Conference Proceeding Open Access
Biodegradation of Free Cyanide Using Bacillus Sp. Consortium Dominated by Bacillus Safensis, Lichenformis and Tequilensis Strains: A Bioprocess Supported Solely with Whey
| Lukhanyo Mekuto, Vanessa Angela Jackson and Seteno Karabo Obed Ntwampe* | |
| Faculty of Applied Sciences, Department of Agriculture and Food Science - Biotechnology, Cape Peninsula University of Technology, Cape Town, South Africa | |
| Corresponding Author : | Seteno Karabo Obed Ntwampe Faculty of Applied Sciences Department of Agriculture and Food Science-Biotechnology Cape Peninsula University of Technology, Cape Town, South Africa Tel: +27 21 460 9097 Fax: +27 21 460 3282 E-mail: NtwampeS@cput.ac.za |
| Received November 09, 2013; Accepted December 16, 2013; Published December 22, 2013 | |
| Citation: Mekuto L, Jackson VA, Ntwampe SKO (2013) Biodegradation of Free Cyanide Using Bacillus Sp. Consortium Dominated by Bacillus Safensis, Lichenformis and Tequilensis Strains: A Bioprocess Supported Solely with Whey. J Bioremed Biodeg S18:004. doi:10.4172/2155-6199.S18-004 | |
| Copyright: © 2013 Mekuto L, et al. This is an open-a ccess article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. | |
Related article at Pubmed Pubmed  Scholar Google Scholar Google |
|
Visit for more related articles at Journal of Bioremediation & Biodegradation
Abstract
Several bacterial species (n=13) were isolated from electroplating wastewater to assess their ability to biodegrade free cyanide (F-CN). A mixed culture mainly dominated by Bacillus sp (Bacillus safensis, Bacillus lichenformis and Bacillus tequilensis) was cultured in nutrient broth for 48 hours at 37°C, to which F-CN as KCN (200 to 400 mg CN-/L) was added in order to evaluate the species ability to tolerate and biodegrade the cyanide. In nutrient broth, the microorganisms were able to degrade 131(65.5%) and 177 (44.3%) mg CN-/L in cultures containing 200 and 400 mg CN-/L over a period of 8 days, respectively. Subsequently, cultures were supplemented solely with agrowaste extracts, i.e. Ananas comosus extract (1% v/v), Beta vulgaris extract (1% v/v), Ipomea batatas extract (1% v/v), spent brewer’s yeast (1% v/v) and whey (0.5% w/v), as the primary carbon sources in 200 and 400 mg CN-/L cultures. The bacterial species were able to degrade F-CN in cultures that were supplemented with whey, whereby 179 (89.5%) and 239 (59.75%) mg CN-/L was biodegraded from 200 and 400 mg CN-/L cultures, respectively.
