Research Article Open Access
Microbial Decolorization and Degradation of Orange 16 Dye by a Newly Isolated Aeromonas Spp. Etl-1949
| Maulin P Shah1*, Patel KA1, Nair SS1, Darji AM1 and Maharaul SJ2 | |
| 1Industrial Waste Water Research Laboratory, Applied & Environmental Microbiology Lab, Enviro Technology Limited (CETP) Plot No: 2413/2414, GIDC, Ankleshwar- 393002, Gujarat, India | |
| 2Laboratory of Environmental Bioremediation, Narmada Clean Tech Limited, Final Effluent Treatment Plant, Surti Bhagor, Near Gujarat Gas, Umarvada Road, Ankleshwar-393001, Gujarat, India | |
| Corresponding Author : | Maulin P Shah Industrial Waste Water Research Laboratory Applied & Environmental Microbiology Lab Enviro Technology Limited (CETP) Plot No: 2413/2414 GIDC, Ankleshwar- 393 002, Gujarat, India Tel: +91-90 999 65504 Fax No: +91-2646-250707 E-mail: shahmp@uniphos.com |
| Received June 08, 2013; Accepted July 22, 2013; Published July 24, 2013 | |
| Citation: Shah MP, Patel KA, Nair SS, Darji AM, Maharaul SJ (2013) Microbial Decolorization and Degradation of Orange 16 Dye by a Newly Isolated Aeromonas Spp. Etl-1949. J Bioremed Biodeg 4:194. doi:10.4172/2155-6199.1000194 | |
| Copyright: © 2013 Shah MP, et al. This is an open-a ccess article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. | |
Related article at Pubmed Pubmed  Scholar Google Scholar Google |
|
Visit for more related articles at Journal of Bioremediation & Biodegradation
Abstract
A bacterial strain ETL-1949, identified as Aeromonas spp. was isolated from sediment from a textile wastewater treatment plant in Ankleshwar, Gujarat, India. The strain could decolorize an azo dye, Orange 16 under the static condition after the shaking culture. In order to increase cell concentration by the shaking culture, the optimum culture condition was investigated for following the decolorization incubation. The dye was decolorized completely up to concentration of 750 mg/l for 50 hr under static condition by using 12 hr-cultured cells at optimal shaking culture condition. Effect of initial cell mass showed that higher cell mass concentration can accelerate decolorization process with maximum of 92% decolorization observed at 2.5 g/l cell mass within 6.5 h. Effect of various metal ions showed Mn & Mg has positive inducing effect whereas Zn strongly inhibited the decolorization process at 5 mM concentration. Analysis of biodegradation products carried out with UV–vis spectroscopy, HPTLC and FTIR confirmed the decolorization and degradation of Orange 16.
| Keywords |
| Aeromonas; Orange 16; Textile waste water; Ankleshwar |
| Introduction |
| One of the major problems that humans are facing is the restoration of the contaminated environment. Textile dyes contribute as the most important environment-polluting agents. Several classes of such contaminants have been synthesized, and still new products are being synthesized now and then. The textile industry is a large water consumer and produces large volumes of contaminated water. One of such examples is the Ankleshwar Industrial Estate, Ankleshwar, Gujarat, India, which is a seriously industrialized area and produces millions of liters of improperly treated