Research Article Open Access
Catabolic Versatility of Sphingobium sp. Strain PNB Capable of Degrading Structurally Diverse Aromatic Compounds
| Madhumita Roy, Pratick Khara, Soumik Basu and Tapan K Dutta* | |
| Department of Microbiology, Bose Institute, Kolkata, India | |
| Corresponding Author : | Tapan K Dutta Department of Microbiology Bose Institute P-1/12 C.I.T. Scheme VII M Kolkata, India Tel: +913325693241 Fax: +913323553886 E-mail: tapan@bic.boseinst.ernet.in |
| Received: October 15, 2012; Accepted: November 22, 2012; Published: November 24, 2012 | |
| Citation: Roy M, Khara P, Basu S, Dutta TK (2013) Catabolic Versatility of Sphingobium sp. Strain PNB Capable of Degrading Structurally Diverse Aromatic Compounds. J Bioremed Biodeg 4:173. doi:10.4172/2155-6199.1000173 | |
| Copyright: © 2013 Roy M, et al. This is an open-a ccess article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. | |
Related article at Pubmed Pubmed  Scholar Google Scholar Google |
|
Visit for more related articles at Journal of Bioremediation & Biodegradation
Abstract
Sphingobium sp. strain PNB isolated from municipal waste-contaminated soil had already been described to
possess catabolic potential of assimilating phenanthrene. Apart from phenanthrene, strain PNB was found to be capable of utilizing naphthalene and biphenyl singly, as sole carbon and energy sources. Based on chromatographic and spectrophotometric analyses, pathways for the degradation of naphthalene and biphenyl were revealed in strain PNB. Anthracene, on the other hand, was found to be partially metabolized, furnishing 9,10-anthraquinone and 3-hydroxy-2-naphthoic acid, which appeared as the dead-end products. In addition, strain PNB was observed to co-metabolize fluoranthene, acenaphthene, benz[a]anthracene, pyrene and benzo[a]pyrene, in presence of phenanthrene indicating metabolic robustness of the strain, which may serve as a candidate organism in the bioremediation of Polycyclic Aromatic Hydrocarbons (PAHs)-contaminated environment.
| Keywords |
| Polycyclic aromatic hydrocarbons; Biphenyl; Biodegradation; Co-metabolism; Sphingobium |
| Introduction |
| Environmental deposit of Polycyclic Aromatic Hydrocarbons (PAHs), having numerous structural configurations is largely due to anthropogenic activities such as crude oil refining, fossil fuel combustion, waste incineration and automobile usage [1-3]. Growing environmental concern about PAHs is primarily because of their biological toxicity, which includes cytotoxicity, carcinogenicity, genotoxicity, and/or mutagenicity [4-6]. Bacterial degradation represents the primary route of transformation and removal of PAHs in the environment, and serves as a promising alternative to physicochemical treatments for the ecological recovery of PAHcontaminated sites. Apart from biodegradation, biotransformation is another major environmental process affecting the fate of PAHs, in both terrestrial and aquatic ecosystems [7]. Microorganisms once co-metabolically transform recalcitrant or more toxic compounds to less toxic or more bioavailable form; they can be metabolized by other microorganisms present at the contaminated sites. |