| Keywords |
| Agro-waste; Bacillus species; Biodegradation; Free cyanide |
| Introduction |
| Cyanide is a natural compound that is produced by living organisms including bacteria, fungi, algae and plants as part of a defence mechanism against predation [1]. However, these natural sources of cyanide are insignificant when compared to cyanide production by anthropogenic activities. The mining, mineral processing, electroplating and plastics industries contribute significantly to cyanide contamination in the environment [2,3]. Over 60,000 tons of sodium cyanide was produced in 2001 and the mining industry utilised 90% for gold extraction. Cyanide is highly toxic to living organisms [4] and thus wastewater containing cyanide does not only pose a threat to aquatic organisms but also to organisms that utilise water on the mainland. Most contaminated wastewater end-up in Wastewater Treatment Plants (WWTPs) in which biological processes are used for bioremediation purposes. The presence of cyanide in the wastewater will thus result in organisms normally employed in WWTPs being susceptible to cyanide toxicity, thus rendering the biological stage of the WWTPs ineffective [5,6]. The resultant treated water from the WWTPs will therefore lead to contamination of potable water resources, as the water from the WWTPs is normally discharged into river systems, ultimately leading to ecological and environmental contamination. A number of treatment methods have been developed to remediate cyanide containing wastewaters. However, these chemical and physical methods are capital intensive and produce excess sludge which requires additional treatment. Furthermore, the by-products that are produced through these methods are hazardous. Therefore, there is a need for the development of alternative methods that are robust and economically viable for the bioremediation of cyanide contaminated wastewater. |
| Biological treatment of free cyanide in industrial wastewaters has been proven as a viable and robust method for treatment of wastewaters containing cyanide [7-9]. Several bacterial species, including Bacillus sp, can degrade cyanide to less toxic products, as these microorganisms are able to use the cyanide as a nitrogen source, producing by-products; ammonia and carbon dioxide [10]. These bacterial species secrete enzymes that catalyse the degradation of cyanide into several endproducts. The end-products of biodegradation can then be utilised by the microorganisms as nutrient sources [11]. This study focused on the biodegradation of F-CN under alkaline conditions. F-CN is the most toxic form of cyanide and its bioremediation was achieved using bacterial species that were isolated from electroplating wastewater. Furthermore, several studies are now directed towards the application of agricultural waste residue for cyanide bioremediation processes [12] and therefore the application of agricultural waste (agro-waste) as a sole supplement to support the activity of the microbial species was also assessed in this study for F-CN biodegradation. |
| Materials and Methods |
| Sample collection and microbial isolation |
| Wastewater samples were collected from an electroplating plant located in Cape Town, Western Cape, South Africa. Previous studies conducted by Santos et al. [13] have demonstrated that the electroplating wastewater is highly contaminated with high concentrations of F-CN (≥ 200 mg CN-/L) and metal-complexed cyanides. The cyanide containing wastewater was obtained from randomly selected sites from an electroplating facility. The obtained wastewater was stored in non-transparent 20 L bottles and immediately stored at 4°C to prevent ultraviolet rays (sunlight) from degrading cyanide in the collected samples. The wastewater (1L) was filtered using a 0.22 μm