Research Article Open Access
Biodegredation of Reactive and Reactive Disperse Dyes by Aspergillius nigar
| Fatma A Mohamed1*, N FAli1, RSR EL-Mohamedy2 and AA Hebeish1 | |
| 1 Dyeing and Printing Department, National Research Center, Dokki, Giza, Egypt | |
| 2 Plant Pathology Department, National Research Center, Dokki, Giza, Egypt | |
| Corresponding Author : | Fatma A Mohamed Dyeing and Printing Department National Research Center, Dokki, Giza, Egypt Tel: +202-3370931 Fax: +2023370951 E-mail: aali_04@hotmail.com |
| Received October 24, 2013; Accepted February 03, 2014; Published February 10, 2014 | |
| Citation: Mohamed FA, FAli N, EL-Mohamedy RSR, Hebeish AA (2014) Biodegredation of Reactive and Reactive Disperse Dyes by Aspergillius nigar. J Bioremed Biodeg 5:215. doi:10.4172/2155-6199.1000215 | |
| Copyright: © 2014 Mohamed FA, et al. This is an open-a ccess article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. | |
Related article at Pubmed Pubmed  Scholar Google Scholar Google |
|
Visit for more related articles at Journal of Bioremediation & Biodegradation
Abstract
The purpose of this paper is to irradiate the dyes by biologically through utilization involve the use of microorganisms such as Aspergillius nigar 1, 2, 3. We have previously synthesised the dyes used, namely, bisazo bifunctional bismonochlorotriazine reactive dyes and reactive disperse dye. The study confirms the potentially of the differentiated fungi in decolourization of dyes and thus opens up a scope for future analysis pertaining to their performance in treatment of textile effluent. Fungi decolourization of some synthesized reactive and reactive disperse dyes using three fungal isolates (Aspergillius nigar 1, 2, 3) were studied. These isolates were used for decolourization activities of reactive and reactive disperse dyes, after it was screened for optimum efficiency along with optimization of the condition of temperature and pH. The effect of initial pH was also tested and it was found that, highest decolorization at acidic pH 5. The results obtained demonstrate that the removal of dyes is dependent on the initial dye concentration of the solution. The isolate of Asprigillus niger 3 recorded the highest biomass accumulation after 10 days and it proves to be the most efficient one in removal the three dyes under investigation.
| Keywords |
| Fungi; Decolourization; Aspergillius nigar; Waste water; Reactive dyes; Reactive disperse dye |
| Introduction |
| Biological methods of removal of dyes involve the use of microorganisms such as fungi, bacteria, algae and actinomycetes to convert the pollutants into non-toxic harmless substances. Biological processes convert organic compounds to water and carbon dioxide. |
| Textile, cosmetics, pharmaceuticals and dyeing industry effluents constitute a major source of water pollution. Dyes or their breakdown products are known to be highly toxic and carcinogenic for living organisms. The wastewater generated from textile industries vary in their characteristics being dependent depends on the process employed such as desiring, scouring, bleaching, mercerizing, dyeing, printing and finishing. |