| Studies have shown that the Low-Molecular-Weight PAHs (LMW PAHs) containing less than four fused benzene rings such as naphthalene and phenanthrene are readily biodegraded, with the exception of poorly water soluble anthracene [2,8-12]. Whereas, the High-Molecular-Weight PAHs (HMW PAHs) containing four or more fused benzene rings such as fluoranthene, chrysene, pyrene, benz[a] anthracene, benzo[a]pyrene, etc. are generally recalcitrant to microbial attack, and thus contributing their persistence in the environment [13]. In addition, the success of most PAHs bioremediation projects has been shown to be limited by failure to remove the HMW PAHs [14,15], and is further compounded by the existence of PAHs in complex mixture [16]. However, studies have shown that HMW PAHs may be biodegraded via co-metabolism, using LMW PAHs or PAH degradative pathway intermediates as carbon and energy sources [17,18]. |
| Therefore, for bioremediation to be an effective tool for the cleanup of PAHs-contaminated soils, greater understanding of the bacterial catabolic machinery, vis-à-vis degradability of large spectrum of PAHs is most important. Among the wide phylogenetic spectrum of microorganisms, sphingomonads, a major group of PAHdegrading bacteria present in various contaminated sites are becoming increasingly important in environmental bioremediation. This is primarily because Sphingomonadaceae are largely reported to harbor versatile metabolic pathways, suggesting that the species belonging to this family have the ability to adapt more quickly and more efficiently, for the degradation of new compounds in the environment than members of other bacterial genera [19,20]. Previously, a phenanthrene degrading bacterial isolate belonging to the genus Sphingobium was reported from this laboratory, describing the degradation pathways in the assimilation of phenanthrene [21]. In the present study, the catabolic versatilities of strain PNB capable of utilizing, and/or cometabolizing other aromatics including HMW PAHs was reported. |
| Materials and Methods |
| Chemicals |
| Phenanthrene, naphthalene, anthracene, biphenyl, acenaphthene, pyrene, benzo[a]pyrene, benz[a]anthracene, fluoranthene, 3-hydroxy- 2-naphthoic acid, benzoic acid and salicylic acid were purchased from Sigma-Aldrich (Milwaukee, WI). Unless stated otherwise, all other chemicals and reagents used in this study were of analytical grade, and used without further purification. |
| Organism and culture conditions |
| Sphingobium sp. strain PNB was isolated in this laboratory [21], from municipal waste contaminated soil by enrichment with phenanthrene in Mineral Salts Medium (MSM). Cells were grown in 250 ml Erlenmeyer flasks containing 50 ml MSM, supplemented with solid naphthalene (0.5 g/l) or biphenyl (1 g/l) individually as sole carbon and energy sources, by incubating them at 28°C on a rotary shaker (180 rpm) for different periods of time. For growth with anthracene (0.5 g/l), culture medium was supplemented with yeast extract (0.2% w/v). For resting cell incubations, cells grown individually on anthracene supplemented with yeast extract, biphenyl or naphthalene were harvested in the late log phase by