Millipore membrane and the bacterial cake on the filter was suspended in a phosphate buffer (pH 7). Serial dilutions were prepared using sterile distilled water subsequent to plating on nutrient agar plates containing 200 mg CN-/L as KCN and cyclohexamine (to prevent fungal growth) with subsequent incubation at 37°C for 48 hours. Colonies were selected according to cultural properties i.e. persistent growth in cyanide containing media, and transferred to nutrient agar to isolate single colonies. The selected bacteria were stored in 20% glycerol at -20°C for further research. Gram reaction, endospore staining, oxidase activity and starch hydrolysis were studied in all the isolated organisms. |
| DNA extraction, Polymerase Chain Reaction (PCR) and 16S rDNA gene sequencing |
| Thirteen bacterial species from nutrient agar plates were isolated and single colonies were inoculated in 1ml nutrient broth and incubated at 30°C. DNA was extracted from the previously grown cells using the ZR Fungal/Bacterial DNA Kit (Zymo Research, California, USA). The presence of the genomic DNA was assessed using 1% molecular grade agarose gel containing 0.5 μg/ml ethidium bromide (EtBr), using 1X Tris-acetate-ethylenediamine tetraacetic acid (TAE) electrophoresis buffer at 100V for 1 hour. |
| PCR was performed using a GeneAmp PCR 9700 System (Applied Biosystems, USA). Amplification of the target DNA by PCR was performed in a total reaction volume of 10 μl containing 0.5 μl ( ± 50 ng/μl) of the purified genomic DNA, 50 mM of the forward and reverse primers and 5 μl of a 2X KapaTaq Readymix solution (KapaBiosystems, South Africa). Bacterial specific primers used were the forward 8F primer 5’-AGAGTTTGATCCTGGCTCAG-‘3 and reverse primer 1492R 5’-GGTTACCTTGTTACGACTT-‘3 [14]. The amplification process included an initial denaturing step at 94°C for 10 min, followed by 36 cycles of 94°C for 30s, 55°C for 30s and 72°C for 1 min. The reaction was completed with a final extension period of 7 min at 72°C followed by cooling and storage at 4°C. Ten microliters of the PCR amplicons were electrophoretically analysed on a 1% molecular grade agarose gel that was stained with ethidium bromide, using 1X TAE electrophoresis buffer at 100V for 1 h, to determine whether the amplification was successful. The PCR amplicons were run on an ABI 3010xl Genetic analyser. The sequences were blasted against the NCBI GenBank database and trees were generated using ClustalX for the alignment and PAUP for the analysis. Distance analysis using the neighbour algorithm and the strength of the branches was calculated with 1000 bootstrap repetitions. |
| Nutrient media preparation and culture conditions |
| The bacterial species (n=13) were mixed and cultured in an Oxoid nutrient broth for 48 hours at 37°C using a rotary shaker at 180 rpm. This culture was used as an inoculum at a cell concentration of 10% (v/v) per batch. Initially, experiments were conducted in batch systems with F-CN concentrations of 200 and 400 mg CN-/L in nutrient broth. This was done to assess the viability of the cultures to effectively biodegrade cyanide. In order to effectively assess cyanide tolerance and culture viability on cyanide biodegradation, the isolated microbial species were assessed in cultures containing alternative carbon sources i.e. agrowaste [Ananas comosus (pineapple) extract (1% v/v), Ipomea batatas (sweet potato) extract(1% v/v) and Beta vulgaris (beetroot extract) extract(1% v/v)], spent brewer’s yeast (SBY) extract (1% v/v) and whey (0.5 w/v) as sole nutrient and energy sources for the microorganisms. The concentration of cyanide in the cultures was set at 200 and 400 mg CN-/L. All the experiments had a final volume of 100 mL in airtight multiport round bottomed flasks with a sampling port while the pH was kept at 9.5 using a phosphate buffer. Potassium cyanide (KCN) was used as a source of F-CN. The biodegraded cyanide taking into account cyanide volatilisation was calculated using the following mass balance equation: |
