Research Article Open Access
A Point of Care Real Time PCR Platform Based on Silicon Technology
Salvatore Petralia, Eloisa Castagna M, Massimo O Spata, Maria Grazia Amore and Sabrina Conoci*
Sabrina Conoci, STMicroelectronics, Stradale Primosole 50, Catania 95121, Italy
- *Corresponding Author:
- Sabrina Conoci, STMicroelectronics
- Stradale Primosole
50, Catania 95121, Italy - Tel: 41 227371000
- E-mail: sabrina.conoci@st.com
Received Date: February 26, 2016; Accepted Date: April 04, 2016; Published Date:April 10, 2016
Citation: Petralia S, Eloisa Castagna M, Spata MO, Amore MG, Conoci S (2016) A Point of Care Real Time PCR Platform Based on Silicon Technology. Biosens J 5:136. doi: 10.4172/2090-4967.1000136.
Copyright: © 2016 Petralia S, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Biosensors Journal
Abstract
The miniaturization of a real time PCR amplification systems are key to genetic point-of-care (POC) diagnostics to offer sample-in answer-out analysis. Actually, miniaturization typically enables shorter analysis times, reduces reagent consumption, minimizes risk of sample contamination, and often enhances assay performance. Here we present the genetic real time PCR PoC system based on silicon microchip able to address the quantitative and qualitative identification of multiple analytes of specific nucleic acids (DNA, RNA) sequences. Thanks to the combination of integrated silicon temperature sensors and heaters, the specific chip design architecture and the smart detection software, an improvement of sensitivity is achieved respect to the commercial tools.
Introduction
var googlewin=dhtmlwindow.open("googlebox", "iframe", https://www.omicsonline.org/socialnetworking/jcst-like.html", "Join with us", "width=250px,height=120px,resize=1,scrolling=1,center=1", "recal") The real-time PCR (Polymerase Chain Reaction) methodology has become an indispensable tool in many fields of molecular diagnostics, including determination of viral or bacterial loads in clinical samples, identification and titers of germs in food, diagnosis of tumors, gene expression analysis, or forensic analyses [1-3]. It is a wide used methodology since it is able to provide in a closed-system the quantitative result of target DNA sequence as it accumulates in real time in the reaction vessel. That allows avoiding time-consuming post- PCR analysis and reduces the risk of cross-contaminations enhancing both accuracy and speed.
.The miniaturization of real time PCR amplification systems is a key point towards the development of the so called “genetic pointof- care (PoC)” that are able to offer sample-in answer-out diagnostic analysis [4,5]. Several solutions based on multiple systems to integrate the sample preparation, nucleic acids amplification and detection via real time PCR [6], LAMP [7] or microarray [8] has been reported in the literature. All these miniaturized devices have been designed to enable shorter analysis times, reduce reagent consumption, minimize risk of sample contamination and enhance the assay performance such as sensitivity, selectivity and limit of detection.
In this context, silicon is very attractive material for biochips due to both its physical aspects for sensing (such as, the low heat capacity, the good thermal conductivity and the possibility to be made porous to increase the surface-area and reaction efficiency) and its technological aspects (consolidated production technologies and industrialization processes, integration of electrodes to the chip, as well as “intelligence on board” with microelectronics circuitry, high-volume production). Additionally, the coming of MEMS (Micro-Electro Mechanical System) technology has open the perspective to integrate micropumps, micronozzle and sensor combining multiple functions to achieve a complete analysis solution, including fluid management, amplification, hybridization or affinity binding, and detection.
For an optically transduced platform the implementation of software for the image analysis is also crucial to achieve the signal improvement. In particular for an optimized fluorescence analysis, it is extremely important to precisely detect the region of interest (ROI) areas where the PCR takes place. The alignment between the chip reaction chambers and the ROI areas of the analysis is therefore fundamental. When a fixed selection of ROI areas is employed, errors in reading fluorescence are generated. In order to solve this problem has been implemented a new flow process, integrating some different algorithm and creating smart decision system which detect automatically ROI areas, computing average values on that areas and calculate PCR for every cycle [9-12].
