Research Article Open Access
Biodegradation of Total Petroleum Hydrocarbon by Aerobic Heterotrophic Bacteria Isolated from Crude Oil Contaminated Brackish Waters of Bodo Creek
| Ichor T*, Okerentugba PO and Okpokwasili GC | |
| Department of Microbiology, University of Portharcourt, Choba, Rivers State, Nigeria | |
| Corresponding Author : | Ichor T Department of Microbiology University of Portharcourt Choba Rivers State, Nigeria Tel: 234 (0)84 817 E-mail: smartichor2012@gmail.com |
| Received July 05, 2014; Accepted July 31, 2014; Published August 04, 2014 | |
| Citation: Ichor T, Okerentugba PO, Okpokwasili GC (2014) Biodegradation of Total Petroleum Hydrocarbon by Aerobic Heterotrophic Bacteria Isolated from Crude Oil Contaminated Brackish Waters of Bodo Creek. J Bioremed Biodeg 5:236 doi:10.4172/2155-6199.1000236 | |
| Copyright: © 2014 Ichor T, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. | |
Related article at Pubmed Pubmed  Scholar Google Scholar Google |
|
Visit for more related articles at Journal of Bioremediation & Biodegradation
Abstract
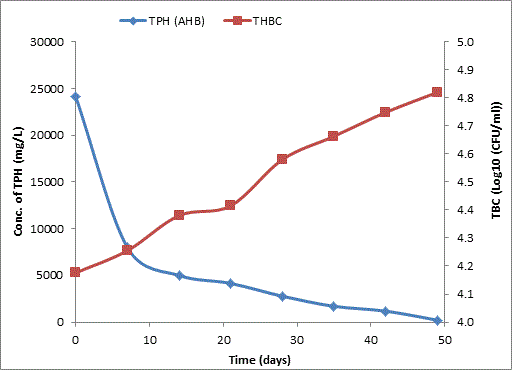
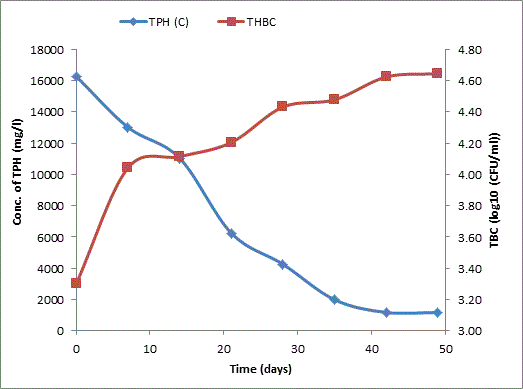
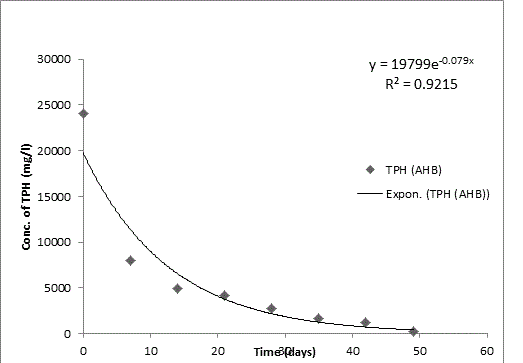
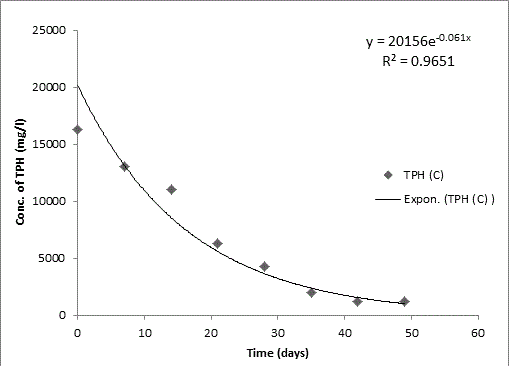
Bodo creek is located in Gokana Local Government Area of Rivers state and is characterised by brackish water which is heavily contaminated with crude oil. Water samples collected from the creek were taken to Environmental Microbiology Laboratory of University of Portharcourt for isolation of aerobic heterotrophic bacteria using serial dilution spread plate technique. Isolation of morphologically distinct pure cultures was done and the isolates obtained were identified molecularly on the basis of 16S rRNA gene sequence analysis. 16S rRNA sequencing for identification of the isolates generated sequences ranging from 500 bp to 1700 bp and a 250 bp size PCR amplified fragment. The nucleotide sequence of the 16S rRNA gene was compared with published 16S rRNA sequences using BLAST search at the data base of National Centre for Biotechnology Information (NCBI). Biodegradation of petroleum hydrocarbon test on the isolates was conducted using Bony light crude oil obtained from Shell Petroleum Development Company and monitored using GC-FID (Agilent 6890 model) for 49 days. The initial quantity of TPH was 24091 and 6706 mg/l on day 0 for aerobic heterotrophic treatment and crude oil contaminated control and decreased progressively to 212.8 and 1174 mg/l respectively on day 49 indicating biodegradation in the treatment and control. Loss of TPH was statistically significant using one way analysis of variance (ANOVA) p<0.05 with time. Model for predicting TPH loss was developed and rate of biodegradation of the isolates determined using Trend line method of Microsoft Excel, 2010. The growth of bacteria cells increased progressively with decrease in TPH implying that the bacteria were responsible for biodegradation. Further application of bioremediation strategies in Bodo creek for biostimulation of the crude oil biodegrading bacteria for reclamation and restoration of an efficient ecosystem structure and function is suggested.