centrifugation (8000×g, 10 min), washed twice with equal volume of potassium phosphate buffer (50 mM, pH 7.0), and finally resuspended in the same buffer to give an optical density (OD660) of 1.0. Anthracene, naphthalene or biphenyl (1 g/l) were added individually to washed-cell suspensions, and incubated at 28°C for different periods of time, up to 48 h. For co-metabolic study in binary substrate system, 125 ml of Erlenmeyer flasks containing 25 ml MSM was supplemented with phenanthrene (0.5 g/l) as growth substrate, along with acenaphthene, fluoranthene, pyrene, benz[a]anthracene (0.5 g/l each) and benzo[a] pyrene (0.1 g/l) individually as co-substrate. Cultures were incubated aerobically up to 15 days in the dark at 28°C, with agitation at 180 rpm to monitor growth of the strain PNB, vis-à-vis biodegradation of the PAHs. Parallel substrate controls have also been incubated to assess abiotic loss of the PAHs. Unless stated otherwise, each experimental set was performed in triplicate. Solid media contained 2% (w/v) agar (HiMedia, India). |
| Analytical methods |
| After incubation, spent cultures and washed-cell suspensions containing unconverted substrates and metabolic intermediates were extracted thrice with equal volume of ethyl acetate, after acidification to pH 1.5-2.0 by 6 N hydrochloric acid. In case of biodegraded samples of binary substrates, organic extracts were spiked individually with 0.1 mg equivalent of n-hexadecane (as external standard), dissolved in ethyl acetate. The organic extracts were dried over anhydrous sodium sulfate and evaporated under reduced pressure. Metabolites in the dried extract were kept in the dark prior to analysis. |
| Metabolites obtained from spent cultures and resting cell suspensions were resolved by Thin Layer Chromatography (TLC) on silica gel GF254 (Merck Limited, India) plates, using benzene: acetic acid (97:3) v/v as the solvent system. Resolved metabolites were detected by a UV lamp at 254 or 365 nm. The identity of resolved product(s) was occasionally determined, by comparison with the TLC profile of reference compounds developed identically. In addition, for the characterization of metabolites, High Performance Liquid Chromatography (HPLC) was done using a Shimadzu model LC20-AT pump system (Shimadzu Corp., Kyoto, Japan), equipped with a diode array model SIL-M20A detector and an analytical Phenomenex C18 reverse-phase column (Phenomenex Inc., Torrance, California, USA), attached to a model SIL-20A autosampler. Biodegraded products were eluted with a programmed gradient of solvent system, at a flow rate of 1.0 ml/min and detected at 254 nm, along with diode array analysis. The mobile phase was a 45 min linear gradient from 50% (v/v) to 95% (v/v) aqueous methanol, with hold at 95% (v/v) aqueous methanol for 10 min, followed by 95% (v/v) to 50% (v/v) aqueous methanol over 5 min. Metabolites were identified by comparing their retention times and UV-visible absorbance spectra, with those of the authentic compounds analyzed under the same set of conditions. On the other hand, Thermo Scientific model TraceGC Ultra column (Thermo Fischer Scientific), with a model Polaris Q mass