| Many dyes and pigments are hazardous and toxic for human as well as for aquatic life in the concentration at which they are being discharged to receiving water bodies. Dyes used in the textile industry are difficult to remove by conventional methods that are recalcitrant against light, oxidizing agents and biodegradation processes. The slow rate of decomposition of dyes present in wastewater desperately needed novel treatment methods to accelerate the process [1]. |
| Microbial metabolism and degradation of dyes depend upon the climatic condition, presence or absence of oxygen, presence of alternate carbon or energy source, optimum pH and temperature etc. |
| The textile industry is one of the industries that generate a high volume of waste water and creates potential for water pollution. Among the many chemicals in textile waste water, dyes are considered important pollutants [2]. |
| The removal of dyes from textile effluents is one of the most significant environmental problems [3,4]. Dyes are used in large quantities in many industries including textile, leather, paper, printing, plastic, food, etc. to colour their products [5]. |
| The extensive use of dyes often passes pollution problems. The presence of very low concentrations of dyes in large water bodies is highly visible and indisputable and also reduces light penetration and photosynthesis. In addition some dyes are either toxic or mutagenic and carcinogenic [6-9]. Wastewater of dyes from textile and dyestuff industries is difficult to treat. This is because dyes usually have a synthetic origin and complex aromatic structures which make them more stable and more difficult to biodegrade [10]. Azo dyes are the most widely used as they account for over 60% of the total number of dye structures known to be manufactured [11]. |
| Recently, a number of studies have focused on microbial biodegradation of dye waste water. In this respect, Fu and Virarghavan [12] reported that Aspergillius nigar was capable of removing dyes from an aqueous solution and bio absorption of dyes was influenced by the functional groups in the fungal biomass and chemical structure of the dyes. Donmeze [13], studied bioaccumulation of the reactive textile dyes Ramazol Blue, Reactive Black and Reactive Red by the yeast species Candida tropicolis growing in molasses medium and found that the increase in dye concentration inhibited growth of yeast and caused a long lag period [14]. |
| Interest in the pollution potential of textile dyes depend on their possibility of toxicity or carcinogenity, cleavage of azo dyes into the corresponding amines many of which are carcinogenic [15-17]. Azo reductases have been shown to be very specific enzymes thus cleaving only the azo bonds of azo dyes. In contrast the phenoxidases, lignin peroxidase, manganese peroxidase and laccase act more unspecifically on the aromatic ring and have the potential to degrade a wide range of aromatic structures [18]. |