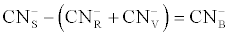 (1) (1) |
| Where |
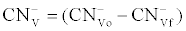 (2) (2) |
Where 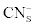 was the initial F-CN concentration in the fermentation broth (mg/L); was the initial F-CN concentration in the fermentation broth (mg/L); 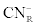 was the residual F-CN measured in the cultures (mg/L); was the residual F-CN measured in the cultures (mg/L); 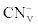 was the cyanide that volatilised during culture incubation (mg/L); was the cyanide that volatilised during culture incubation (mg/L); 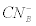 was the F-CN that was bioremediated (mg/L); was the F-CN that was bioremediated (mg/L); 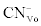 was the initial cyanide in control cultures; was the initial cyanide in control cultures; 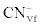 was the final cyanide concentration in the control cultures (mg/L). was the final cyanide concentration in the control cultures (mg/L). |
| Preparation of agro-waste extracts |
| Agro-waste extracts were prepared by drying the agro-waste at 80°C for 7 days followed by milling to less than 100 μm. The dried waste was mixed with water and autoclaved at 116°C for 20 minutes. The solution was filtered through a 0.22 μm Millipore filter and the debri free extract was used for the experiments at the required concentration. |
| Sampling and analysis |
| Samples (2 mL) were obtained from the flask and centrifuged at 14500 rpm for 5 minutes. The centrifuged supernatant was analysed for cyanide and ammonium concentration using Merck cyanide (CN-) (09701) and ammonium (NH4+) (00683) test kits. The residual concentrations for both cyanide and ammonium were quantified using the Merck Spectroquant Nova 60 instrument. |
| Results and Discussions |
| Isolation and characterization of cyanide-degrading bacteria |
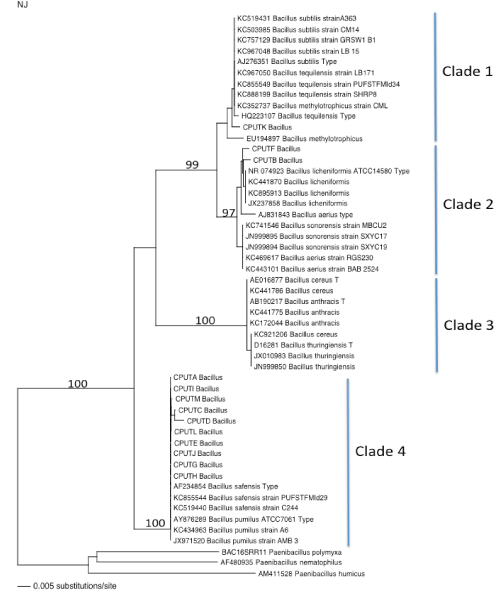
| All the bacterial isolates (labelled CPUT A to CPUT M) were slender, aerobic, oxidase positive, gram positive and endospore-forming rods. The spores were located centrally in all the isolates. All the isolates could not hydrolyze starch, which was indicative of organisms that belong to the Bacillus genus as described by Meyers et al. [10]. The phylogeny of the representative organisms in the GenBank was analysed using the Neighbour-joining algorithm in ClusterX (Figure 1). The phylogenetic tree was constructed using organisms that were a representative of the isolates. |
| The sequence of the bacterial species labelled CPUT-K was found to belong to the Bacillus subtilis group (Figure 1, Clade 1), while organisms labelled CPUT-F & B were found to belong the Bacillus licheniformis group (Figure 1, Clade 2). The organisms labelled CPUT-A, C, D, E, G, H, I & J were identified as species belonging to the Bacillus safensis/pumilus group (Figure 1, Clade 4). Since the 16S rDNA gene sequence does not distinguish between species belonging to the Bacillus genera, more especially to Bacillus safensis and Bacillus pumilus species, it is advisable to use the gyrase gene (gyrB gene) for the identification of species belonging to the Bacillus genus, especially to the Bacillus subtilis group as this is the most efficient method of distinguishing between Bacillus species [15]. |