Here we present the miniaturized Bio2Bit Q3 Real time PCR platform able to perform quantitative and qualitative identification of multiple analytes of specific nucleic acids (DNA, RNA) sequences in a PoC format. The platform is based on silicon microchip that integrates in the same device both real time PCR micro reactors (6-15 μl in volume) and silicon temperature sensors and heaters. Thanks to the integrated silicon temperature sensors and heaters, the chip allows a temperature control accuracy of ±0.2°C, heating rate of 15°C/s and cooling rate of 8°C/s. Additionally, a specific chip design architecture [13] enhances the optical flouroscence signal in combination with a smart detection software, so that an improvement of sensitivity is achieved respect to the commercial tools. The chip is thermally and optically driven by a miniaturized instrument specifically developed for it design.
Here we report real time PCR applicative results related to the detection of hepatits B virus (HBV) using a specific HBV clone as sample material.
Materials and Methods
qRTPCR platform
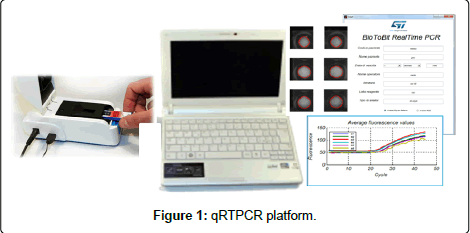
The real time PCR System is composed by the following components: (a) a disposable chip that contains reaction chambers for the real time PCR process; (b) a portable and customized thermacycler instrument (Q3 reader) that thermally and optically drives the chip during the real time PCR process; (c) a software package that manages the PCR process and carried out the data analysis.
Figure 1 reports the picture of the platform.
Microchip fabrication
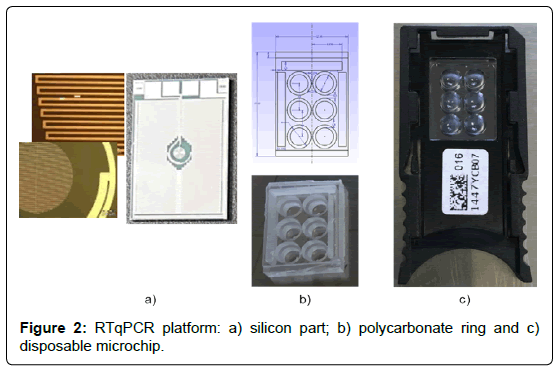
The core of the platform is a silicon-plastic hybrid microchip (Figure 2) composed by:
(a) A bottom part made in silicon by the VLSI (Very-large-scale integration) technology on a six inch silicon wafer that contains the temperature sensors and heaters integrated on the bottom of the device (Figure 2a). They consist in AlCu metal strips (size 4.0 ± 0.1, spaced 5.0 ± 0.1 μm for the temperature sensor, size 115.0 ± 0.1 μm spaced 30.0 ± 0.1 μm for the heater). The device perform thermal cycling in combination with the driving Instrument (Q3-reader below described) with the following performances: temperature sensor maximum power consumption 55 mW, heater maximum power consumption 10 mW, temperature control accuracy of ± 0.2°C, heating rate of 15°C/s, cooling of rate: 8°C/s and temperature resolution: 0.1°C.
(b) A top part (ring) made in polycarbonate. The ring was manufactured by molding technology and comprises an architecture containing 6 reaction chambers with cylindrical shape featured by a volume of 25 μL (Figure 2b). The ring is glued onto the silicon part by silicone glue. The microchip is finally mounted on a plastic holder to facilitate the chip handling (Figure 2c).
During the PCR cycles the thermal sealing is guarantee by 15 μl of paraffin wax dispensed on each chamber.
Q3 Reade
The Q3 instrument reader was developed by STMicroelectronics (Figure 1). It thermally and optically drives the chip during the real time PCR process. It has a size of 14 L × 7.1 W × 8.7 H (cm) (weight: 322 g). The optical module contains up to 2 independent optical channels, for multiple Fluorescent reporters (FAM, VIC®). The LED light sources are centered at wavelength 470, 530 nm, respectively. The emission is detected by a CMOS camera that contains a high-pass filter at 520 nm or 20 nm wide band-pass filter centered at 556. Finally, the thermal Module drives the temperature sensor and heater of the chip to reach the temperature performance characteristics described in the section 2.2.