effluents that are released directly without giving proper treatment. Synthetic dyes released into the environment in the form of effluents by textile, leather, food, paper and printing industries cause severe ecological damages. Wastewater resulting from dyeing and finishing processes has an adverse impact in terms of total organic carbon, biological oxygen demand and chemical oxygen demand. Azo dyes are the main constituents of such pollution because of their wide applicability and usages, and therefore, these are present majorly in textile industrial effluents. Moreover their toxicity and resistance to degradation offer great challenge for removal technologies. In many cases the products formed after the degradation of the parent azo dye molecule are more toxic. These products are mainly in aromatic amine form. Azo dyes have been shown to be mutagenic to the human hepatoma cell line where frame shift mutation was observed [1,2]. Induction in the micronuclei formation in human lymphocytes and in HepG2 cells after treatment with azo dye was also observed [3]. The effluent from a dye-processing plant was shown to be responsible for the mutagenic activity detected in a Brazilian river [4]. Methyl red is mutagenic in nature, and most microbial degradation studies reveal the formation of N, N-dimethyl-phenylenediamine, a toxic and mutagenic aromatic amine [5] that remains untreated in the culture [6]. Acid violet 7 has a significant ability to induce chromosome aberrations, lipid peroxidation and inhibition of acetylcholinesterase. Its toxicity increases extensively after static biodegradation with Pseudomonas putida, due to the corresponding azo reduction metabolites 4’ -aminoacetanilide and 5-acetamido-2-amino-1-hydroxy-3,6- naphthalene disulphonic acid [7]. Therefore azo dyes are the prime attention of the researchers because of their toxicity perspectives. Several physicochemical and microbial methods have been developed for the removal and detoxification of azo dyes. However the developed physicochemical methods present several drawbacks such as high cost, high generation of sludge, high-energy-requiring irradiation methods requires a lot of dissolved O2, etc. [8,9]. Biological methods represent more proper way of textile azo dye removal. Several microorganisms such as algae, yeast, filamentous fungi and bacteria individually or in consortium are shown to degrade the azo dyes in the presence of nutrients [10-15]. However only yeast species Saccharomyces cerevisiae MTCC 463 has been shown to decolorize azo dye in plain distilled water [16]. Rapid industrialization has necessitated the manufacture and use of different chemicals in day to day life [17]. The textile industry is one of them which extensively use synthetic chemicals as dyes. Wastewaters from textile industries pose a threat to the environment, as large amount of chemically different dyes are used. A significant proportion of these dyes enter the environment via wastewater [18]. Approximately 10,000 different dyes and pigments are used industrially and over 0.7 million tons of synthetic dyes are produced annually, worldwide [19]. In this study efficiency of a bacterial species Aeromonas