| However, the gyrB gene does not differentiate between Bacillus safensis and Bacillus pumilus as these organisms share 92% of the gyrB sequence similarity [16]. Meyer et al. [10] investigated the capability of the isolated cyanide-adapted Bacillus pumilus to degrade free cyanide and observed that the organism was able to degrade 100 mg CN-/L in 2 hours. Furthermore, Bacillus pumilus was found to possess the cyanide dihydratase enzyme which utilises the hydrolytic pathway for degradation of F-CN [10,17]. There are currently no reports on the degradation of F-CN by Bacillus subtilis and Bacillus licheniformis. However, it has been reported that Bacillus subtilis is able to degrade organic cyanides (nitriles) through the action of nitrile hydratase [18]. Additionally, it has been reported that the organism is able to express the nitrilase enzyme [18] and this enzyme has been observed to facilitate the degradation of F-CN [19,20]. |
| Biological degradation of free cyanide |
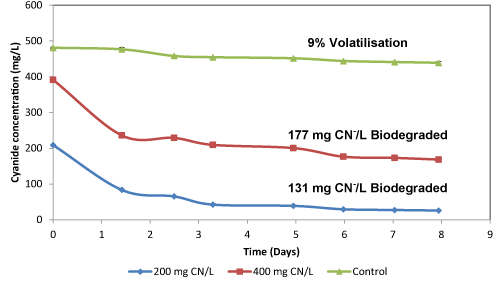
| The mixed culture of the 13 isolates was prepared and assessed for the degradation of F-CN under alkaline conditions in nutrient broth. The residual F-CN in the medium containing 200 and 400 CN-/L after 8 days was 26 and 170 CN-/L respectively, with degradation efficiency of 65.5% and 44.3%, respectively (Figure 2a). Using Equation. 1, it was observed that the microorganisms were able to degrade 131 and 177 mg CN-/L of F-CN from 200 and 400 mg CN-/L of F-CN solutions, respectively, where it was observed that the spore forming Bacillus species were able to withstand and degrade F-CN concentration in cultures containing F-CN concentration of up to 400 mg CN-/L. It has been reported that the maximum concentration of F-CN that microorganisms can tolerate and be able to degrade is 200 mg CN-/L [21]. However, this report has proven that the microbial community employed in this system were able to degrade and tolerate cyanide concentrations of 400 mg CN-/L. This is a first report to date on bacterial species that are able to tolerate and degrade F-CN efficiently in water containing such cyanide concentration. Cabuk et al. [22] evaluated the ability of the white rot fungus Trametes versicolor to degrade F-CN at a maximum cyanide concentration of 400 mg CN-/L, where the degradation efficiency was less than 30% (Table 1). Furthermore, the immobilised cells of Klebsiella oxytoca were evaluated for the degradation of F-CN at a maximum concentration of 157 mg CN-/L where the microbe had obtained a degradation efficiency of 26% when cellulose acetate (CTA) was used [19]. Furthermore, in studies conducted by Cabuk et al. [22] and [19] aeration was used; however, it has been reported that the introduction of air in cyanide biodegradation systems results in the volatilisation of cyanide [23]. Therefore, the work that was reported as being due to biological degradation might be due to volatilization of cyanide and not biodegradation. |
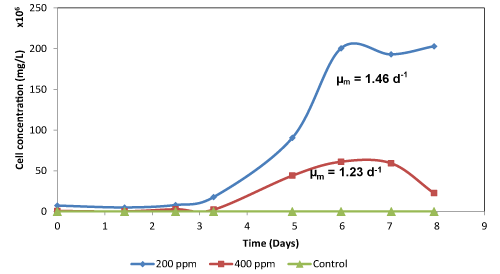
| In this study, the degradation of cyanide was accompanied by increasing microbial growth, with cultures containing 200 mg CN-/L showing high growth resulting in cell concentration exceeding 2×108 CFU/mL with the maximum specific growth rate of 1.46d-1, while the medium containing 400 mg CN-/L had a maximum cell concentration of 6.12×107 CFU/mL with a maximum growth rate of 1.23d-1. The microbial growth was highly dependent on the initial cyanide concentration in the cultures, with exponential growth being observed during day 3 to 6 in both the cultures containing 200 and 400 mg CN-/L. Generally, the growth rate decreased as the cyanide concentration was increased in the cultures. As a result of this, the lag phase for cellular multiplication in the cultures increased (Figure 2b). The bacterial isolates were able to degrade cyanide in media solely supplemented with agro-waste. However, the degradation efficiency was poor in media that were supplemented with B. valgaris, I. batatas, A. comosus and spent brewer’s yeast