Research Article Open Access
Metabolism and Preferential Utilization of Phenylacetic acid and 4-Hydroxyphenylacetic Acid in Pseudomonas putida CSV86
| Rahul Shrivastava1, Hemant Purohit2 and Prashant S Phale1* | |
| 1Department of Biosciences and Bioengineering, Indian Institute of Technology-Bombay, Powai, Mumbai 400076, India | |
| 2Environmental Genomics Unit, National Environmental Engineering Research Institute, Nagpur-440020, India | |
| Corresponding Author : | Dr. Prashant S Phale Department of Biosciences and Bioengineering Indian Institute of Technology-Bombay Powai, Mumbai 400076, India Tel: +91-22-2576 7836 Fax: +91-22-2572 3480 E-mail: pphale@iitb.ac.in |
| Received March 29, 2011; Accepted June 20, 2011; Published June 21, 2011 | |
| Citation: Shrivastava R, Purohit H, Phale PS (2011) Metabolism and Preferential Utilization of Phenylacetic acid and 4-Hydroxyphenylacetic Acid in Pseudomonas putida CSV86. J Bioremed Biodegrad 2:120. doi:10.4172/2155-6199.1000120 | |
| Copyright: © 2011 Shrivastava R. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. | |
Related article at Pubmed Pubmed  Scholar Google Scholar Google |
|
Visit for more related articles at Journal of Bioremediation & Biodegradation
Abstract
Metabolic pathways for phenylaceticacid (PA) and 4-hydroxyphenylacetic acid (4-HPA) from Pseudomonas putida CSV86 were elucidated. PA grown strain CSV86 cells showed higher oxygen-uptake on PA as compared to hydroxyphenylacetic acids. Detection of phenylacetyl-CoA ligase and absence of 4-hydroxyphenylacetic acid hydroxylase (4HPAH) and 3,4-dihydroxyphenylacetic acid-dioxygenase (3,4-DHPADO) activity supports this observation. 4-HPA grown cells showed oxygen-uptake on 4-HPA and 3,4-dihydroxyphenylacetic acid (3,4-DHPA) but showed significantly lower respiration rates on 2,5-dihydroxyphenylacetic acid and PA. Detection of 4HPAH and 3,4-DHPADO activities support the respiration data. Metabolic studies indicate that strain CSV86 metabolizes PA via phenylacetyl-CoA while 4-HPA via 3,4-DHPA pathway. Glucose grown cells showed lower activity of enzymes and respiration on PA, 4-HPA and 3,4-DHPA, suggesting that pathways are inducible. When grown on double carbon sources such as PA or 4-HPA plus glucose, CSV86 cells showed diauxic growth pattern with oxygen uptake on PA, 4-HPA and 3,4-DHPA in the first log-phase which was reduced during the second log-phase with increase in the respiration rate on glucose. This observation was supported by detection of high 4HPAH and 3,4-DHPADO activity in the first log-phase and glucose-6-phosphate dehydrogenase activity in the second log-phase. These results suggest that strain CSV86 utilizes PA and 4-HPA preferentially over glucose.
| Introduction |
| Conventional pathway for the degradation of aromatic compounds by bacteria involves oxidation and hydroxylation reactions catalyzed by specific mono- or dioxygenases yielding central aromatic intermediates like catechol, protocatechuate, gentisate, homoprotocatechuate, homogentisate etc [1]. However, recent studies suggest the existence of an alternative pathway for the degradation of aromatic compound via CoA-thioesters, which is also referred as 'aerobic hybrid pathway' [2]. The best studied example is benzoate degradation in Azoarcus evansii where all the proposed intermediate metabolites are CoA-thioesters and the ring-cleaving reaction does not involve the molecular oxygen [2-4]. |