spectrometer, equipped with a 30 m×0.25 mm (0.25-μm film thickness) DB-5MS capillary column, was used for analysis of various metabolites by Gas Chromatography-Mass Spectrometry (GC-MS). The ion source was kept at 230°C, and both the inlet temperature and the transfer line temperature were kept at 280°C. The temperature program comprised 2 min hold at 70°C, increase to 200°C at 10°C/min, followed by hold for 1 min at 200°C, further increase to 325°C at 5°C/min-1, and 15 min hold at 325°C. The injection volume was 1 μl, and the carrier gas was helium (1 ml/min). The mass spectrometer was operated at electron ionization energy of 70 eV. Metabolites were identified by comparing their retention time and mass fragmentation pattern, with those observed for authentic compounds. However, for commercially unavailable authentic compounds, structures were suggested based on mass fragmentation patterns and related information available in literature. In co-metabolism studies, unconverted PAHs were quantified based on n-hexadecane as the external standard. |
| Preparation of cell-free extracts and enzyme assays |
| Cells grown individually on biphenyl and naphthalene, as described above, were harvested at mid-log phase by centrifugation at 8000×g for 10 min. The pellets obtained were washed two times with phosphate buffer (50 mM, pH 7.0), finally resuspended in the same buffer, and were loaded into a pre-cooled french press, Constant Cell Disruption System, One Shot Model (Constant System Ltd. United Kingdom), fitted with a 8.0 ml cell, followed by lysis at 30,000 psi for two cycles. Extracts were subjected to centrifugation at 20,000×g for 40 min at 4°C, and the resultant clear supernatant was used as cell-free enzymes for further studies. Cell-free extracts were examined for the presence of catechol 1,2-dioxygenase, catechol 2,3-dioxygenase and salicylaldehyde dehydrogenase. Assays were performed by monitoring disappearance of substrates and appearance of products by UV-visible spectrometry, on the basis of spectral properties reported earlier on these enzyme systems [22-24]. Spectral changes were recorded at 25°C with Cary 100 Bio UV-visible spectrophotometer (Varian Australia Pty Ltd.), using 1 cm path length quartz cuvettes. Data were analyzed with the Varian Cary Win UV Scan application software. Protein was measured by using the Lowry method, with BSA as the standard [25]. |
| Results and Discussion |
| Organism and growth |
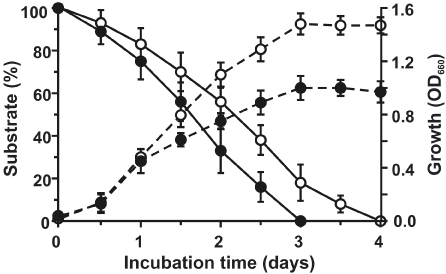
| Strain PNB isolated from municipal waste-contaminated soil sample, capable of degrading phenanthrene as sole sources of carbon and energy, was characterized as Sphingobium sp., and the metabolic pathways involved in the degradation of phenanthrene was reported earlier [21]. Apart from phenanthrene, Sphingobium sp. strain PNB was found to be capable of utilizing naphthalene and biphenyl individually as sole sources of carbon and energy. Strain PNB can also form clearing zones on anthracene, biphenyl and naphthalene coated agar plates, similar to that on phenanthrene. Blue colored colonies were observed when indole was sprayed onto cells grown on LB agar plates, and induced individually