Biodegradation of petroleum hydrocarbon in the environment is a complex process. The quantitative and qualitative aspects depend on the nature and amount of the oil or hydrocarbon present, the composition of autochthonous microbial community [5,6]. Hydrocarbon degrading bacteria are widely distributed in marine freshwater, soil habitat and their use in bioremediation of hydrocarbon contaminated soils, which exploits their ability to degrade and/or detoxify organic contaminants, has proven to be an efficient, economical, versatile and environmentally friendly treatment [7].
There is a broad spectrum of distribution of bacteria that have ability to degrade or transform hydrocarbons [8]. [5,9,10] isolated varying types of bacteria from brackish water capable of petroleum hydrocarbon degradation and these include members of the genera Achromobacter, Acinetobacter, Alcaligenes, Arthrobacter, Bacillus, Brevibacterium, Corynebacterium, Enterobacter, Escherichia, Flavobacterium, Norcadia, Pseudomonas, Staphylococcus, and Vibrio.
Brackish waters have a higher percentage of salinity over freshwater and the salt concentrations are reported to constitute a stressful agent for most of the known living organisms. Organisms that grow best in such environment are called halophiles, while the ones that grow in non-saline environments but are able to grow in hypersaline environments are regarded as halotolerant organisms. Bodo creek of Bodo community in Ogoniland of Gokana LGA of Rivers State is characterised by brackish water and has been contaminated with petroleum hydrocarbons due to oil exploration activities and spillages. Gokana LGA is over the major oil producing areas in Rivers state and is located within the south east senatorial district of the state, comprising both the riverine and upland communities [2]. The aim of our study was to isolate and identify aerobic heterotrophic bacteria from brackish waters of hydrocarbon polluted Bodo creek and test them for biodegradation potentials, which justifies the relevance of the study.
Biodegradation activities were monitored for a period of 49 days for petroleum hydrocarbon degradation using Gas chromatography.
16S rRNA sequencing for identification of the isolates generated sequences ranging from 500 bp to 1700 bp and a 250 bp size PCR amplified fragment. The nucleotide sequence of the 16S rRNA gene was compared with published 16S rRNA sequences using BLAST search at the data base of National Centre for Biotechnology Information (NCBI).
The result of the study clearly demonstrated the ability of the consortium of the isolated aerobic heterotrophic bacteria to degrade total petroleum hydrocarbon which justifies the essence of their presence in this crude oil contaminated brackish water habitat. Previous studies have reported on biodegradation of petroleum hydrocarbons in both terrestrial and marine environment under oxic conditions [5,24-26].
The bacteria isolated and identified with their accession numbers include; Alcaligenes faecalis (KF056900.1), Bacillus pumilus (KF717600.1), Bacillus pumilus (JX680132.1), Bacillus pumilus (HQ334985.1), Myroides odoratimimus (GU186112.2) Bacillus cohnii (KC813164.1), Citrobacter murliniae (KF254752.1), Bacillus pumilus (HM744710.1), Bordetella sp (KF601910.1), Lysinibacillus fusiformis (KC329831.1), Bacillus pumilus KF387715.1), Myroides pelagicus (NR041042), Bacillus pumilus (JQ833612.1) and Enterobacter cloacae (JQ832514.1) and belonged to the families of Enterobacteriaceae, Bacilliceae, Alcaligenaceae, Pseudomonadaceae, Flavobactericeae and Planococcaceae which members have been implicated in petroleum hydrocarbon biodegradation even at higher salinities compared to the to the present area under study [27]. Microbial degradation is the major and ultimate natural mechanism by which petroleum hydrocarbon pollutants can be cleaned from the environment [17,28,29]. It is proven by researchers to be more effectively done by a consortium of bacteria which out performs single species isolated even from the consortium and can occur under oxic and non oxic conditions [30].