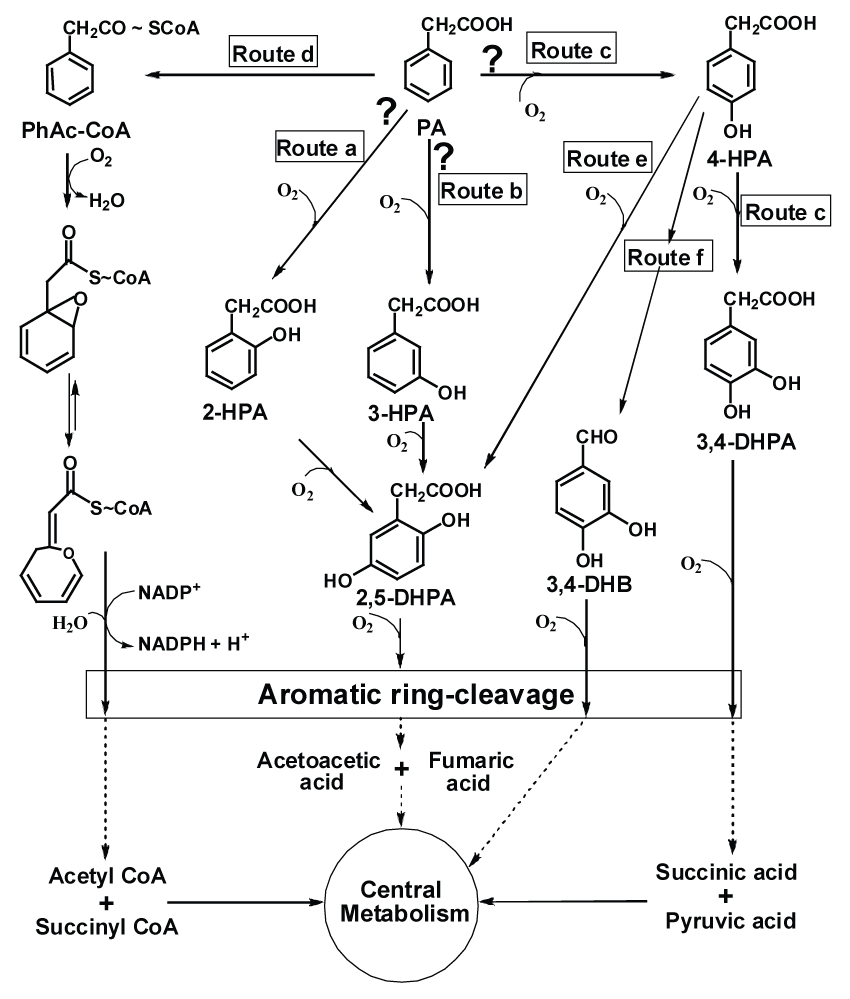
| Phenylacetic acid (PA) and hydroxyphenylacetic acids (HPA) are ubiquitous and found in the nature as metabolic intermediates of aromatic amino acids, lignin, styrene etc. [5]. PA, particularly is biotechnological relevant because of its extensive usage in the production of b-lactam antibiotics. However, due to their industrial application they released into the environment thus leading to pollution. Microbes are known to degrade PA and HPA. Hence, it is important to study and understand pathway adapted by various microbes that can utilize these compounds as carbon source, thus preventing their excess accumulation in the environment. In bacteria, metabolic pathways of PA assimilation have been widely studied and considered to follow either of four possible routes (Figure 1). Utilization of PA via hydroxy intermediates (2-, 3-, or 4-hydroxyphenylacetic acid, Figure 1, route a, b or c, respectively) has been proposed in several microorganisms like Aspergillus, Nocardia, Flavobacterium, Trichosporon, Streptomyces, a chloridazon-degrading bacterium, P. putida, E. coli, Klebsiella, Acinetobacter and other unrelated microbes [6-12]. The metabolism of PA via hydroxylation of the aromatic ring was proposed in these strains. However, due to lack of evidences like presence of enzymes or ability to grow on hydroxylated intermediates, the hydroxylation pathways of PA metabolism are inconclusive so far. An unconventional 'aerobic hybrid pathway' of PA catabolism encoded by paa gene cluster has been initially identified in P. putida U [13-15] and E. coli W [16,17] where in PA is activated to its CoA-thioester (phenylacetyl-CoA) by phenylacetyl-CoA ligase (Figure 1, route d). This key enzyme has unequivocally been shown to be directly involved in the degradation of PA and its presence has been observed across the bacterial phylum [5,7,18-21]. A series of CoA-thioester intermediates have been involved in this pathway which ultimately leads to the formation of succinyl-CoA and acetyl-CoA that enters the central metabolic pathway [1] (Figure 1, route d). Recently, detailed metabolic route of PA degradation via CoA intermediate has been elucidated in E. coli K12 and Pseudomonas sp. strain Y2 [22]. This is the only pathway which has shown the complete utilization of PA via CoA intermediates (Figure 1, route d) and the activities of concerned enzymes. The PA pathway is the core of the phenylacetyl-CoA catabolon, a functional unit that integrates peripheral catabolic pathways that converts several structurally related aromatic compounds such as styrene, 2-phenylethylamine, tropic acid, and phenylacetyl esters and amide to the common intermediate phenylacetyl-CoA [23]. |
| The metabolic pathway of 4-hydroxyphenylacetic acid (4-HPA) has been elucidated in several bacteria. 4-HPA is metabolized either via 3,4-dihydroxyphenylacetic acid (3,4-DHPA or homoprotocatechuate pathway, Figure 1, route c) or 2,5-dihydroxyphenylacetic acid (2,5-DHPA or homogentisate pathway, Figure 1, route e). Homoprotocatechuate pathway involves initial hydroxylation of 4-HPA at C3 position to yield 3,4-DHPA, which is subsequently ring-cleaved to yield semialdehyde intermediate. This yellow colored intermediate ultimately leads to the formation of succinic and pyruvic acid (Figure 1, route c) [14,24-27]. |