in presence of phenanthrene, anthracene and naphthalene (data not shown). Utilization of naphthalene and biphenyl individually as growth substrates by strain PNB was further confirmed by their removal from respective substrate-mineral salt medium, with a corresponding increase in bacterial biomass; while complete degradation of naphthalene and biphenyl was achieved within 3 and 4 days respectively (Figure 1). Among the possible metabolic pathway intermediates, salicylic acid and benzoic acid supported growth of strain PNB, when provided individually as sole carbon sources. However, growth was not observed with 3-hydroxy-2-naphthoic acid, phthalic acid, protocatechuic acid, gentisic acid or catechol. |
| Identification of metabolites |
| TLC analysis of the ethyl acetate extracts of the spent medium and resting cell incubation cultures of anthracene, biphenyl and naphthalene, indicated the presence of various polar metabolites in the reaction mixture (data not shown). Based on the comparison of the Rfs and UV-fluorescence properties of standard compounds, 3-hydroxy- 2-naphthoic acid (Rf 0.48, bluish black, fluorescent), was identified as the major intermediate of the anthracene degradation pathway. Salicylic acid (Rf 0.79, blue, fluorescent), salicylaldehyde (Rf 0.42, bluish fluorescent) and trace of catechol (Rf 0.21, black, non-fluorescent) were observed as metabolic intermediates in the degradation of naphthalene. On the other hand, TLC profile indicated the presence of benzoic acid (Rf 0.53, blue fluorescent) and trace of catechol as the metabolites in the degradation of biphenyl. |
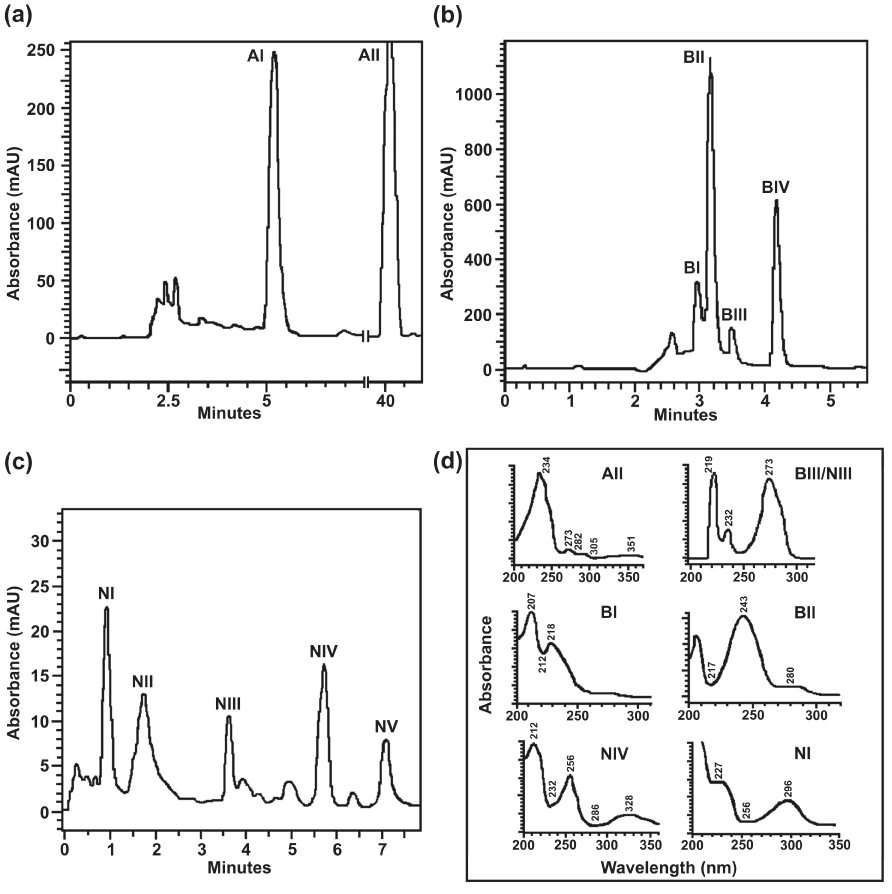
| Results of TLC analyses were substantiated by data obtained from HPLC analyses of metabolites obtained from resting cell incubations of individual compounds. HPLC analysis of the ethyl acetate extract of the resting cell incubation-mediated transformed products of anthracene with anthracene (supplemented with year extract) grown culture, using a reverse-phase C18 column, indicated the presence of 3-hydroxy-2-naphthoic acid (AI), unresolved products eluted around 2.5 min and unutilized anthracene (AII) (Figure 2a). While metabolites obtained from resting cell incubation of biphenyl, grown on the same substrate furnished benzoic acid (BI), unidentified product (BII), catechol (BIII) and unconverted biphenyl (BIV) (Figure 2b). On the other hand, salicylic acid (N1), co-eluted unidentified products (NII), catechol (NIII), salicylaldehyde (NIV) and unutilized naphthalene (NV) were resolved by HPLC analysis of the products obtained from resting cell incubation of naphthalene with naphthalene-grown culture (Figure 2c). Few other minor peaks obtained from resting cell incubation of naphthalene could not be identified. UV-visible spectra of the identified products and major unidentified product were shown in figure 2d. Metabolites were identified by comparing the retention times, co-elution profile and UV-visible spectra (obtained from diode array analysis), with that of the authentic compounds analyzed under identical conditions. It may be mentioned that the products that could not be identified from HPLC analyses are primarily due to unavailability of appropriate pathway intermediates as reference compounds. |
