Research Article Open Access
Biosensors for Indirect Monitoring of Foodborne Bacteria
Diana C Vanegas1, Carmen Gomes2* and Eric S McLamore3
1Food Engineering, Universidad del Valle, Colombia
2Biological and Agricultural Engineering, Texas A&M University, USA
3Agricultural and Biological Engineering, University of Florida, USA
- *Corresponding Author:
- Carmen Gomes
Biological and Agricultural Engineering
Texas A&M University, USA
Tel: 979845-2455
E-mail: carmen@tamu.edu
Received date: March 05, 2016; Accepted date: May 04, 2016; Published date: May 05, 2016
Citation: Vanegas DC, Gomes C, McLamore ES (2016) Biosensors for Indirect Monitoring of Foodborne Bacteria. Biosens J 5: 137. doi:10.4172/2090- 4967.1000137
Copyright: © 2016 Vanegas DC, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Biosensors Journal
Abstract
Microbiological safety assessment in food samples is often a cumbersome and time consuming task that has to be done in a regular basis in most food manufacturing facilities. Standard microbiological techniques for evaluating critical levels of bacteria take several days to yield results, thus the food products have to be withheld in the processing factories until the safety inspection is concluded. The food industry requires faster and reliable tools for more efficient microbiological testing that can reduce the safety inspection lag significantly. Biosensors that target specific molecules associated with bacterial activity are amongst the most promising technologies for addressing this need. State-ofthe- art biosensors allow for instantaneous quantification of molecular biomarkers. This review focuses on describing the operation mechanism and technological challenges and opportunities for developing high performance biosensors capable of real-time monitoring of biomarkers from foodborne bacteria.
Keywords
Foodborne pathogens; biomarkers; nanomaterials
Introduction
The rich nutrient content and high water activity of foods (particularly in fresh and minimally processed products) offer a propitious environment where bacteria can live and thrive. There are at least 31 known pathogens that cause foodborne illnesses. Nontyphoidal Salmonella, Escherichia coli, Listeria monocytogenes, and Campylobacter spp. remain the most prevalent strains in the USA that cause food poisoning resulting in hospitalizations and death [1]. In order to reduce the prevalence of foodborne infections, regulatory agencies have been strengthening the safety standards by requiring more exhaustive protocols of microbiological inspection in the food processing plants [2]. Direct assessment of bacterial contamination is typically performed based on microbiological counts. Usual microbiological methods for detecting bacteria are laborious, requiring sample preparation/dilution, selective enrichments and subsequent culturing on a selective medium to obtain colonies; and usually take several days to produce results. These microbiological tests are often followed by biochemical and/or serological confirmation and differentiation of species [3]. There are a number of modern detection technologies that enable significantly faster analysis (in the order of minutes to hours) [4]; some of these technologies are designed for directly assessing bacteria via the detection of cell components such as membrane proteins or DNA (e.g. PCR, aptasensors, and phagesensors), whereas other devices aim to detect bacteria indirectly by measuring molecular markers produced by specific strains (e.g. chromatographic methods, and enzymatic biosensors). While direct monitoring of pathogens is the preferred approach, biomarker monitoring is a more rapid and less expensive technique that can provide early screening of large volumes of food samples. Herein, we provide a concise overview of state-of-the-art biosensors for indirect monitoring of foodborne bacteria, and describe the operating mechanism and current limitations of this fast-evolving technology.
Operating principle of biosensors
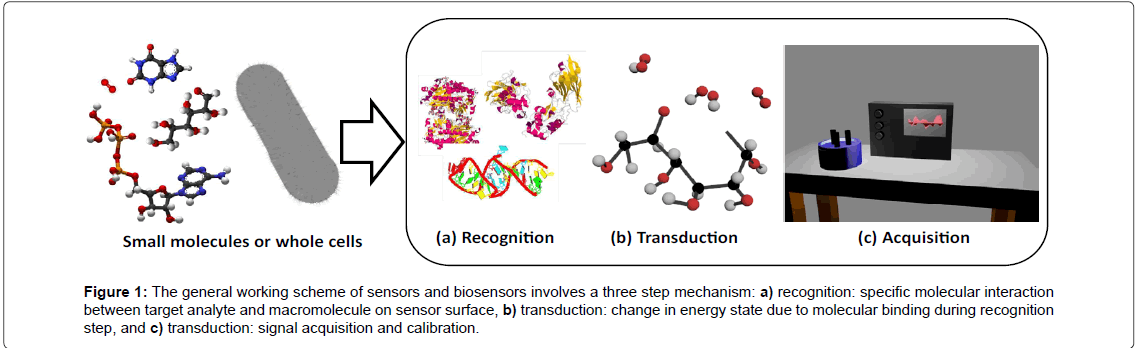
Recent technological advancements in the field of food safety biosensors have focused on real-time monitoring-technologies such as aptamer-based assays, fluorescent probes, and electrochemical sensors (Figure 1). The working scheme for all sensors and biosensors involves a three step process, including i) recognition, ii) transduction, and iii) signal acquisition. In the recognition step (panel a), a molecular interaction between the target (i.e., analyte) and a macromolecular structure on the sensor surface results in highly specific binding. In the transduction step (panel b), selective binding of the target produces a change in mass, photon transport, and/or electron transport. In the acquisition step (panel c), the change in energy state as a result of specific molecular interactions is measured and an output is correlated to analyte concentration.