| Homogentisate pathway of 4-HPA involves hydroxylation of 4-HPA at C1 position with migration of the carboxymethyl side-chain to C2 position due to 'NIH-shift' reaction [28]. The resulting intermediate (2,5-DHPA) yields maleylacetoacetic acid in subsequent steps. This ring cleaved structure ultimately results in the formation of acetoacetic acid and fumaric acid (Figure 1, route e) [11,27,29-31]. Other alternative pathways for 4-HPA metabolism have been reported (Figure 1, route f) in which degradation of the side-chain proceeds before oxidative ringcleavage occurs [5,32]. |
| A soil bacterium, P. putida strain CSV86 utilizes various aromatic compounds like naphthalene, methylnaphthalene, benzylalcohol, benzoate etc. as the sole carbon and energy source [33,34]. Strain CSV86 is reported to utilize aromatic compounds in preference to glucose [35]. In the present study, we report the metabolic pathways of PA and 4-HPA degradation in strain CSV86. Metabolic and biochemical studies demonstrate that the strain utilizes PA and 4-HPA preferentially over glucose. |
| Materials and Methods |
| Chemicals |
| 4-Hydroxyphenylacetic acid (4-HPA), 3,4-dihydroxyphenylacetic acid (3,4-DHPA, Homoprotocatechuate) and 2,5-dihydroxyphenylacetic acid (2,5-DHPA, homogentisate), ATP, Coenzyme-A, hydroxylamine hydrochloride and Tris were purchased from Sigma-Aldrich (USA). Phenylacetic acid (PA) was purchased from Spectrochem (India). NADH, NAD(P)H and FAD were purchased from SRL (India). All other chemicals were of analytical grade and purchased locally. |
| Growth conditions |
| Pseudomonas putida CSV86 was grown on 150 ml mineral salt medium (MSM) at 30°C on a rotary shaker [33]. PA, 4-HPA (0.1%) or glucose (0.25%) was added aseptically as the carbon source either alone or in combination. Growth was monitored at 540 nm spectrophotometrically (Perkin Elmer Lambda 35) using MSM as the blank. |
| Chemical estimations |
| Reducing sugar estimation in the medium was performed as described by Miller using glucose as the standard [36]. The protein content was estimated as described by Bradford using bovine serum albumin as the standard [37]. |
| Metabolite isolation and identification |
| Metabolites from the spent medium were extracted as described previously [33]. Metabolites were resolved by thin layer chromatography (TLC) on 0.5-mm silica gel coated plates using solvent system- benzene: acetone: acetic acid (6:3:1) and identified by comparing Rf and UV-fluorescence properties at 254 nm with the authentic compounds [PA (Rf, 0.96; whitish); 4-HPA (Rf, 0.83; black non fluorescent); 3,4-DHPA(Rf , 0.67; black non fluorescent); 2,5- DHPA (Rf , 0.68; black non fluorescent)]. |
| Biotransformation and oxygen uptake |
| CSV86 cells grown on the respective carbon source until the latelog phase were harvested by centrifugation (12,000g for 10min), and washed twice with phosphate buffer (50 mM, pH 7.5). Biotransformation experiment using whole-cells was carried out as described previously for 3h [33]. To monitor O2 uptake, a cell suspension was prepared by re-suspending 100 mg of cells (wet weight) in 1 ml of phosphate buffer. Respiration rates were monitored as described [38] using an Oxygraph (Hansatech, UK) fitted with Clark's type O2 electrode. The reaction mixture (2 ml) contained cells (4 mg, wet weight), substrate (200 µM) and phosphate buffer (50 mM, pH 7.5) at 30°C. The rates were corrected for endogenous O2 consumption and expressed as nmol O2 consumed min-1 mg-1 cells (wet weight). |