| Biodegraded products obtained from the organic extracts of the spent cultures and resting cell incubations (48 h) with anthracene, naphthalene and biphenyl were analyzed by GC-MS, and the results are summarized in table 1. GC-MS data correlated well with those obtained from TLC and HPLC analyses. Apart from that, 6,7-benzocoumarin, 1-methoxy-2-hydroxyanthracene were detected in the anthraceneincubated culture, and the mass spectral data were found to be similar to those reported earlier [26,27]. In addition, the organic extract of resting cell incubation with anthracene, indicated the presence of trace amount of a metabolite that consisted of molecular ion at an m/z of 212, and characteristic fragment ions at m/z values of 194, 168, 166, 165 and 140, accounting for anthracenedihydrodiol (possibly cis-1,2- dihydroxy-1,2-dihydroanthracene), by comparison to data reported earlier [27]. However, the position of hydroxyl substitutions in this metabolite could not be confirmed further at this stage due to its low level of production. Moreover, 9,10-anthraquinone was identified as one of the anthracene-degraded metabolites. It is believed that apart from 1,2-dioxygenation of anthracene, strain PNB is supposed to metabolize anthracene by dioxygenating at 9 and 10 positions, forming anthracene 9,10-dihydrodiol, which may be converted non-enzymatically to the dead-end product 9,10-anthraquinone, as reported in strain PYR1 [27]. In the degradation of biphenyl, apart from others, GC-MS analysis indicated the presence of 2,3-dihydroxybiphenyl, a metabolite obtained from biphenyl, by the actions of biphenyl 2,3-dioxygenase and biphenyl 2,3-dihydrodiol dehydrogense, which are the peripheral enzymes in the biphenyl metabolic pathway, and have been described in several studies. GC-MS analyses of the organic extract obtained from spent culture supernatant and resting cell incubation of naphthalene with naphthalene-grown cells further confirmed salicylaldehyde, salicylic acid and catechol, as the metabolic intermediates. |
| Enzyme assay |
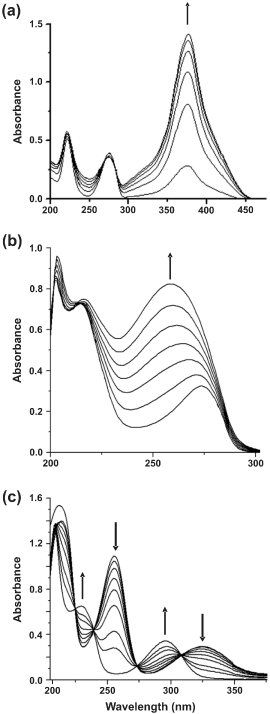
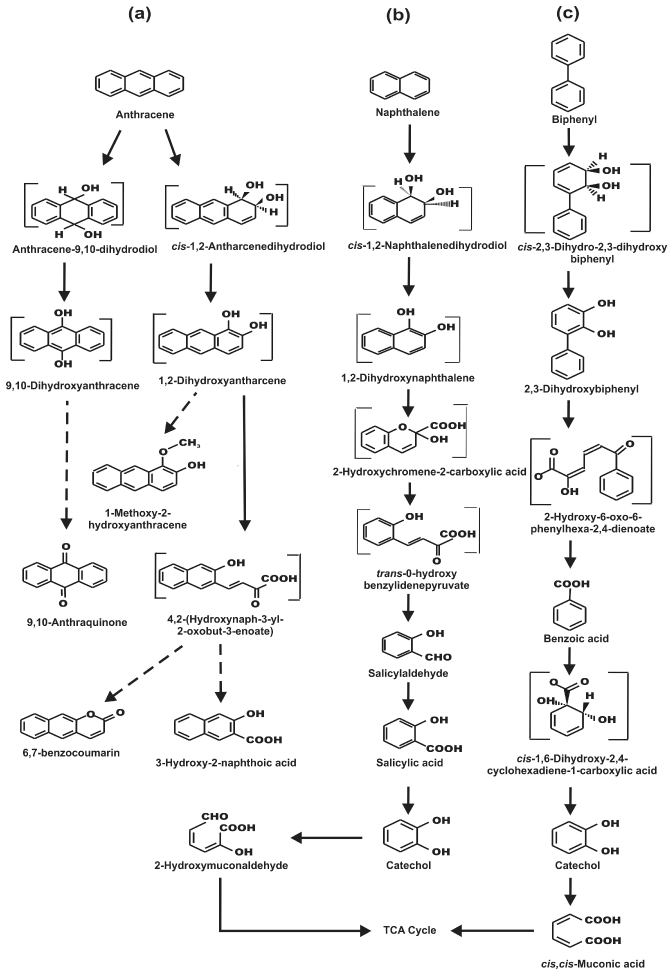
| Metabolic pathways involved in the degradation of naphthalene and biphenyl were further examined by the cell-free extract-mediated transformations of metabolites. In biphenyl degradation pathway, catechol was found to be metabolized by ortho cleavage pathway, while in naphthalene, catechol was observed to be metabolized by meta cleavage pathway. Appearance of a yellow-colored product with λmax at 374 nm is due to the formation of 2-hydroxymuconaldehyde acid (Figure 3a), indicative of the meta cleavage of catechol by catechol 2,3-dioxygenase (specific activity 118 nmol min-1 (mg protein)-1) [24], while a shift of λmax from 275 nm to 260 nm for catechol is signifying the ortho cleavage of catechol by catechol 1,2-dioxygenase (specific activity 35 nmol min-1 (mg protein)-1) (Figure 3b), in the formation of cis, cis-muconic acid [23]. In addition, cell-free extract-mediated NAD+- dependent conversion of salicylaldehyde to salicylic acid was