Smart-software for image analysis
The smart-software was properly developed for the image analysis, it is based on three main core modules: 1) Fully automatic image quantification: a set of algorithms for the digital image analysis process; 2) Numerical computation: a set of algorithms for the numerical analysis of raw data coming from real time PCR, 3) Graphical output: a graphical user interface to show the fluorescence curves for the PCR.
Chemicals
Hepatitis B virus (HBV) clone (ref. product CLO-05960116 HBV Complete Genome) and all the reagents for the HBV Real time PCR were purchased from Clonit (kit ref. product CLO-FO2 HBV MMIX KIT 48) and used according to the instruction for use.
RTPCR experimental set up
Real time PCR experiments were performed using a Master Mix solution (volume 10 μL) containing Clonit buffer (1×) and Taq DNA polymerase, 0.5 μM of each forward and reverse primers, 2 μL of HBVclone (105 copies/μL). Different amount of probe (0.01, 0.05, 0.1 and 0.2 μM) were tested. The PCR cycling was performed in the portable Q3-thermocycler-reader by using the following thermal program: preheating period of 10 min. at 95°C, followed by a maximum of 45 cycles of 95°C for 15s and 60°C for 60s. Different temperature of annealing were used for testing (60°C -63°C). Negative samples were prepared substituting the HBV clone with the same amount of water. Each experiment was repeated 10 times.
Same real time PCR experiments (10 μL of the above reported Master Mix) were executed on standard 96 well reaction plate in gold standard Applied Biosystem 7500 real time PCR equipment as reference.
Results and Discussion
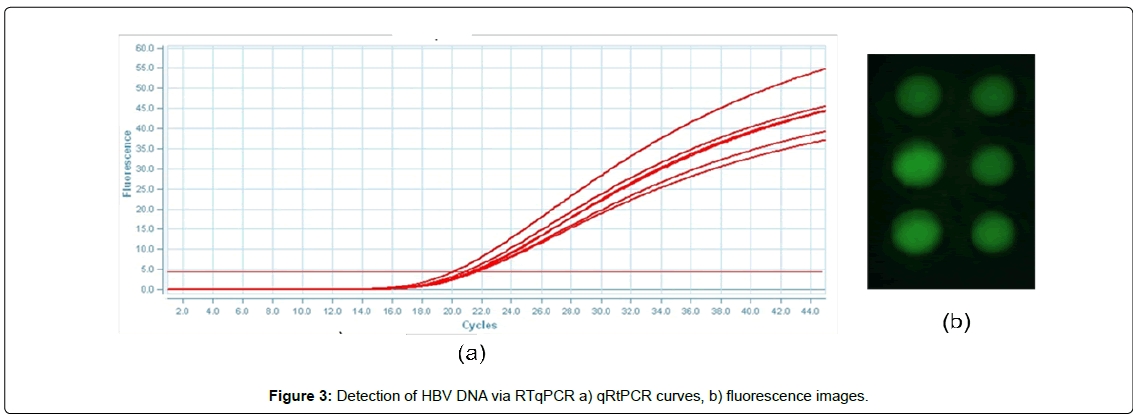
In order to demonstrate the ability of the system to detect the pathogen species HBV, we conduct real time PCR experiments with different conditions. In particular in order to optimize the device performance, we explored different annealing temperatures and probe concentrations. Figure 3a and 3b reports a typical qRT-PCR curves and fluorescence images obtained during the above described experiments
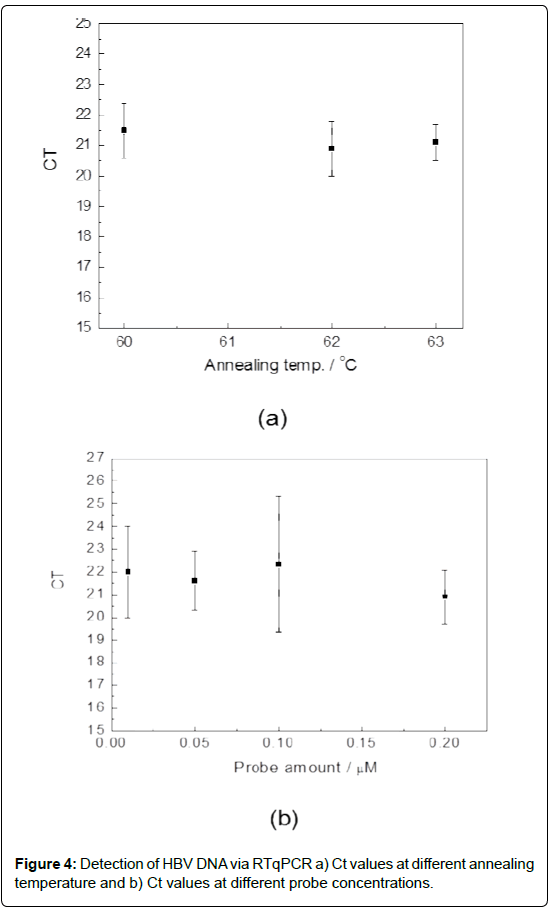
More in details, an amount of 15 μl of PCR master mix (containing HBV-clone 105 cps/μL) was loaded on each chamber (the loading process was performed at 70°C in order to melt the paraffin wax) and the amplification at three different annealing temperatures (60, 62 and 63°C) was conducted. The fluorescence images after each cycles was analyzed by the smart-software and the Ct values calculated. The results are reported in Figure 4a. The data indicate no relevant difference in term of Ct for the three temperatures investigated with an average