| Preparation of cell-free extracts and enzyme activities |
| Cell-free extracts were prepared from CSV86 cells grown on the appropriate carbon source (aromatics or glucose) till the late log-phase. Cells were harvested and washed twice with phosphate buffer (50 mM, pH 7.5). Cells (1 g wet wt) were suspended in 4 ml ice-cold phosphate buffer and sonicated at 4°C with four cycles of 15 pulses each (1s pulse, 1s interval, cycle duration 30s, output 15 W) using an Ultrasonic processor (model GE130, USA), and centrifuged at 20,000 g for 20 min. The clear supernatant obtained was referred to as the cell-free extract and used for monitoring various enzyme activities. Specific activities of enzymes are expressed as nmol min-1 mg-1 of protein. |
| Phenylacetyl-CoA ligase was measured essentially as described earlier [13]. In brief, the reaction mixture contained MgCl2 (0.2 M, 12.5 µl), ATP (0.1 M, 50 µl), CoA (20 mM, 30 µl), potassium phenylacetate (0.2 M, 30 µl), hydroxylamine solution pH 8.0 (50 µl, prepared by mixing 1 ml hydroxylamine hydrochloride (5 M), 250 µl of distilled water, and 1.25 ml KOH (4 M). All substrates except MgCl2 were dissolved in Tris (50 mM) and adjusted to pH 8.2. In control reactions, potassium phenylacetate, ATP, MgCl2, or CoA were omitted. After 5 min of temperature equilibration in a water bath at 30°C, 100 µl of cell-free extract was added and the incubations were carried out for 20 min at 30°C. Reactions were stopped by adding ferric-chloride reagent (450 µl, prepared by mixing ferric chloride (0.37 M), trichloroacetic acid (200 mM) and hydrochloric acid (0.66 M)) and kept on ice for 30 min. The reactions were centrifuged in a microcentrifuge for 2 min. The activity was followed by measuring the rate of formation of phenylacetylhydroxamate (red-purple color) at 540 nm spectrophotometerically. The extinction coefficient of phenylacetylhydroxamate under these conditions was 0.9 mM-1 cm-1 [13]. One unit of enzyme activity is defined as the catalytic activity leading to the formation of 1 nmol of phenylacetylhydroxamate in 1 min. |
| 4-Hydroxyphenylacetic acid hydroxylase (4-HPAH) activity was measured by monitoring the rate of decrease in the absorbance at 340 nm (εNADH = 6220 M-1 cm-1, [39]) spectrophotometrically. The reaction mixture (1 ml) contained phosphate buffer (50 mM, pH 7.5), NADH (100 µM), FAD (200 µM) and 4-HPA (1 mM). A unit of enzyme activity is defined as the amount of enzyme required to oxidize 1 µmol of NADH/min under standard conditions. |
| Phenylacetic acid hydroxylase (PAH) activity was measured by monitoring the rate of decrease in the absorbance at 340 nm (εNADH = 6220 M-1 cm-1) spectrophotometrically. The reaction mixture was identical to that described for the assay of 4-HPAH except that the substrate was PA (100 µM to 1 mM). The activity was determined as described for 4-HPAH. |