observed by cells grown on naphthalene, where an increase in the absorbance at 296 nm and simultaneous decrease in absorbance at 254 and 330 nm was monitored (Figure 3c), due to the formation of salicylic acid [22]. Because salicylaldehyde itself has absorbance around 340 nm, specific activity of salicyladehyde dehydrogenase, based on the formation of NADH (λmax at 340 nm) from NAD+ could not be measured during this transformation. However, none of these enzyme activities could be detected in the cell-free extract obtained from succinate-grown cells,indicating inducible nature of these catabolic enzymes. Based on the identification of metabolites by TLC, HPLC, GCMS and UV-visible spectral studies, metabolic pathways of degradation of naphthalene, anthracene and biphenyl by strain PNB have been elucidated (Figure 4). |
| Co-metabolic potential of strain PNB |
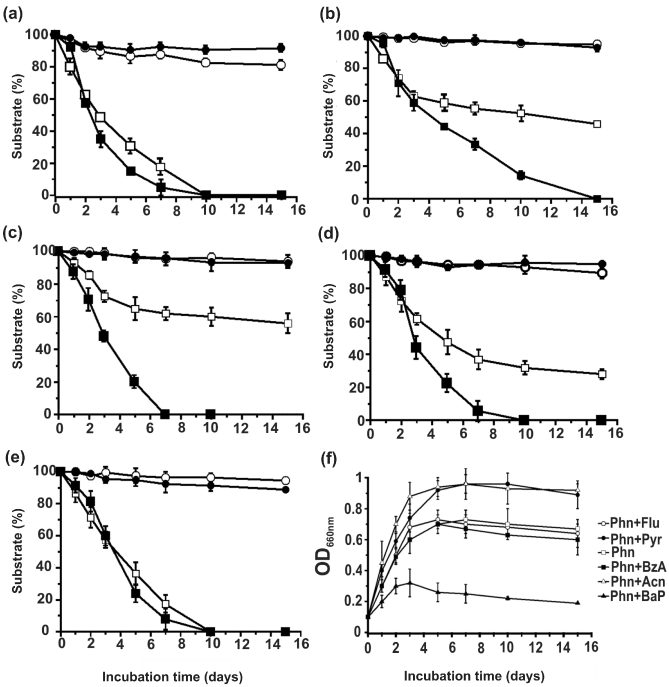
| Acenaphthene, fluoranthene, pyrene, benz[a]anthracene and benzo[a]pyrene, which did not serve individually as growth substrates, were observed to be degraded co-metabolically in the presence of phenanthrene by the strain PNB. Co-metabolism of all these PAHs (0.5 g/l, except 0.2 g/l for benzo[a]pyrene) individually, in presence of phenanthrene (0.5 g/l) as binary substrate system is shown in figure 5. Among all the PAHs, complete degradation was observed with acenaphthene and pyrene, during 10 days of incubation (Figure 5a and 5e), followed by fluoranthene (65% degradation within 14 days) (Figure 5d). Although benzo[a]anthracene was least degraded (40%) among the co-metabolic PAHs, but it was observed that the presence of benzo[a]anthracene had no adverse effect in the degradation kinetics of phenanthrene in this binary substrate system (Figure 5c). However, benzo[a]pyrene even at a much lower concentration adversely affected the phenanthrene degradation kinetics, and a 50% degradation of benzo[a]pyrene was achieved during 15 days of incubation (Figure 5b). Figure 5f shows the growth profile of strain PNB in individual binary substrate systems, in comparison to growth profile with phenanthrene alone, as the sole carbon and energy sources. With the same concentration of phenanthrene, higher growth profile was observed only with acenaphthene-phenanthrene and pyrene-phenanthrene binary substrate systems, while fluoranthene-phenanthrene and benzo[a]anthracene-phenanthrene binary substrate systems showed almost similar growth profile, in comparison to phenanthrene alone. However, a drastically lower growth was observed with benzo[a] pyrene-phenanthrene binary system, indicating possible growth inhibitory factor generated in this binary substrate metabolic process, which was also reflected in their degradation profiles (Figure 5b). |