| A great number of white rot fungi have been reported to produce the lignin-degrading enzymes LiP, MnP, and laccase, or at least one of these enzymes [19]. |
| In this study, two reactive dyes and reactive disperse dye that were synthesised previously and used for dyeing textile fabrics tested for degradation by micro organisms to remove it from textile waste water and exhausted dye baths were treated by three species of microbial isolate of Aspergillius nigar from Egyptian soil. The efficiency of these fungal isolates in decolourization of the said dyes was investigated. |
| Experimental |
| Materials |
| Chemicals: H-acid, γ-acid and N, N-dimethylformamide (DMF) were obtained from Fluka Chemie AG. Cyanuric chloride (98%) was obtained from Merck-Schuchardt. 2,4-Diaminobenzene sulphonic acid was obtained from DyStar. All other chemicals used in the study were of reagent grade and applied without further purification. Fungal isolate belonging to (Aspergillius nigar) was obtained from plant pathology department, National Research Centre, Egypt. |
| Synthesis of dyes |
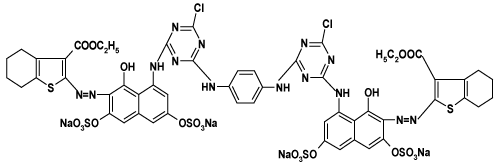
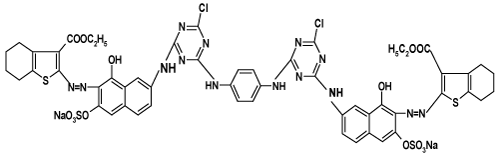
| Synthesize of bisazo bifunctional bismonochlorotriazine reactive dyes 1, 2: We previously prepared these dyes by adding a solution of cyanuric chloride (1.88 g, 0.01mol) which was stirred in acetone (50 ml) containing a few amount of crushed ice at 0-5°C in an ice/salt bath to this a solution of each of the monoazo dye in water (30 mL) was added drop wise after being adjusted at pH 6-7 by sodium carbonate solution. The reaction mixture was stirred at 0-5oC for 4h then a neutral solution of 1,4-phenylendiamine (0.58 g, 0.005 mol) in ethanol (40 mL) was added drop wise to the whole solution. The respective bisazo bifunctional bismonochlorotriazine reactive dyes (bis MCT) 1and 2 were precipitated by adding sodium chloride solution (10% w/v), The dyes were filtered off and dried at 40°C under vacuum [20]. |
| C54H42N14O18S6Na4Cl2(1530.25), Calcd: C, 42.38; H, 2.77; N, 12.81; S, 12.57%, found: C, 42.36; H, 2.81; N, 12.82; S, 12.12%. IR (ν, cm-1): 3520 (2 OH) 3468-3338 (4NH), 3055 (CH aromatic), 1690, 1640 (2 bis C=O), 1500 (2 bis N=N). 1HNMR (δ, ppm): 1.16 (t, 6H, 2 x ester CH3), 3.44-4.22 (m, 16H, 2 × 4CH2), 4.72 (q, 4H, 2 × ester CH2), 7.85-7.97(s, 3H, 2 × naphthyl H), 10.11(s, 2H, 2 x NH), 10.31 (s, 2H, 2 × NH),13.21 (s, 2H, 2x ×OH). |
| C54H44N14O12S4Na2Cl2 (1326.16), Calcd: C, 48.91; H, 3.34; N, 14.79; S, 9.67%, found: C, 48.66; H, 3.55; N, 14.82; S, 9.92%. IR (ν, cm-1): 3468- 3338 (OH, 4NH), 3055 (CH aromatic), 2895, 2865 (CH3, CH2), 2228, 2220 (2CN), 1690 (C=O), 1520 (N=N). 1HNMR (δ, ppm): 1.18 (t, 6H, 2 × ester CH3), 2.46-2.85 (m, 16H, 2 × 4CH2), 4.34 (q, 4H, 2 × ester CH2), 7.12 ( s, 2H, 2 × naphthyl H), 7.54 ( t, 2H, 2 × naphthyl H), 7.91-8.13 (d, 4H, 2 × naphthyl H), 10.11 (s, 2H, 2 × NH), 10.41 (s, 2H, 2 × NH), 13.51 (s, 2H, 2 × OH). |
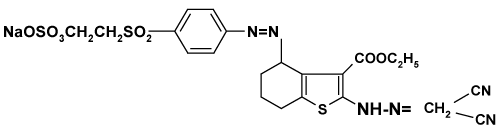