spp. ETL-1979 was evaluated for decolorization, and degradation of textile azo dye Orange 16. A bacterial strain, ETL-1949 isolated from sediment of a textile industrial wastewater treatment plant, showed strong ability to remove the color of Orange16 under static condition. Preliminary experiments of this bacterial cultivation suggested that the initial cell concentration was a very important factor to decolorize Orange 16 following static incubation. So we employed a serial 2-step culture condition. After growing the bacterial cells in the medium aerobically with the low concentration of the dye, the cultured medium was shifted to the static condition with addition of the dye for the decolorization. A set of experiments has been carried out in batch culture to optimize the cell growth under aerobic condition. The effect of several culture conditions such as temperature, initial pH, and nutrient concentration of the medium were investigated. |
| Materials and Methods |
| Measurement of dye and cell growth |
| Orange 16 was obtained from local textile industry of Ankleshwar, Gujarat, India. Concentration of Orange 16 in the medium was determined by measuring the absorbance at 492 nm corresponding γ max of this dye. Bacterial growth of the cultures was measured as OD at 600. |
| Microorganism and culture condition |
| Bacterial strain ETL-1949 used in this work was isolated from sediment of the wastewater treatment facilities of a textile industry in Ankleshwar, Gujarat, India. The medium used for screening of bacteria with 0.02 g/l Orange 16 decolorizing potential contained 5 g/l glycerol, 4 g/l yeast extract, 0.5 g/l ammonium sulfate, 0.2 g/l MgSO4.7H2O, 0.5 g/l K2HP O4, and 1.5 g/l K2HPO4. The initial pH was adjusted to 8.0. The strain was cultivated in a reciprocal shaker at 30°C. The culture conditions including the medium composition were varied to determine the optimum condition for the bacterial growth. Batch culture experiments under the aerobic condition were carried out in 500 ml conical flask, containing 100 ml medium inoculated with a loopful of I-day grown-cells from the slant. Flasks were shaken at 125 rpm in a reciprocal shaker. Aliquots collected at intervals were used for measurement of the cell growth. For decolorization experiments, the bacterial cells under optimal growth condition were transferred to 100 ml of Erlen-Meyer flask. Orange 16 was added to give 0.15-2 mg/l and further incubated at 30°C under the static condition. |
| Identification of bacterial strain ETL-1949 |
| The taxonomical studies were carried out according to Bargey’s manual.The 16s rRNA gene in this strain was amplified by 30 cycle of PCR using genomic DNA from strain ETL-1949 as the template and two specific primers, 20F (5’-GATTTTGATCCTGGCTCA-3’) and 1500R (5’-GTTACCTTGTTACGACTT-3’). Each cycle was carried out at 95°C for 30 sec, at 55°C for 20 sec, and at 72°C for 90 sec. An amplified DNA fragment was sequenced by the dideoxynucleotide chain termination method using Big Dye terminator Cycle Sequencing Kit (Perkin-Elmer, Foster City, California) and a DNA sequencer (model 310, Perkin-Elmer). Homology of the DNA sequence was searched by using the BLASTn program at DDBJ. |