extracts at all the cyanide concentrations evaluated (Table 2).The biodegraded cyanide using cultures with an initial cyanide concentration of 200 mg CN-/L, for the B. valgaris, I. batatas, A. comosus and spent brewer’s yeast extracts was 27 ± 1.25, 47 ± 1.08, 60 ± 0.92 and 32 ± 0.78 mg CN-/L, respectively. Similarly, in cultures containing 400 mg CN-/L, only 23 ± 1.08, 23 ± 1.25, 67 ± 1.58 and 18 ± 1.25 mg CN-/L, respectively, were biodegraded (Table 2). The degradation efficiency using these waste materials for microbial support was below 40% (Table 2), with the media that was supported with A. comosus extract showing the highest degradation efficiency of approximately 38%. The high degradation efficiencies experienced in the media that were supported by A. comosus extract are largely due to the high reducing sugars present in the extract [12]. |
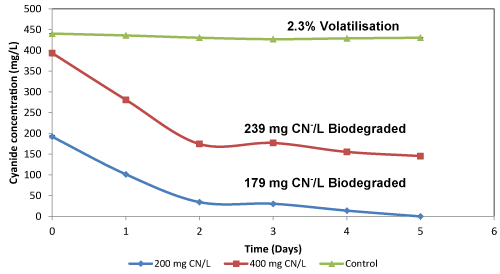
| For cultures supplemented with whey, the biodegradation efficiency was significantly increased. The residual cyanide in the media with 200 and 400 mg CN-/L was 0.009 mg CN-/L and 143 mg CN-/L, respectively after 5 days. This was translated into degradation efficiencies of 90% and 60% respectively (Figure 2d). The biodegraded cyanide from both cultures was 179 and 239 mg CN-/L, respectively. The degradation of cyanide in both the nutrient broth and whey-supplemented medium occurred within a period of 3 days. It is hypothesised that the degradation was due to the expression of cyanide degrading enzymes in response to the high cyanide concentrations at the beginning of the experiments. Similarly, Akcil et al. [24] evaluated the ability of the isolated Pseudomonas sp. to degrade high cyanide (Weak Acid Dissociable, WAD) concentrations (400 mg CNWAD/L) and observed that the species were able to degrade cyanide efficiently, with degradation efficiency of approximately 90% occurring within a period of less than 5 days. However, the results presented did not contain any control for comparison purposes since cyanide is highly volatile. Furthermore, the use of shake flasks introduces aeration into the system, thus resulting in the chemical oxidation of cyanide [23]. |
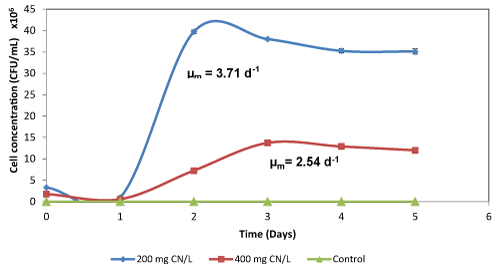
| The maximum cell concentration in a medium supplemented with whey at 200 mg CN-/L was found to be 3.93×107 CFU/mL, while the maximum cell concentration at 400 mg CN-/L was 1.36×107 CFU/mL (Figure 2e). The observed maximum growth rate in both culture media containing 200 and 400 mg CN-/L was 3.71 d-1 and 2.54°d-1, respectively. The cell growth in a whey containing 400 mg CN-/L was similar to cell growth in the medium that contained less cyanide concentration, which suggested whey as a suitable substrate for the biodegradation of cyanide. The cell concentration observed was similar to that obtained using nutrient broth, with a reduced lag phase period. Although the organisms were able to grow well on the agro-waste extracts (Table 2), the media that were supported with A. comosus extract showed high cell concentrations, with a maximum cell concentration of 1.25×107 CFU/mL and cyanide degradation of 37.6% (60 to 67 mg CN-/L) which was comparable to the low F-CN degradation efficiencies that were observed in several studies [19] where refined carbon sources were utilised for the degradation of cyanide (Table 2). However, this was not observed for SBY, B. vulgaris and I. batatas supplemented cultures. |
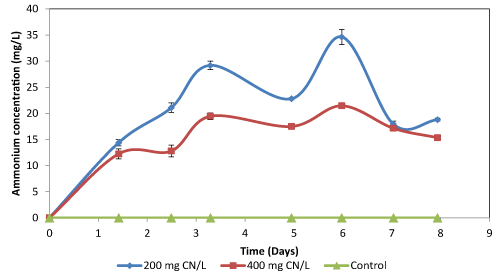
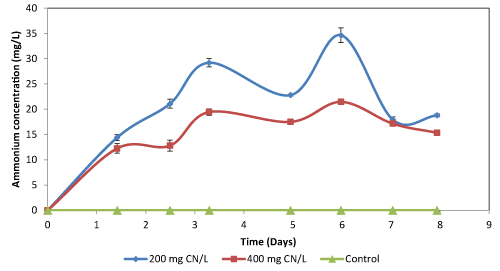