| 3,4-Dihydroxyphenylacetic acid dioxygenase (3,4-DHPADO) was monitored by following the increase in the absorbance at 380 nm due to the formation of the product 5-carboxymethyl-2- hydroxymuconic semialdehyde (CHS) (εCHS = 38,000 M-1 cm-1, [39]) spectrophotometrically. The reaction mixture (1 ml) contained phosphate buffer (50 mM, pH 7.5), 3,4-dihydroxyphenylacetic acid (500 µM) and appropriate amount of enzyme. A unit of enzyme activity is defined as that required to form 1 µmol of the muconic semialdehyde/ min under standard assay conditions. |
| 2,5-Dihydroxyphenylacetic acid dioxygenase (2,5-DHPADO) assay was performed by monitoring the rate of increase in the absorbance at 330 nm [40] spectrophotometrically. The reaction mixtures (1 ml) contained potassium phosphate buffer (50 mM, pH 7.5), ascorbic acid (2 mM), FeSO4 (50 µM) and homogentisic acid (200 µM) and appropriate amount of enzyme solution. |
| Glucose-6-phosphate dehydrogenase (Zwf) was monitored spectrophotometrically by measuring the rate of increase in the absorbance at 340 nm due to the appearance of NADPH (εNADPH = 6220 M-1 cm-1, [41]). Reaction mixture (1 ml) contained Tris-HCl buffer (100 mM, pH 8.5), glucose 6-phosphate (4 mM), NADP (200 µM) and appropriate amount of enzyme. |
| All experiments described were performed atleast three times with readings in triplicates. The observed standard deviations (SD) are represented by error bars at appropriate places. |
| Results and Discussion |
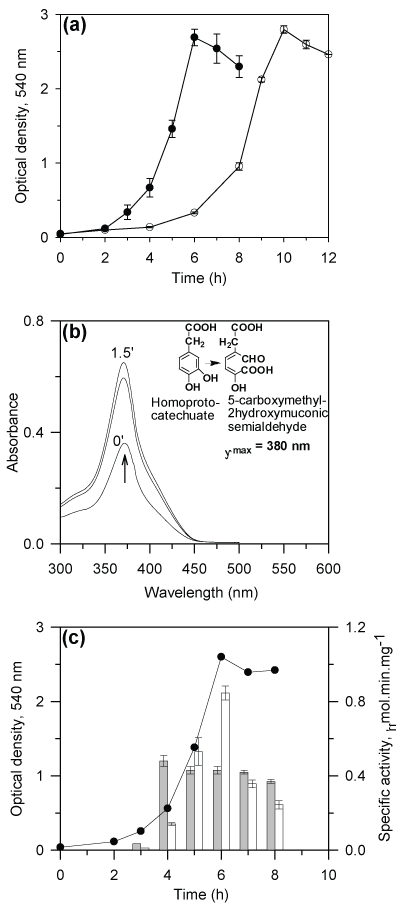
| P. putida CSV86 utilizes naphthalene, 1- and 2-methylnaphthalene, benzylalcohol, 2-hydroxybenzylalcohol, 4-hydroxybenzylalcohol, p-hydroxybenzoate, benzoate, salicylate and benzaldehyde as the carbon source. The metabolic pathways of degradation of these aromatic compounds have been elucidated [33,34]. Besides these compounds, strain CSV86 has been found to utilize phenylacetic acid (PA) and 4-hydroxyphenylacetic acid (4-HPA) as the sole carbon and energy source (Figure 2a). The observed specific growth rate (µ, h-1) and doubling time (h) were found to be: PA (0.8, 0.87) and 4-HPA (1.0, 0.7). These specific growth rates were higher than those observed earlier [42] on naphthalene (0.51, 1.36), salicylate (0.34, 2.04), benzylalcohol (0.54, 1.28), benzoate (0.59, 1.17) glucose (0.22, 3.15) and succinate (0.76, 0.91), suggesting that strain CSV86 utilizes PA and 4-HPA faster than other carbon sources. |
| Metabolism of phenylacetic acid and 4-hydroxyphenylacetic acid |
| The catabolic pathways for PA and 4-HPA were elucidated by isolating and identifying intermediates from the spent media using TLC, monitoring enzyme activities in the cell-free extracts, whole-cell respiration and biotransformation studies. Strain CSV86 cells grown on PA showed significantly higher oxygen uptake on PA compared to 4-HPA, 3,4-DHPA, 2,5-DHPA and succinate (table 1). Cell-free extract obtained from cells grown on PA showed activity of phenylacetyl- CoA ligase. The activity of PA hydroxylase, 4-HPAH, 3,4-DHPADO and 2,5-DHPADO were not detected in the cell-free extract (Table 2). Spent medium analysis by TLC from cells grown on PA failed to show any metabolite spot corresponding to 4-HPA, 3,4-DHPA or 2,5-DHPA during the early log- and late log-phase of the growth cycle suggesting that PA is not metabolized through hydroxylation pathway (data not shown). TLC analysis of biotransformation reaction using PA as substrate failed to show any metabolite spots related to 4-HPA and 3,4- DHPA. Similarly, biotransformation of 4-HPA from PA grown cells failed to show the spots of 3,4-DHPA