value for Ct of about 21.5. The reference samples processed on gold standard 96-well reaction plate in Applied Biosystem 7500 equipment report for the testing at different annealing temperature an average value of Ct 22.4 ± 0.5. These data indicate an improvement on PCR efficiency performed on microchip device of about 1 Ct respect to the commercial equipment. This advance was explained by the specific microchip design architecture that include a chamber designed to collect the paraffin wax (15 μL) on the upper cylindrical part (diameter 3.5 mm) while the biological mix (10 μL) is collected on the lower zone (diameter 3.2 mm): this shape in combination with a smart detection software improve the final signal enhances the optical fluorescent signal of platform.
In order to investigate the robustness of the assay at the annealing temperature of 60°C (since no difference in the annealing temperature was found-vide supra), additional experiments were performed with different probe amounts. With the same method above described, an amount of 15 μl of PCR of master mixes containing a concentration of 105 cps/μL HBV-clone and different probe amount (0.01, 0.05, 0.1 and 0.2 μM), were loaded into the microchip pre-heated chambers and real time PCR amplifications carried out. Figure 4b reports the related experimental results. Also in this case no variation of Ct was found with an average value for Ct of about 21.3. No fluorescence signal was registered for the negative sample (data not showed). The reference samples processed on gold standard 96-well Applied Biosystem 7500 show an average value of Ct 22.7 ± 0.4. These results proves the assay is robust in the experimental condition ranges confirming the improvement of sensitivity of about 1 Ct compared to the commercial equipment. Additionally, they indicate that the platform allows to use very low amount of probe (0.01 μM) without any efficiency variation.
Conclusions
Here we presented a miniaturized platform for nucleic acids detection by Real time PCR. The core of the platform is a hybrid microchip composed by a silicon part and a polycarbonate portion mounted on the plastic ring. Each PCR chamber with a total volume of volume 25 μL is pre-loaded with 15 uL of paraffin wax to guarantee the sealing during the thermal protocol. The chamber has a particular shape suitable to accommodate an amount of 10 μL of biological master mix. The experiments were conducted at different annealing temperature and probe amounts and the results here discussed indicate an improvement of sensitivity of about 1 Ct, thanks to the smartsoftware implemented for the image analysis and to the chamber shape.