| Synthesise of reactive disperse of Ethyl 2-hydrazomalononitrilo- 4-(sodium 2-(4-aminophenylsulfonyl) ethylsufato-4,5,6,7- tetrahydrobenzo[b]thiophene: The method was carried out by initial preparation of the 1-aminobenzene-4-β-sulphatoethylsulphone PABSES (2.86 g; 0.01 mol) in water (50 ml) then treated with 10 ml HCl/0.69 g NaNO2 to produce the diazonium salt SES. The resulting solution of the diazonium salt was then added slowly to a stirred solution of the dye intermediates a (3.02 g; 0.01 mol). The desired reactive disperse dye 3 was precipitated by slow addition of sodium chloride (15% w/v), filtered and dried under vacuum at 40°C [21] (Figure 3). |
| C22H21N6NaO8S3 (616.62), Calcd: C,42.85; H,3.43; N,13.63; S, 15.60%, found: C, 42.66; H, 4.81; N, 13.82; S, 15.92%. IR (ν, cm-1): 3468- 3338 (NH), 3055 (CH aromatic), 2895, 2865 (CH3, CH2), 2228, 2220 (2CN), 1690 (C=O)), 1638 (C=C). 1HNMR (δ, ppm): 1.16 (t, 3H, ester CH3), 2.23-2.74 (2m, 6H, 3CH2), 3.44 (t, 1H, cyclohexene CH), 4.22 (q, 2H, ester CH2), 3.88, 4.72 (2t, 4H, 2CH2), 8.11 (s, 1H, NH). |
| Preparation of dye solution |
| The dye stock solution was prepared by dissolving accurate weight of the dye in distilled water to the concentration of 500 mg/L. Different concentrations were prepared from the stock solution (0.02-0.1 g/L). |
| Fungal isolates and dye degradation: Aspergillius nigar isolates were isolated and identified at Plant Pathology department, National Research Centre, Cairo, Egypt. These isolates were tested for their high dye decolourization potential in earlier report [22]. |
| Dye Decolourization |
| The dyes were added to cultures as aliquots of concentrated stock solutions. Decolourization was measured spectrophotometrically at the wavelength of peak absorbance of each dye using UV/ Vis recording spectrophotometer. The experiments were carried out in 250 ml flasks containing 100 ml of SDA medium, at different concentrations of tested dyes (0.02-1g/L). The pH was adjusted (3-7), and then the flasks were autoclaved at atmospheric pressure 1.5 for 30 min. The autoclaved flasks were incubated with 5mm discs of 7 day old fungi of cultures of the tested isolates. They incubated for two weeks, after the end of this period 20 ml of the dye solution was centrifuged at 5000 rpm for 15 min. Then the maximum absorption (λ max) was measured using spectrophotometer. |
| Decolourization Assay: The decolourization assay was expressed in the terms of decolourization% using spectrophotometer. The decolourization percentage in dye concentration was calculated for each treatment (dye concentration and fungal isolate) as follows: |
| Decolourization%= 100 Χ(Co - Ct )/Co |
| Where, Co is the initial concentration of the dye (control), Ct is the concentration at time t. |
| Results and Discussion |
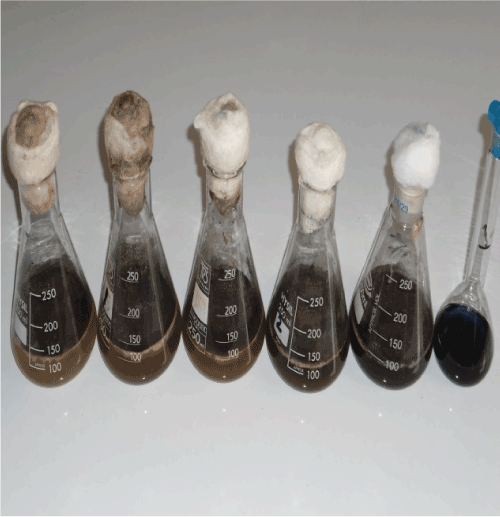
| Dye concentration Fungi decolourization of three synthesized dyes was studied. The colour removal of the reactive dyes can be achieved by treating with Asperigillus nigar to cause biodegradation of the colour of dyes under investigation [23]. The non incubated flasks of each dye concentration were set as control (Figures 1-3). Differences in efficiency of decolourization of the three dyes under investigation due to the use of vary fungi Asperigillus nigar 1, 2, 3 can be realized by making a comparison among the photographs represented in Figures 1-3. |