| Effect of physicochemical parameters |
| Effect of different pH was studied by adjusting the pH of the distilled water (with 0.1 M HCl and 0.1 M NaOH) prior to cells and dye addition. Similarly, to study the effect of different temperatures on decolorization performance, flasks containing cells suspended in distilled water were kept at respective temperatures for 15 min before dye addition to attain the temperature. After dye addition the incubation was continued at respective temperatures. The decolorization was monitored after 5 h of incubation. The percent decolorization was measured at different time intervals. |
| Effect of cell mass |
| Impact of various cell mass concentrations on decolorization rate was studied here. Experiments were carried out at cell concentration viz. 0.5, 1.0, 1.5, 2.0 and 2.5 gl-1. Flasks were incubated at respective conditions, and decolorization was monitored at various time intervals. |
| Effect of heavy metal ions |
| To inspect the influence of various metal ions such as Mg (MgCl2), Co(CoCl2), Zn (ZnSO4), Mn (MnCl2) and Hg (HgCl2) on decolorization activity, experiments were carried out in presence of these metals at concentration 1 and 5 mM. Metal ions from stock solutions were added to the cell suspension and incubated for 15 min, and then flasks were added with the dye. Decolorization was monitored at 30°C for different time intervals. |
| UV-Vis spectral analysis, HPTLC, FTIR |
| UV-visible spectral analysis of cell free sample, before and after dye decolorization, was carried by using Shimadzu UV spectrophotometer (Shimadzu UV 1800, Japan), and changes in its absorption spectrum in the visible range (400-800 nm) were recorded. HPTLC analysis was carried out by using HPTLC system (CAMAG, Switzerland). Samples of dye and its biodegradation metabolites (dissolved in HPLC-grade methanol) were loaded on precoated HPTLC plates (Silica gel 60F 254, Merck, Germany), by using nitrogen as a spraying gas and TLC sample loading instrument (CAMAG LINOMAT 5). TLC plate was developed in solvent system toluene to ethyl acetate to methanol (2:4:13). After development, the plate was observed in UV chamber (CAMAG) and scanned at 254 nm with slit dimension 5×0.45 mm by using TLC scanner (CAMAG). The results were generated by using HPTLC software WinCATS 1.4.4.6337. For FTIR analysis, residue obtained after evaporation of solvent extracts was mixed with stereoscopically pure KBr and analysis was carried out using Shimadzu 8400S spectrophotometer in the mid-infrared region of 400-4,000 cm-1 with 16-scan speed. Identification of metabolites formed after degradation of Orange 16 was carried out using a QP2010 gas chromatography coupled with mass spectroscopy (Shimadzu, Japan). The ionization voltage was 70 eV, and helium was used as carrier gas with a flow rate 1 mlmin-1 and 33 min run time. Gas chromatography was conducted in temperature programming mode with a Resteck column (0.25×30 mm; XTI-5). Column oven temperature was 280°C, and injection temperature was 200°C. Temperature was hold at 50°C for 1 min then rose to 280°C at 10°C rise per min. |
| Statistical analysis |