| The residual ammonium concentration in nutrient broth fluctuated throughout the experiments, with the highest concentration being below 40 mg NH4+/L (Figure 2c and 2f). For cultures grown using broth, the amount of ammonium formed was dependent on the amount of cyanide that was biodegraded. The fluctuating concentration of ammonium was hypothesised to be largely dependent on ammonium utilisation by the microorganisms. It was however observed that the high ammonium concentration in the broth inhibited cyanide biodegradation as the organisms prefer the ammonium rather than cyanide. The residual ammonium concentration in cultures supplemented with agro-waste extracts was high (Table 2). Since the microorganisms preferred the ammonium as a source of nitrogen than cyanide, which resulted in low cyanide degradation efficiency as the microbial species utilised the ammonium already present in the culture broth rather than the cyanide. The concentration of ammonium in all the flasks fluctuated with an observed concentration range of 20 to 90 NH4+/L. The fluctuating ammonium concentration was due to formation and subsequent degradation of the ammonium in the cultures. The degradation of both cyanide and ammonium by the Bacillus sp. showed that these endospore-forming organisms are both cyanide-degraders and nitrifiers, and therefore, can be used in the reduction of residual ammonia. |
| Conclusion |
| All the isolated microorganisms from the cyanide containing wastewater belonged to the Bacillus genus. These microorganisms were found to be able to tolerate and degrade high concentrations of F-CN. This is a first report of bacterial species that can tolerate and degrade F-CN concentrations of 400 mg CN-/L. Cyanide biodegradation was successful even at elevated concentrations, with subsequent production of ammonium. However, the presence of ammonium was found to have an inhibitory effect on cyanide degradation as the microorganisms preferred utilising the ammonium over cyanide, thus reducing the efficacy of the cyanide biodegradation process. This phenomenon requires further studies on the application of these cyanide degrading microorganisms for the assimilation of ammonium that is generated through cyanide biodegradation. However, the formation of ammonium from cyanide degradation is of concern. It was observed that the cyanide-degrading organisms can alternate their metabolism depending on either the presence of cyanide or ammonium, thus utilising both substrates by activation/deactivation of metabolic pathway depending on the substrate that is present. The application of the same microbial species for both cyanide biodegradation and nitrification stages can add value to the designed process, increasing process efficiency and thus lowering operational costs of the process. The application of agrowaste, particularly whey-containing waste from the dairy industry, has been shown to successfully sustain microbial activity which in-turn resulted in an efficient process for the bioremediation of F-CN. |
| Acknowledgement |
| This research was sponsored by the Cape Peninsula University of Technology, University Research Fund (URF RK16). |
References
|
Tables and Figures at a glance
| Table 1 | Table 2 |
Figures at a glance
 |
 |
 |
 |
| Figure 1 | Figure 2a | Figure 2b | Figure 2c |
 |
 |
 |
| Figure 2d | Figure 2e | Figure 2f |
Relevant Topics
- Anaerobic Biodegradation
- Biodegradable Balloons
- Biodegradable Confetti
- Biodegradable Diapers
- Biodegradable Plastics
- Biodegradable Sunscreen
- Biodegradation
- Bioremediation Bacteria
- Bioremediation Oil Spills
- Bioremediation Plants
- Bioremediation Products
- Ex Situ Bioremediation
- Heavy Metal Bioremediation
- In Situ Bioremediation
- Mycoremediation
- Non Biodegradable
- Phytoremediation
- Sewage Water Treatment
- Soil Bioremediation
- Types of Upwelling
- Waste Degredation
- Xenobiotics
Recommended Journals
Article Tools
Article Usage
- Total views: 18521
- [From(publication date):
specialissue-2014 - Aug 16, 2025] - Breakdown by view type
- HTML page views : 13790
- PDF downloads : 4731