and the only spot observed was of 4-HPA (data not shown). These results indicated that in strain CSV86, PA metabolism followed route d as shown in Figure 1, i.e. by activating PA to phenylacetyl-CoA. Till recently, the bacterial pathway involved in aerobic PA metabolism was not clear. Although hydroxylated phenylacetic acids were detected in some PA degrading bacteria [8,9,11,12], neither ring-hydroxylating nor ring-cleaving reactions for PA metabolism could be demonstrated. Recently, PA 'aerobic hybrid pathway' via CoA-thioesters as intermediates has been reported in E. coli W and P. putida Y2 [22]. This pathway does not involve utilization of molecular oxygen and resembles the anaerobic mode of metabolism. The key enzyme, phenylacetyl-CoA ligase, responsible for initial activation in this pathway has been reported in several bacterial strains [5,13,17,20] and the gene coding for phenylacetate-CoA ligase has been identified in these species [5,15,17]. |
| CSV86 cells grown on 4-HPA showed higher oxygen uptake on 4-HPA and 3,4-DHPA but failed to respire on PA and 2,5-DHPA (table 1). The cell-free extract obtained from 4-HPA grown cells showed significant activities of 4-HPAH and 3,4-DHPADO, however, the activity of 2,5-DHPADO and phenylacetyl-CoA ligase were not detected (Table 2). The ring-cleavage product of 3,4-DHPA was identified by performing time-dependent changes in the spectral properties during the enzyme reaction. The cell-free extract prepared from cells grown on 4-HPA showed an increase in the absorbance at 380 nm due to the formation of a yellow colored product, 5-carboxymethyl-2-hydroxymuconic semialdehyde when 3,4-DHPA was used as the substrate (Figure 2b). On the other hand with 2,5- DHPA as substrate, an increase in the absorbance at 330 nm (due to the formation of maleylacetoacetic acid) was not observed (data not shown), suggesting the absence of 2,5-DHPADO. These observations showed the 4-HPA metabolism followed homoprotocatechuate pathway in strain CSV86 (Figure 1, route c). Moreover, growth versus specific activity profiles from cells grown on 4-HPA showed higher activity of 4-HPA hydroxylase during the early phase of the growth (3- 5h), while the activity of 3,4-HPA dioxygenase was maximum during later phase of the growth (6h) suggesting their sequential induction (Figure 2c). Spent medium analysis from cells grown on 4-HPA showed the presence of 3,4-DHPA during early log and late log-phase while 2,5-DHPA was not detected in the spent medium which suggests that 4-HPA was hydroxylated to 3,4-DHPA and not 2,5-DHPA during its metabolism (data not shown). Biotransformation of 4-HPA and PA was carried out using 4-HPA grown cells. During biotransformation on 4-HPA, TLC gave spots with Rf and fluorescence properties identical to authentic 4-HPA and 3,4-DHPA. However, no intermediates were detected with PA as substrate (data not shown). These results indicated that 4-HPA metabolism in strain CSV86 occurs via 3,4-DHPA and succinate (homoprotocatechuate pathway) and not via 2,5-DHPA (homogentisate pathway). |
| Results presented provide evidences that in strain CSV86, the metabolism of PA (Figure 1, route d) and its hydroxylated derivative (4-HPA) (Figure 1, route c) followed two different unrelated routes. A bacterial strain capable of utilizing PA and 4-HPA via different metabolic routes has been demonstrated earlier in P. putida U and Azoarcus evansii [5,14]. |