| The comparison reveals that Asperigillus nigar 3 reduces the highest biomass accumulation after 10 days incubation and, in deeded, it is the most efficient in removal of each of the three dyes used. It is also observed that dye 2 displays the highest percent decolourization as compared to dye 1 and dye 3 for the three isolates used. |
| The comparison outputs as obtained above could be interpreted in terms of nature of the dye (Figure 4-6) as well as nature of the fungi. The nature of the subsistent on the aromatic ring has been shown to influence enzymatic oxidation. Electron donating subsistent as methyl, methoxy and amino groups enhance enzymatic degradation of azo phenols while electron withdrawing subsistent as chloro, nitro and hydroxyl inhibited oxidation [24]. Hydroxy and amino groups enhance decolourization. |
| The azo dye reduction may involve different mechanisms or locations like enzymatic, non-enzymatic, me-diated, intracellular and various combinations of these mechanisms and locations. Oxidative biodegradation takes place upon action of enzymes such as peroxidases and laccases. The involvement of fungal peroxidases and laccases for the oxidation of sulfonated azo dyes has been reported earlier [25]. |
| The decolonization efficiency of fungi can be due to the presence of chitin with hydroxyl and amino groups in their cell wall, which make them an efficient adsorbent of dye effluent. Differences in the capacity of dye decolonization between fungi have been related to inter and intraspecific variations, the molecular complexity of the dye and culture conditions [10,26]. |
| The results exhibit a general trend which demonstrates that the decolourization percent increases as the dye concentration increases respective of the dye used. Nevertheless, the two reactive dyes are more susceptible to biological decolourization than the reactive disperse dye and within the category of reactive dyes, dye 2(N,�?�?-Bis[3-ethyl2- azo(2-amino-8-hydroxy-nathalene-7-yl-6-sulphonicacid)-4,5,6,7- tetrahydrobenzo[b]thiophene-3-carboxylato-(N-4-chlorotriazino- 6-yl)]-1,4-phenylenediamine) is more amenable to decolourization than dye 1(N, �?�?-Bis[3-ethyl 2-azo(1-amino-8-hydroxy-nathalene- 7-yl-3,5-disulphonic acid)-4,5,6,7-tetrahydrobenzo[b]thiophene-3- carboxylato-(N-4-chlorotriazino-6-yl)]-1,4-phenylenediamine). The three dyes show different efficiency of decolourization which brings into focus the following order dye 2> dye 1>dye 3. |
| The isolate of Asprigillus niger 3 reduced the highest biomass accumulation after 10 days and it is the most efficient one in removal the reactive dye for the three dyes used. It is also noticed that dye 2 recorded the highest percent as compared to dye 1 and dye 3 for the three isolates used. |
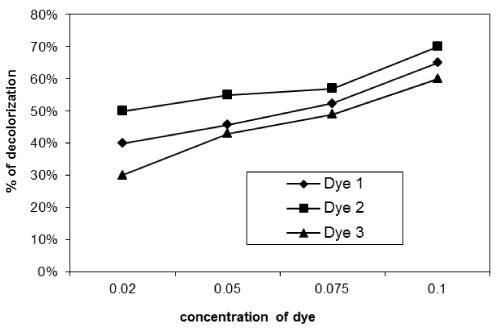
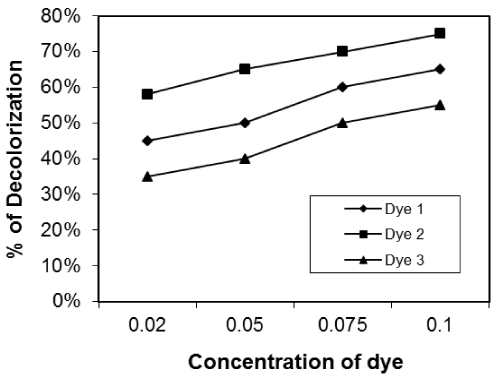
