, 10.88 and 12.11 can be seen. Mass spectra corresponding to the peak with RT value of 7.71 indicates a fragmentation pattern with m/z 120, 92 and 65 signals which confirm the presence of Methyl Salicylic acid. Methyl salicylic acid might have formed from Salicylic acid during the extraction process with ethyl acetate. Fragments with m/z 110 and 81 signals for RT value of 10.88 may be due to the presence of catechol. A small peak with the RT value of 15.04 gave mass spectra with m/z 115, 101 and a strong signal at 43 may be due to the presence of β-ketoadipic acid [46]. |
| The bacterial degradation of azo dyes was reported to start with the reduction of azo bond which is generally a non-specific and presumably extracellular process in which the reducing equivalents from an external electron donor (biologically or chemically generated) are transferred to the dye resulting in the production of aromatic amines [47-49]. In this case, metanilic acid and peri acid identified through TLC and 1,4-diaminonaphthalene (not identified). This reduction usually takes place in the absence of oxygen and many microorganisms are known to possess an enzyme azoreductase that may be oxygen insensitive [32,37,50-52]. These aromatic amines require the presence of oxygen for further breakdown as these oxygen molecules has to be inserted into such molecules which initiate other metabolic degradation [53]. The presence of salicylic acid confirmed by GC-MS clearly indicates that the naphthalene part of the dye must have undergone oxidative degradation followed by decarboxylation to catechol whose presence has also been identified in the GC-MS spectra [54]. The catechol might have come from the benzene part of the dye through oxidative deamination. This catechol must have taken up the ortho pathway reaction producing β-ketoadipic acid which enters the TCA cycle. These reports clearly indicated that this consortium has also taken up a degradation pathway similar to Pseudomonas aeruginosa in degrading Acid Blue 113 [11]. |
| Conclusion |
| Water pollution control is presently one of the major areas of scientific activity. Effluent discharge from textile and dyestuff industries to neighboring water bodies and wastewater treatment systems is currently causing significant health concerns to environmental regulatory agencies. Color removal, in particular, has recently become of major scientific interest, as indicated by the multitude of related research reports. As microbes, the poweRful weapon of bioremediation process, offer advantages of completely mineralizing the pollutants, investigations are warranted in the present research work focused on the possibilities of using constructed aerobic microbial consortium for decolorization and complete mineralization of azo dyes in dye laden textile effluent. This consortium was effective in degrading the azo dye Acid Blue 113 to β-ketoadipic acid which can be utilized by the bacteria for their normal metabolism. |
| Acknowledgements |
| The authors would like to thank the Department of Biotechnology, India for the financial support and also the Management of Sathyabama University for the encouragement and opportunity to carry out this work. |
References
|
Tables and Figures at a glance
| Table 1 | Table 2 | Table 3 |
Figures at a glance
 |
 |
 |
| Figure 1 | Figure 2 | Figure 3 |
Relevant Topics
- Anaerobic Biodegradation
- Biodegradable Balloons
- Biodegradable Confetti
- Biodegradable Diapers
- Biodegradable Plastics
- Biodegradable Sunscreen
- Biodegradation
- Bioremediation Bacteria
- Bioremediation Oil Spills
- Bioremediation Plants
- Bioremediation Products
- Ex Situ Bioremediation
- Heavy Metal Bioremediation
- In Situ Bioremediation
- Mycoremediation
- Non Biodegradable
- Phytoremediation
- Sewage Water Treatment
- Soil Bioremediation
- Types of Upwelling
- Waste Degredation
- Xenobiotics
Recommended Journals
Article Tools
Article Usage
- Total views: 16140
- [From(publication date):
September-2012 - Nov 29, 2025] - Breakdown by view type
- HTML page views : 11198
- PDF downloads : 4942