| Data were analyzed by one-way analysis of variance (ANOVA) with the Tukey-Kramer multiple comparisons test [20]. |
| Results and Discussion |
| Isolation of bacteria with high dye-decolorizing potential |
| A small portion of the sediment samples was inoculated to 100 ml of the basal medium and cultivated at 30°C for 24 hr with shaking and then flasks were subsequently incubated at 30°C for 24 hr under the static condition. The samples showing decolorization of Orange 16 were spread on the agar plate containing Orange 16 to observe clear zone formation. Isolated bacteria were cultivated again as described above to check the decolorization potential. Isolated strain ETL-1949 showed the highest decolorization rate. |
| Identification of a bacterial strain ETL-1949 |
| Isolated bacterial strain ETL-1949 was rod-shaped and negative in Gram-staining. The nucleotide sequence of 16s rDNA gene showed high homology to Aeromonas caviae, A. frola, and A. hydrophila, respectively. From these results, the bacterial strain ETL-1949 was identified as Aeromonas sp. |
| Decolorization of Orange 16 at various physicochemical conditions |
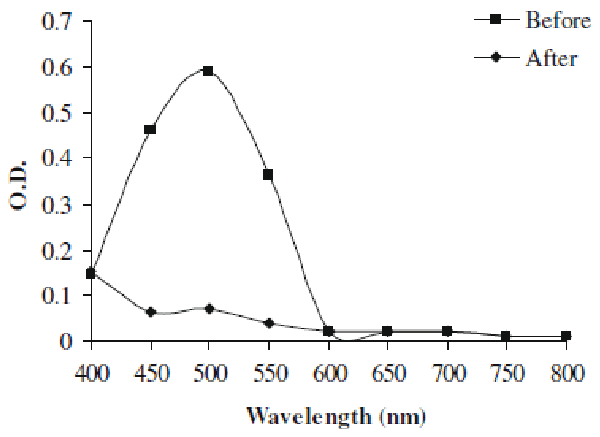
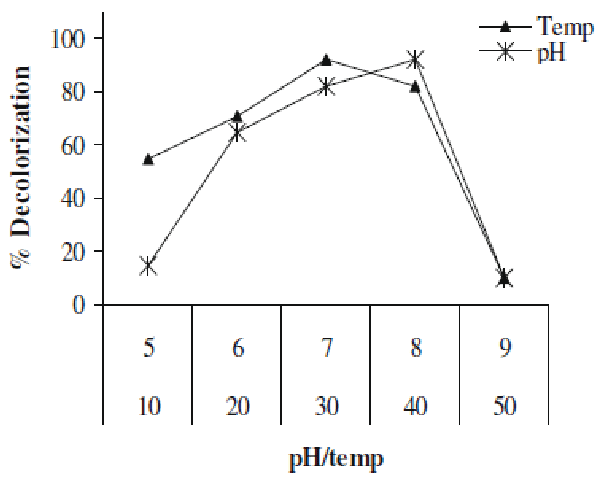
| In present study we investigated the decolorization potential of bacterium Aeromonas spp. ETL-1949 towards textile azo dye Orange 16. Bacterium was highly efficient in decolorizing the dye in absence of organic or inorganic nutrients. To confirm the decolorization of dye, UV-vis spectroscopic analysis was carried out. Figure 1 denotes the absorption spectra of Orange 16 before and after its bacterial treatment in visible range. Peak responsible for absorption maxima of parent dye (495 nm) was found to be almost completely disappeared in the sample obtained after dye decolorization. Only yeast species S. cerevisiae MTCC 463 has been shown to decolorize the azo dye methyl red without adding any organic or inorganic nutrients [16]. In the present study, Orange 16 was found to be adsorbed only initially and no adsorption was observed visually after dye decolorization. Enzymatic analysis in the extracellular sample as well as intracellular filtrate, before and after dye decolorization, exposed the induction in extracellular activity of tyrosinase and NADHDCIP reductase. Intracellular activities of laccase, veratryl alcohol oxidase, NADHDCIP reductase and tyrosinase were also induced (data not shown). These results clearly propose that dye was decolorized both by extracellular enzymatic degradation plus simultaneous adsorption to cell and decolorization by intracellular enzymes. The mechanism of simultaneous adsorption and then decolorization can be seen reported for various microbial species [21-23]. Figure 2 represents the optimum pH and temperature for the Orange 16 decolorization which came out to be 8 and 30°C, respectively. When dye decolorization was observed at shaking condition (120 rpm), no dye decolorization occurred. This designates that static condition was necessary as is the case for most of the microbial species for azo reduction. The general mechanism for the azo dye degradation is through reduction of azo bond by azoreductase under anaerobic conditions [24]. However there are