Acknowledgments
This work has been funded by MIUR by means of the national Program PON R&C 2007-2013, project “Hyppocrates-Sviluppo di Micro e Nano-Tecnologie e Sistemi Avanzati per la Salute dell'uomo” (PON02 00355) and PRIN 2010-2011 “Metodologie chimiche innovative per biomateriali intelligenti”.
References
- Heid CA, Stevens KJ, Livak J, Williams PM (1996) Real time quantitative PCR. Genome Res 6: 986-994.
- Foglieni B, Brisci A, San Biagio F, Di Pietro P, Petralia S (2010) Integrated PCR amplification and detection processes on a Lab-on-Chip platform: a new advanced solution for molecular diagnostics. Clinical chemistry and labora tory medicine 48: 329-336.
- Sun Y, Quyen TL, Hung TQ, ChinWH, Wolffa A, et al. (2015) Lab-on-a-chip system with integrated sample preparation and loop-mediated isothermal amplification for rapid and quantitative detection of Salmonella spp. in food samples. Lab Chip 15:1898-1904.
- Foudeh AM, Didar TF, Veresa T, Tabrizian M (2012) Microfluidic designs and techniques using lab-on-a-chip devices for pathogen detection for point-of-care diagnostics. Lab Chip 12: 3249-3266.
- Schumacher S, Nestler J, Otto T, Wegener M, Ehrentreich-Förster, et al. (2012 ) Highly-integrated lab-on-chip system for point-of-care multiparameter analysis. Lab Chip 12: 464-473
- Xu Y,Yan H, Zhang Y, Jiang K, Lu Y, et al. (2015) A fully sealed plastic chip for multiplex PCR and its application in bacteria identification. Lab Chip 15: 2826-2834.
- Hsieh K, Ferguson BS, Eisenstein M, Plaxco KW, Soh HT (2015) Integrated Electrochemical Microsystems for Genetic Detection of Pathogens at the Point of Care Integrated Electrochemical Microsystems for Genetic Detection of Pathogens at the Point of Care Acc. Chem Res 48: 911-920.
- Petralia S, Verardo R, Klaric E, Cavallaro S, Alessi E, et al. (2013) In-Check system: A highly integrated silicon Lab-on-Chip for sample preparation, PCR amplification and microarray detection of nucleic acids directly from biological samples. Sensors and Actuators B: Chemical 187: 99-105
- Maramis C, Delopoulos A (2010) Efficient Quantitative Information Extraction from PCR-RFLP Gel Electrophoresis Images Pattern Recognition (ICPR). 20th International Conference on Digital 2560-2563.
- Rutledge RG (2014) Sigmoidal curve-fitting redefines quantitative real-time PCR with the prospective of developing automated high-throughput applications. Nucleic Acids Research 32: 178
- Ziou D, Tabbone S (1998) Detection Techniques: An Overview.International Journal of pattern recognition and image analysis 8: 537-559.
- Spata MO, Castagna ME, Conoci S (2015) Image data analysis in qPCR: A method for smart analysis of DNA amplification. Sensing and Bio-Sensing Research 6: 79-84.
- Lazzara A, Castagna ME, Giuffrida R, Conoci S (2015) MiniaturisedLabonChip for Bio-Chem Analysis.
--
Relevant Topics
- Amperometric Biosensors
- Biomedical Sensor
- Bioreceptors
- Biosensors Application
- Biosensors Companies and Market Analysis
- Biotransducer
- Chemical Sensors
- Colorimetric Biosensors
- DNA Biosensors
- Electrochemical Biosensors
- Glucose Biosensors
- Graphene Biosensors
- Imaging Sensors
- Microbial Biosensors
- Nucleic Acid Interactions
- Optical Biosensor
- Piezo Electric Sensor
- Potentiometric Biosensors
- Surface Attachment of the Biological Elements
- Surface Plasmon Resonance
- Transducers
Recommended Journals
Article Tools
Article Usage
- Total views: 13558
- [From(publication date):
June-2016 - Aug 24, 2025] - Breakdown by view type
- HTML page views : 12375
- PDF downloads : 1183