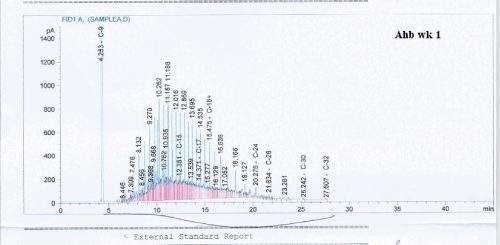
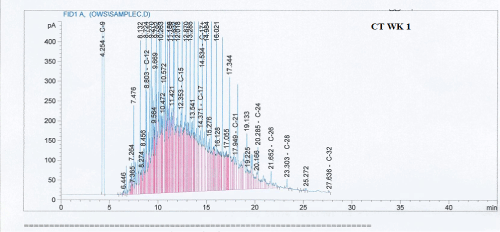
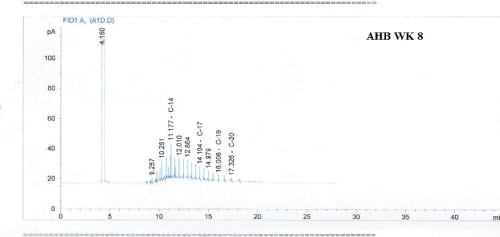
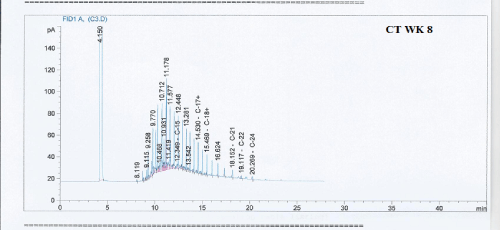
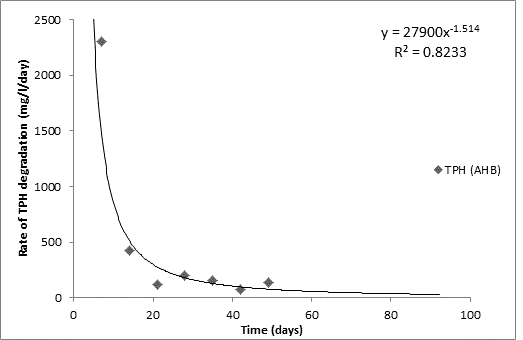
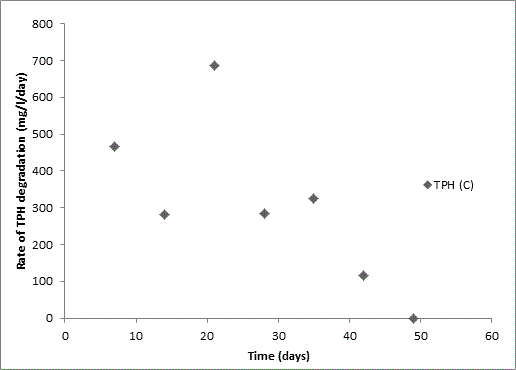
Total petroleum hydrocarbon degradation as observed in this study provides evidence that the consortium of bacteria isolated from the crude oil contaminated and the uncontaminated site used as control both possess an inherent capability for biodegradation of petroleum as monitored using GC-FID. This corroborates report of on the isolation of hydrocarbon degraders from environment [30-32]. The reduction in TPH was more fiercely observed for the treatment and control during the first seven days as observed from Figure 5 and 6 implying that volatile components of the crude oil had evaporated. TPH loss was statistically significant (p<0.05) with time for the treatment and control throughout the experimental period using one way analysis of variance though samples on day 42 and 49 showed no decrease in TPH.