| Preferential utilization of PA and 4-HPA |
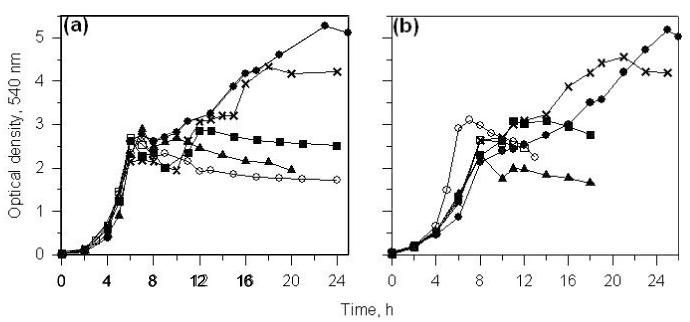
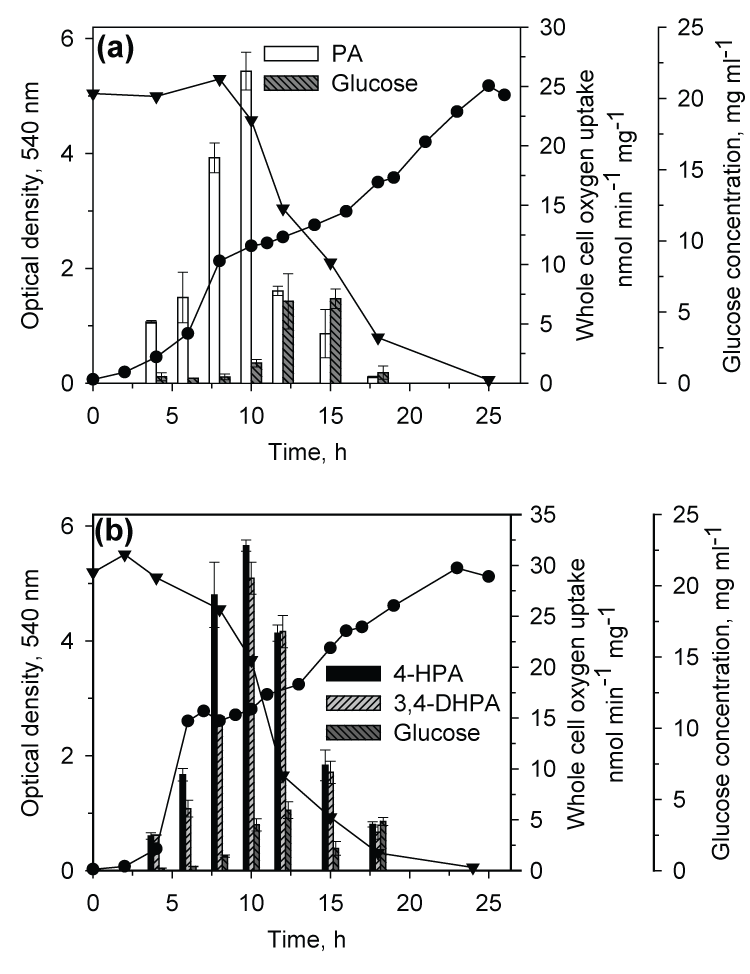
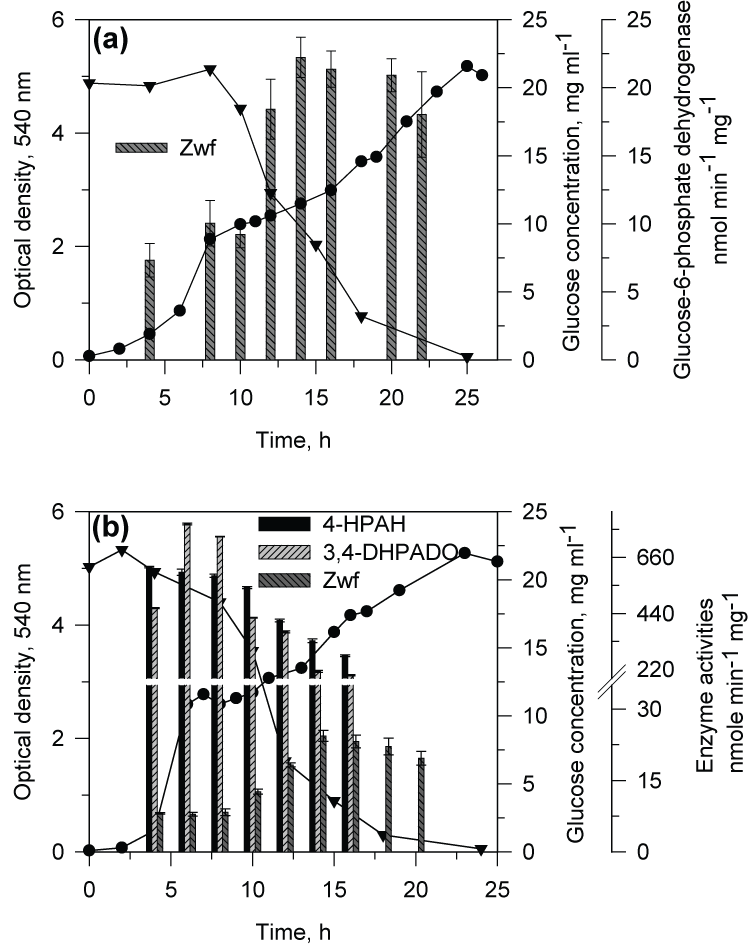
| When CSV86 cells grown on PA or 4-HPA were inoculated in MSM containing PA plus glucose or 4-HPA plus glucose, culture showed distinct diauxic profiles at higher glucose concentration (1 and 2%) where it utilized PA or 4-HPA during the first log-phase and glucose during the second log-phase (Figure 3a and b). Cell respiration and enzyme activity studies were carried out from cells grown on double carbon source, i.e. PA plus glucose or 4-HPA plus glucose. During the growth on PA (0.1 %) plus glucose (2%), high respiration rate was observed on PA during the first log-phase while the cell respiration rate was increased significantly on glucose during the second log-phase. The glucose concentration in the medium was constant during PA utilization and decreased only during the second log phase (glucose utilization) suggesting the sequential utilization of these two compounds (Figure 4a). Cell-free extracts prepared from cells grown on PA plus glucose showed low activity of glucose-6- phosphate dehydrogenase (Zwf) during the first log- phase but the activity increased as cells entered into the second log glucose utilization phase (Figure 5a). Similar results were observed when cells were grown on 4-HPA (0.1%) plus glucose (2%). Cells showed higher respiration rates on 4-HPA and 3,4-DHPA during the first log- phase with low rates for glucose which increased significantly in the second log-phase (Figure 4b). Growth versus enzyme specific activity analysis showed the presence of 4-HPAH and 3,4-DHPADO in the first log-phase while the activity of glucose metabolizing enzyme (Zwf) increased during the second log-phase (Figure 5b). |
| At low glucose concentration (0.1 and 0.25%), the second lag- and log-phase were not distinct. This could be due to high number of cells at the end of first log-phase which consume glucose rapidly without showing significant increase in the cell mass. However, analysis of glucose concentration in the spent medium revealed that the amount of glucose remained constant till 6 (4-HPA plus glucose) and 10 h (PA plus glucose); and then decreased. These results indicated that glucose utilization starts during the late log-phase of first growth cycle when low concentrations of glucose was used with PA or 4-HPA. Hence, although the diauxic growth pattern is not clear at low glucose concentrations, the culture utilized PA and 4-HPA earlier than glucose. Increasing the concentrations of glucose resulted in the longer durations of the second log-phase and constant durations of the first log-phase with a distinct second lag-phase. The sequential utilization of aromatics and glucose has also been determined using oxygen uptake and enzyme activity analysis and the results suggest that strain CSV86 utilizes PA and 4-HPA preferentially over glucose. |