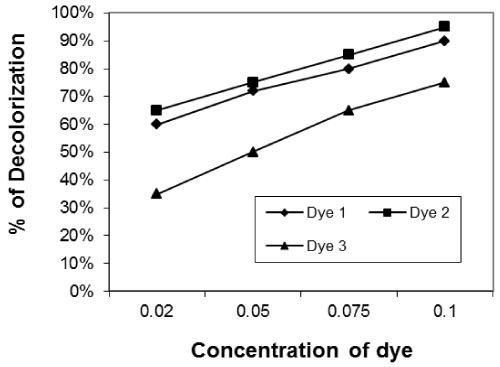
| Effect of initial dye concentration on decolourization% dye: The effect of initial dye concentration of dye in solution on removal of dyes was studied. We used different concentration of dyes (0.02, 0.04, 0.08 and 0.1g/L). The removal of dyes was clearly dependent on the initial dye concentration of the solution (Figures 7-9) for the three isolates used. Figures 7-9 show the effect of dye concentration on decolourization percent for ten days incubation of the reactive dye solution using Asperigillus nigar 1, 2, 3 respectively. |
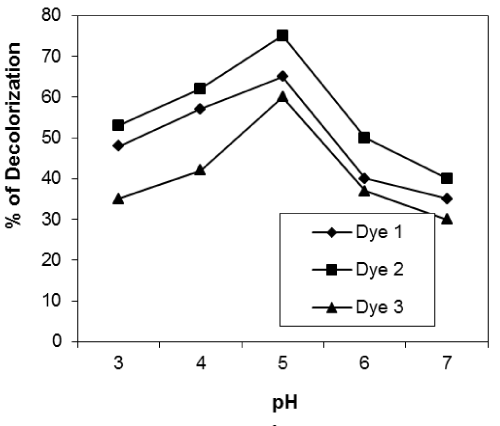
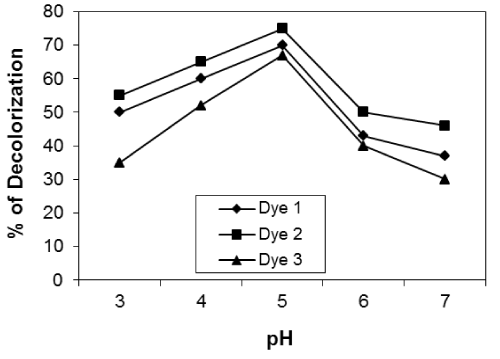
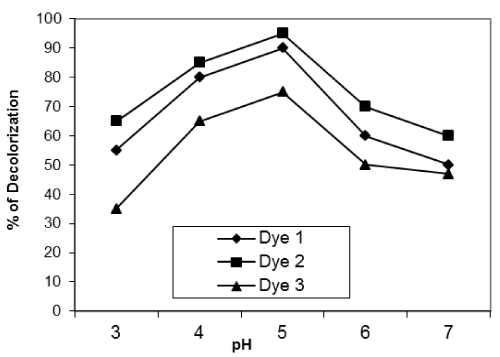
| Effect of pH of the dye solution: Figures 10-12 show the effect of pH of the dye solution on the decolourization% of microorganisms within the range of pH 3-7. The results show that the decolourization reaches maximum at pH 5 for reactive dyes for all isolates used. Many authors stated that pH is very important for fungal growth. |
| Fungi can grow at low pH from 4 to 5. Higher uptake obtained at lower pH value may be due to the electrostatic attraction between positively charged cell surface and dye anions.The optimum pH for decolourization by unidentified White-rot fungus was pH 4-5 [24]. |
| Conclusion |
| Fungi decolourization of reactive dyes could be successfully achieved by treating with Aspergillius nigar. The latter is capable of removing approximately 95% dye colour after 10 days incubation. The most efficient removal was observed with the reactive dye 3 at high concentrations. The results revealed also that the decolourization attains maximum at pH 5 with respect to the three two reactive dyes and the reactive disperse dye used. |
| The removal of dyes was clearly dependent on the initial dye concentration of the solution. The isolate of Asprigillus niger 3 displays the highest biomass accumulation after 10 days and it is the most efficient in removal of the reactive colour of the three dyes used. It was also noticed that dye 2 exhibited the highest percent as compared to dye 1 and dye 3 for the three species used. |
References
|
Figures at a glance
 |
 |
 |
 |
| Figure 1 | Figure 2 | Figure 3 | Figure 4 |
 |
 |
 |
 |
| Figure 5 | Figure 6 | Figure 7 | Figure 8 |
 |
 |
 |
 |
| Figure 9 | Figure 10 | Figure 11 | Figure 12 |
Relevant Topics
- Anaerobic Biodegradation
- Biodegradable Balloons
- Biodegradable Confetti
- Biodegradable Diapers
- Biodegradable Plastics
- Biodegradable Sunscreen
- Biodegradation
- Bioremediation Bacteria
- Bioremediation Oil Spills
- Bioremediation Plants
- Bioremediation Products
- Ex Situ Bioremediation
- Heavy Metal Bioremediation
- In Situ Bioremediation
- Mycoremediation
- Non Biodegradable
- Phytoremediation
- Sewage Water Treatment
- Soil Bioremediation
- Types of Upwelling
- Waste Degredation
- Xenobiotics
Recommended Journals
Article Tools
Article Usage
- Total views: 14940
- [From(publication date):
February-2014 - Aug 24, 2025] - Breakdown by view type
- HTML page views : 10306
- PDF downloads : 4634