certain oxygeninsensitive or aerobic azoreductases that have been reported from aerobic microorganisms [25,26]. But notably in our study, no activity of azoreductase was observed, suggesting different mechanism of this azo dye reduction other than azoreductase. It decolorized reactive, azo, diazo and triphenylmethane dyes. Viability of bacterial strain was the main question in this study, because it was the decolorizing variety of dyes in absence of nutrients. We checked the viability of the strain qualitatively by streaking the loop full of suspension from the dye decolorized flask. We observed normal growth pattern with colonies observed on all the four quadrants of agar plate, indicating that strain remains viable. These studies along with decolorization studies carried out with heat-killed cell suspension signify that dye decolorization was mediated by live culture. |
| Effect of cell mass concentration |
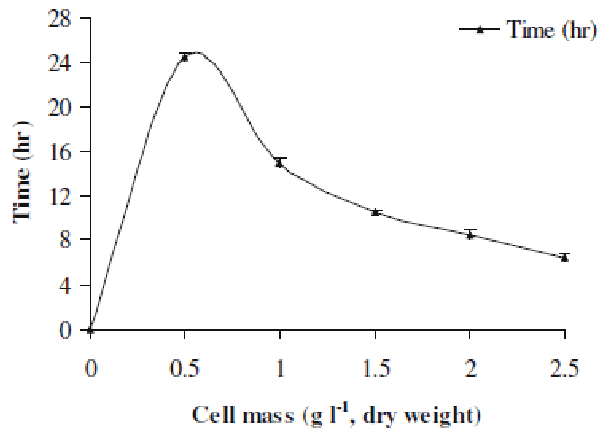
| Here we investigated the effect of various cell mass concentrations in the range 0.5-2.5 gl-1 (dry cell weight) on the decolorization performance. Figure 3 represents the results of this study. It was evident that as the cell mass increased from lower to higher concentration, the time required for decolorization of Orange 16 went on decreasing. The decolorization time for 0.5 gl-1 cell mass was 24.5 h, whereas it was reduced to 6.5 h for 2.5 gl-1 cell mass. Simply at higher concentration, there was more number of cells present which utilized the available dye molecules in minimum time as compared to that of lower cell mass concentration. This indicates that higher cell mass accelerated the decolorization process. |
| Effect of heavy metal ions |
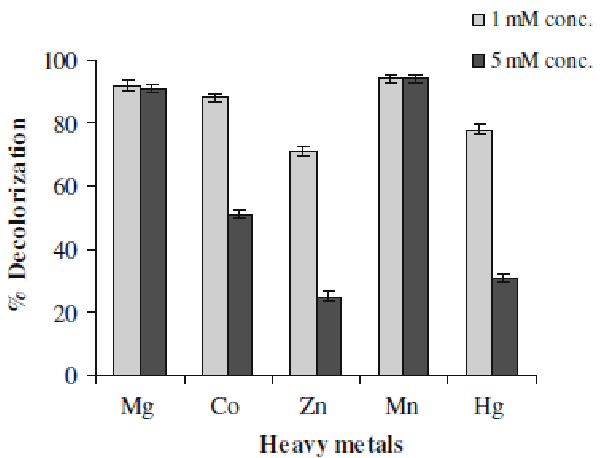
| Textile effluents often contain some amounts of heavy metals which make them more toxic. Some metals ions can be beneficial to the microorganisms but some can be harmful. The direct impact is on the proteins or enzymes because these forms complexes with protein molecules which render them inactive, for example inactivation of enzymes [27]. Hence we examined the effect of presence of some metal ions, at 1- and 5-mM concentrations, on decolorization activity of Aeromonas spp. ETL-1949. It can be observed in Figure 4 that Mn & Mg showed positive effect on decolorization performance (94%). Whereas Zn was the most inhibiting of the tested metals at both concentrations (25% decolorization at 5-Mm concentration). Co and Hg also exhibited the strong negative effect at 5-mM concentration. |
| Dye degradation analysis |
| HPTLC and FTIR Analyses |
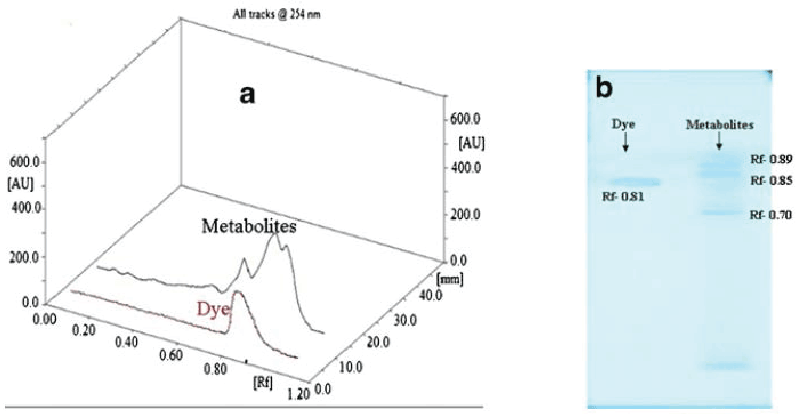
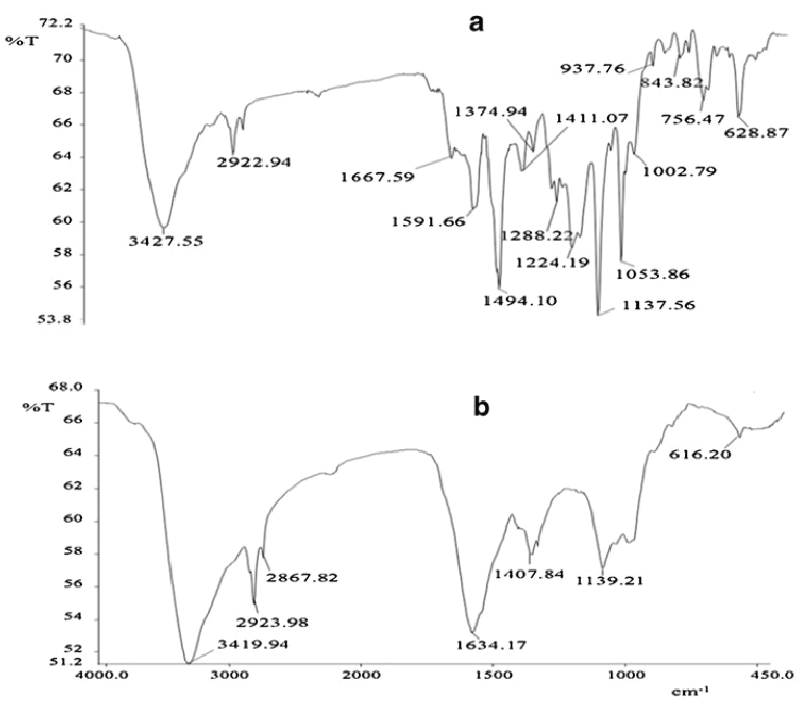
| Figure 5 gives the HPTLC analysis of parent dye compound and metabolites formed after dye degradation. Single peak was obtained for the parent dye, whereas three new and different peaks were observed for the metabolite sample in 3D graph of Rf values vs. absorbance (Figure 5a). It is also visible from fluorescent TLC plate that single band of dye at Rf value at 0.81 was almost disappeared in the metabolite sample. Three new bands with Rf values 0.89, 0.85 and 0.70 (scanned at 254 nm) appeared in the