| It has been reported that the degradation of PA has been repressed due to the carbon catabolite repression. Glucose particularly has been shown to repress the activity of phenylacetyl-CoA ligase in P. putida [13] and PA degradation by suppressing the transport of phenylacetic acid inside the cell in P. putida U [43]. However, the preferential utilization of aromatic compounds has been reported in strain CSV86 [35], the observation of preferential utilization of PA is novel as it isthe first report showing preferentiality of an aerobic hybrid pathway. Moreover, phenylacetyl-CoA pathway is regarded as the central route of convergence of a complex functional unit termed as phenylacetyl- CoA catabolon [15,23]. This unit integrates different upper or peripheral pathways involved in the degradation of several compounds that are structurally related to PA like styrene, 2-phenylethylamine, 2-phenylethanol, tropic acid, esters of PA etc. Hence, it may be hypothesized that in strain CSV86, a functional unit is being operated preferentially even in the presence of simple structure carbon sources like glucose. |
| Microorganisms in nature show the preference for structurally simple carbon sources like glucose and unless they are completely depleted, alternative energy sources like polysaccharides or aromatic compounds are not degraded [44]. This preferential utilization of simple carbon source represses the utilization of complex compounds leading to their accumulation in nature thereby locking the carbon thus aggravating the pollution. P. putida CSV86 utilizes aromatic compounds or organic acids in preference to glucose and co-utilizes aromatics and organic acids [35]. The uptake of glucose in CSV86 is by an active transport, and its metabolism is via an inducible intracellular phosphorylative pathway [42]. Also, it has been reported that the preferential utilization of aromatic compounds over glucose in CSV86 involves the suppression of the activity of glucose 6-phosphate dehydrogenase (Zwf) by aromatics and the inability of glucose to suppress the activity of aromatic degrading enzymes [42]. Additionally, at the transport level of glucose, a 43kDa peri-plasmic glucose binding protein [45] and a glucose inducible 40kDa outer membrane protein [46] have been found to be modulated when organism grown on double carbon source of aromatics and glucose. Hence, the ability to utilize various range of aromatic compounds preferentially make strain CSV86 an ideal candidate for the biotechnological and bioremediational purposes. In addition, these results may also be utilized to generate genetically engineered strains having aromaticsuppressible glucose transport systems which may prove to be more proficient for environmental degradation of aromatic compounds. |
References
|
Tables and Figures at a glance
| Table 1 | Table 2 |
Figures at a glance
 |
 |
 |
 |
 |
| Figure 1 | Figure 2 | Figure 3 | Figure 4 | Figure 5 |
Relevant Topics
- Anaerobic Biodegradation
- Biodegradable Balloons
- Biodegradable Confetti
- Biodegradable Diapers
- Biodegradable Plastics
- Biodegradable Sunscreen
- Biodegradation
- Bioremediation Bacteria
- Bioremediation Oil Spills
- Bioremediation Plants
- Bioremediation Products
- Ex Situ Bioremediation
- Heavy Metal Bioremediation
- In Situ Bioremediation
- Mycoremediation
- Non Biodegradable
- Phytoremediation
- Sewage Water Treatment
- Soil Bioremediation
- Types of Upwelling
- Waste Degredation
- Xenobiotics
Recommended Journals
Article Tools
Article Usage
- Total views: 16613
- [From(publication date):
June-2011 - Dec 23, 2025] - Breakdown by view type
- HTML page views : 11770
- PDF downloads : 4843