| Thus, it appears that few individually non-utilizable substrates were utilized at least partially by the strain PNB for its growth during cometabolism, in presence of phenanthrene, indicating enzymes induced by one substrate allow the organism to derive additional carbon and energy from a second compound that cannot serve as a growth substrate alone. Such a situation of co-metabolic analysis has been reported in the degradation of chlorobenzoates [28]. In natural ecosystems, a variety of related molecules are present at low concentrations, and it would be inefficient to evolve a specific induction system for the degradation of each compound, rather induction of a nonspecific enzyme by one component of a mixture would facilitate the organism to derive carbon and energy from a suite of related compounds [29]. In this context, sphingomonads has come out as a versatile group for their exceptionally diverse degradation capacity towards various natural and xenobiotic compounds [30,19,20]. It appears that the degradative genes in sphingomonads are complexly arranged and the genes for the individual degradative pathways are scattered through several gene clusters [19]. It is believed that the presence of multiple ring-hydroxylating, as well as ring cleavage dioxygenases, and/or broad substrate range dioxygenases in sphingomonads make them more quickly, and/or more efficiently adapted for the degradation of new compounds in the environment than members of other bacterial genera, signifying strains belonging to spingomonads are ideal for bioremediation. |
| In conclusion, the present study describes a Sphingobium sp. that can utilize a number of LMW PAHs and biphenyl individually as sole carbon sources, elucidating metabolic pathways in the complete degradation of naphthalene and biphenyl, apart from a partial pathway in the metabolism of anthracene. In addition, studies on co-metabolic potential of the strain towards a number of HMW PAHs, have distinctly qualified the catabolic robustness of strain PNB. However, metabolic pathways and their interactions need to be studied in greater depth, which will further contribute knowledge towards the understanding of complex induction and regulation pattern in strain PNB, in particular, and sphingomonads, in general. |
| Acknowledgements |
| The financial support of the Department of Biotechnology (DBT, grant BT/ PR9138/BRB/10/542/2007), Government of India, and Bose Institute, Kolkata, India is gratefully acknowledged. Madhumita Roy, Pratick Khara and Soumik Basu are also grateful to the Council of Scientific and Industrial Research, Government of India, DBT and Bose Institute, respectively, for awards of Junior and/or Senior Research Fellowships. |
References
|
Tables and Figures at a glance
| Table 1 |
Figures at a glance
 |
 |
 |
 |
 |
| Figure 1 | Figure 2 | Figure 3 | Figure 4 | Figure 5 |
Relevant Topics
- Anaerobic Biodegradation
- Biodegradable Balloons
- Biodegradable Confetti
- Biodegradable Diapers
- Biodegradable Plastics
- Biodegradable Sunscreen
- Biodegradation
- Bioremediation Bacteria
- Bioremediation Oil Spills
- Bioremediation Plants
- Bioremediation Products
- Ex Situ Bioremediation
- Heavy Metal Bioremediation
- In Situ Bioremediation
- Mycoremediation
- Non Biodegradable
- Phytoremediation
- Sewage Water Treatment
- Soil Bioremediation
- Types of Upwelling
- Waste Degredation
- Xenobiotics
Recommended Journals
Article Tools
Article Usage
- Total views: 14926
- [From(publication date):
January-2013 - Dec 20, 2025] - Breakdown by view type
- HTML page views : 10195
- PDF downloads : 4731