metabolite sample (Figure 5b). In FTIR analysis of dye compound (Figure 6a), presence of azo bond was revealed with the peak value 1591.66 cm-1. Peak at 628.87 cm-1 was responsible for C-S stretching in sulphur-containing compounds. Peaks at 1,053.86 and 1,124.19 cm-1 are representing the SOO asymmetric stretching in sulphonic acid groups. These three peak values confirm the presence of -SO3Na group in the dye structure. Trisubstituted C-H deformation was indicated by the peak at 1,224.19 cm-1. Peaks at 1,374.94 and 2,922.94 cm-1 reflect the alkanes C-H deformation and asymmetric stretching, respectively. Aromatic nature was exposed by the peak at COC in-plane vibrations for homocyclic compounds. Overall the structural appearance of the dye was reflected in this analysis. In analysis of metabolite sample (Figure 6b) peak at 616.20 cm-1 was responsible for C-S stretching and peak at 1,339.21 cm-1 for S0O symmetric stretching, suggesting that there may be an intermediate which was a sulphur-containing compound. Peaks at 1,407.84 cm-1 for C-O stretching plus O-H deformation in -COOH and 1,634.17 cm-1 for COO stretching in aryl carboxylic acid defines that there may be formation of carboxylic acid derivative. Peak at 2,923.98 cm-1 was indicating the O-H stretching. Peak value of 2,867.82 cm-1 demonstrated symmetrical stretching C-H in alkane. Notably there was absence of the peak responsible for azo (NON) group proposing that removal of azo bond after dye decolorization. Overall both these analytical studies (HPTLC and FTIR) point out that there was structural degradation of the dye molecule which leads to the decolorization. |
| Conclusion |
| Current investigation has confirmed the decolorization of Azo dye Orange 16 by the isolated Aeromonas spp. under in vitro conditions. Thus the study has confirmed the potential of Aeromonas spp. ETL-1979 in the decolorization of the dye indicating their possible application for treatment of textile effluents. By shifting the culture condition from the aerobic growth to the static condition, which is an oxygen limiting or anaerobic condition, Orange16 was efficiently decolorized. |
| Acknowledgement |
| Authors are highly grateful to the management of Enviro Technology Limited., Ankleshwar, Gujarat, India for allowing us to carry out such a noble work for the sustainable environment. |
References
|
Figures at a glance
 |
 |
 |
| Figure 1 | Figure 2 | Figure 3 |
 |
 |
 |
| Figure 4 | Figure 5 | Figure 6 |
Relevant Topics
- Anaerobic Biodegradation
- Biodegradable Balloons
- Biodegradable Confetti
- Biodegradable Diapers
- Biodegradable Plastics
- Biodegradable Sunscreen
- Biodegradation
- Bioremediation Bacteria
- Bioremediation Oil Spills
- Bioremediation Plants
- Bioremediation Products
- Ex Situ Bioremediation
- Heavy Metal Bioremediation
- In Situ Bioremediation
- Mycoremediation
- Non Biodegradable
- Phytoremediation
- Sewage Water Treatment
- Soil Bioremediation
- Types of Upwelling
- Waste Degredation
- Xenobiotics
Recommended Journals
Article Tools
Article Usage
- Total views: 16101
- [From(publication date):
July-2013 - Nov 16, 2025] - Breakdown by view type
- HTML page views : 11078
- PDF downloads : 5023