The growth of bacteria in the treatment and control increased progressively with decrease in TPH throughout the 49 days period monitored. The continued bacteria growth in the control irrespective of stalled biodegradation of TPH on day 42 and 49 could be attributed to other favorable environmental factors, presence of limiting nutrients which were adjusted to mimic the natural environmental conditions from which the bacteria were isolated and other metabolites suspected to have been generated during the biodegradation which our subsequent research seeks to unravel. A negative correlation between growth of aerobic heterotrophic bacteria and TPH loss was observed for both the treatment and the control (p<0.05) implying that bacterial growth increased with decreasing quantity of TPH for the treatment and the control. Latha and Kalaivani [33] reported a correlation between increased oil degradation to an increase in cell number of bacterial indicating that the isolates were responsible for the degradation which corroborates the result of our finding.
The rate of biodegradation has been widely reported to be affected by a number of factors. Our study considered these factors and adjusted our set up to reflect the factors at the site where the pollution occurred and samples were obtained in order to ascertain near accuracy the activities of the autochthonous bacteria at the pollution site. The result established the fact that bacteria isolated from crude oil polluted Bodo creek have efficient ability for utilizing crude oil as a source of carbon and energy. Application of bioremediation strategies such as biostimulation and bioaugmentation as reported in various laboratory researches should be applied to field experiment to breach the gap between bench findings and real solutions to environmental challenges that stimulated the research initially for reclamation of such environment for efficient ecosystem balance and function.
References
- Okoh A1, Ajisebutu S, Babalola G, Trejo-Hernandez MR (2001) Potential of Burkholderiacepacia RQ1 in the biodegradation of heavy crude oil.IntMicrobiol 4: 83-87.
- UNEP (2011) Environmental setting in Ogoniland and the Niger Delta: Environmental Assessment of Ogoniland. United Nations Environment Programme, Nairobi, Kenya, 30-33.
- Mishra S, Jyot J, Kuhad RC, Lal B (2001) Evaluation of inoculum addition to stimulate in situ bioremediation of oily-sludge-contaminated soil.Appl Environ Microbiol 67: 1675-1681.
- O’Reilly KT, Magaw RI,Rixey WG (2001) Predicting the Effect of Hydrocarbon and Hydrocarbon-Impacted Soil on Groundwater. American Petroleum Institute.
- Leahy JG, Colwell RR (1990) Microbial degradation of hydrocarbons in the environment.Microbiol Rev 54: 305-315.
- Atlas RM (1995) Petroleum biodegradation and oil spill bioremediation. Marine Pollution Bulletin 31:178-182.
- Margesin R, Schinner F (1997) Efficiency of indigenous and inoculated cold-adapted soil microorganisms for biodegradation of diesel oil in alpine soils.Appl Environ Microbiol 63: 2660-2664.
- Desai A,Vyas P (2006) Hydrocarbon degradation. Applied Microbiology: Petroleum and Hydrocarbon Microbiology. M.S. University of Baroda Vadodara. 1-22.
- Foght JM, Westlake DWS (1987) “Biodegradation of hydrocarbons in freshwater,” In: Oil in Freshwater: Chemistry, Biology, Countermeasure Technology, (Vandermeulen, J. H. and Hrudey, S. B. Eds.), Pergamon Press, New York, NY, USA, 217-230.
- Atlas RM,Bartha R (1992) Hydrocarbon biodegradation and oil spill bioremediation. Advances in Microbial Ecology 12: 287-338.
- Cheesbrough M (2002) District Laboratory Practice in Tropical Countries 1 & 2. Cambridge University Press, 614-661.
- Abed RMM (2010)Interaction between cyanobacteria and aerobic heterotrophic bacteriain the degradation of hydrocarbons. International Biodeterioration and Biodegradation64:58-64.
- Moore DM,Dowhan D(2002) Current Protocols in molecular biology. John Wiley and Sons inc.
- King C, Scott-Horton T (2008) Pyrosequencing: a simple method for accurate genotyping.J Vis Exp .
- Pruesse E, Quast C, Knittel K, Fuchs BM, Ludwig W, et al. (2007) SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB.Nucleic Acids Res 35: 7188-7196.
- Abed RMM, Koster J (2005) The direct role of aerobic heterotrophic bacteria associated with cyanobacteria in the degradation of oil compounds. InternationalBiodeterioration and Biodegradation 55:29-37.
- Jones DM, DouglasAG, ParkesRJ, TaylorJ, Giger, W et al. (1983) The recognition of biodegraded petroleum-derived aromatic hydrocarbons in recent marine sediments. Marine Pollution Bulletin 14: 103-108.
- Adebusoye SA, Ilori MO, AmundOO, Teniola OD,OlatopeSO (2007) Microbial degradation of petroleum hydrocarbons in a polluted tropical stream.World Journal of Microbiology and Biotechnology 23. 1149–1159.
- Bartha R,Bossert I (1984) The treatment and disposal of petroleum wastes. (Atlas R.M. edn) Petroleum Microbiology. New York, NY, USA,Macmillian553-578.
- Rahman KS, Rahman TJ, Kourkoutas Y, Petsas I, Marchant R, et al. (2003) Enhanced bioremediation of n-alkane in petroleum sludge using bacterial consortium amended with rhamnolipid and micronutrients.BioresourTechnol 90: 159-168.
- Brooijmans RJ, Pastink MI, Siezen RJ (2009) Hydrocarbon-degrading bacteria: the oil-spill clean-up crew.MicrobBiotechnol 2: 587-594.
- Yakimov MM1, Timmis KN, Golyshin PN (2007) Obligate oil-degrading marine bacteria.CurrOpinBiotechnol 18: 257-266.
- Uzoigwe CI,Okpokwasili GC (2012) Biodegradation of oil spill dispersant in natural aquatic ecosystem. International Journal of Physical Sciences 7:5477-5484.
- Luiz FM, Raquel SP (2012) Biodegradation of petroleum hydrocarbons in hypersaline environments. Brazilian Journal of Microbiology 43:865-872.
- Perez-Pantola D,Gonzále B, Pieper DH (2010) Microbial Degradation of aromatic Hydrocarbons:Hand Book of Hydrocarbon and and Lipid Microbiology.(Timmis KN., McGenity T.J., van der Meer J.R., de Lonrezo V. Berlin Heidelbergeds), 909-924.
- Van Hamme JD1, Singh A, Ward OP (2003) Recent advances in petroleum microbiology.MicrobiolMolBiol Rev 67: 503-549.
- McGenity TJ, Folwell BD, McKew BA, Sanni GO (2012) Marine crude-oil biodegradation: a central role for interspecies interactions.AquatBiosyst 8: 10.
- Amund OO,Nwokoye N (1993) Hydrocarbon potentials of yeast isolates from a polluted Lagoon. Journal of Scientific Research and Development 1: 65-68.
- Lal B1, Khanna S (1996) Degradation of crude oil by Acinetobactercalcoaceticus and Alcaligenesodorans.J ApplBacteriol 81: 355-362.
- Wang J, Xu H, An M, Yan G (2008) Kinetics and characteristics of phenanthrene degradation by a microbial consortium. Petroleum Science5: 73-78.
- Ron E, Rosenberg E (2010)Acinetobacter and Alkannindiges. In Hand Book of Hydrocarbon and and Lipid Microbiology, (Timmis, K.N., McGenity, T.J., van der Meer, J.R., de Lonrezo, V. Berlin Heidelbergeds), 1800-1803.
- vanBeilen JB1, Funhoff EG (2007) Alkane hydroxylases involved in microbial alkane degradation.ApplMicrobiolBiotechnol 74: 13-21.
- Latha R,Kalaivani R (2012) Bacterial degradation of crude oil by gravimetric analysis. Advances in Applied Science Research 3:2789-2795.
Figures at a glance
 |
 |
 |
 |
 |
 |
| Figure 1 | Figure 2 | Figure 3 | Figure 4 | Figure 5 | Figure 6 |
 |
 |
 |
 |
| Figure 7 | Figure 8 | Figure 9 | Figure 10 |
Relevant Topics
- Anaerobic Biodegradation
- Biodegradable Balloons
- Biodegradable Confetti
- Biodegradable Diapers
- Biodegradable Plastics
- Biodegradable Sunscreen
- Biodegradation
- Bioremediation Bacteria
- Bioremediation Oil Spills
- Bioremediation Plants
- Bioremediation Products
- Ex Situ Bioremediation
- Heavy Metal Bioremediation
- In Situ Bioremediation
- Mycoremediation
- Non Biodegradable
- Phytoremediation
- Sewage Water Treatment
- Soil Bioremediation
- Types of Upwelling
- Waste Degredation
- Xenobiotics
Recommended Journals
Article Tools
Article Usage
- Total views: 16080
- [From(publication date):
September-2014 - Aug 28, 2025] - Breakdown by view type
- HTML page views : 11165
- PDF downloads : 4915