Figure 1: The general working scheme of sensors and biosensors involves a three step mechanism: a) recognition: specific molecular interaction between target analyte and macromolecule on sensor surface, b) transduction: change in energy state due to molecular binding during recognition step, and c) transduction: signal acquisition and calibration.
Enzymatic biosensors for measuring bacterial biomarkers
Enzymatic biosensors are capable of real-time monitoring and ultrasensitive detection. The working mechanism of this type of device is based on the direct measurements of biochemical reactions at the sensor-sample interphase. Amperometric biosensors are based on the catalytic activity of immobilized enzymes that either produce hydrogen peroxide during the course of an oxidative reaction (Equation 1), or generate the reduced form of β-nicotinamide adenine dinucleotide (NADH) when the target analyte undergoes reduction at the sensor surface (Equation 2).
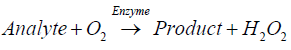 (1)
(1)
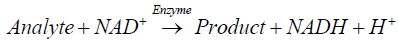 (2)
(2)
Based on the principle described above, a number of portable electrochemical biosensors have been developed for detecting bacterial biomarkers like xanthine (a biomarker of meat spoilage) [5], D-aminoacids (biomarkers of racemization in protein-rich foods) [6], histamine and putrecine (biomarkers of shellfish decomposition) [7], and botulinum neurotoxin (a biomarker of Clostridium botulinum contamination in canned foods) [8] among others. Table 1 depicts some other recent examples of biosensors capable of tracking bacterial biomarkers for food quality and safety analyses.
| Biosensor Platform | Analyte/Sample | Response Time | Operating Range | Reference |
|---|---|---|---|---|
| XO-CD/pAuNP/SWNT/GCE | Xanthine/NT | 5 s | 0.05-9.5 µM | [9] |
| CGi/SWNT/p-DNA | t-DNA/NT | NR | 100 fM-1 μM | [10] |
| DAAO/c-MWCNT/CuNPs/PANI/Au | D-amino acid/fruit juices | 2 s | 0.001-0.7 mM | [11] |
| MIP/Gr-AuNPs/GCE | Glycoprotein/NT | 8 min | 1×10-11 -1×10-5 g/mL | [12] |
| GS-CS-PtPd/ChOx | Cholesterol/meat, fish oil, margarine and eggs | <7 s | 2.2 - 520 µM | [13] |
| NT: not tested in food samples. NR: not reported in the manuscript. | ||||
Table 1: Biosensors capable of monitoring markers of bacterial contamination in food.
Villalonga et al. demonstrated a biosensor enhanced with single walled carbon nanotubes (SWNT) and gold nanoparticles (pAuNP) for the direct determination of xanthine, which is a commonly found marker of bacterial spoilage in meat. The operating mechanism of this electrochemical device is based on an enzymatic reaction catalyzed by xanthine oxidase (XO), immobilized on the surface of a pAuNP/SWNT glassy carbon electrode (GCE) [9]. Ko et al. presented a biosensor consisting of a chip functionalized with gold nano-islands (CGi), single-walled carbon nanotubes (SWNT) and a 25 mer p-DNA for selective detection of t-DNA. The sensing principle is based on changes in the overall conductance occurring after hybridization of t-DNA with p-DNA on the chip surface [10]. Lata et al. developed a D-amino acid biosensor based on covalent immobilization of D-amino acid oxidase (DAAO) onto a gold electrode (Au) modified with multi-walled carbon nanotubes (c-MWCNT), copper nanoparticles (CuNPs) and polyalinine (PANI). In this study, the authors only aimed to quantify the level of amino acids in fruit juices as a measure of their nutritional value; nonetheless, the device could also be used to estimate presence and maybe even viability of bacteria based on the relative amounts of D and L amino acids and the rate of racemization [11]. Wang et al. created a biomimetic sensor based on graphene, gold nanoparticles (Gr-AuNPs) and molecularly imprinted polymers (MIP) onto a glassy carbon electrode (GCE) for detecting glycoproteins. In this case, a BSA template was used as model protein for technology demonstration. This amperometric device could be applied for detecting cell membrane glycoproteins which are known to be important virulence factors in pathogenic microorganisms such as E. coli and Streptococcus spp. [12]. Cao et al. built an electrochemical biosensor functionalized with graphene, platinum, palladium, chitosan (GS-CS-PtPd) and cholesterol oxidase (ChOx). The probe was designed for cholesterol sensing in food samples, and has a potential use for monitoring the transformation of sterols by specific spoilage microorganisms [13].
Challenges and opportunities
Enzymatic biosensors for monitoring biomarkers are highly accurate, but adoption of these technologies has been slow. In large part, this is due to a lack of demonstration under field conditions, or poor shelf life (primarily due to enzyme denaturation). Maintaining enzyme activity in thin films is a major challenge in the field, particularly when sensors are used in complex solutions during field applications. Problems inherent to enzyme immobilization include loss of purity, stability, activity, and/or specificity [14]. Bio-based nanomaterials provide durability and stability in the thin protein film formed on the electrode surface [15].
New biomaterials and nanomaterials that can enhance the performance of biosensors in terms of sensitivity, response time, shelf life, and biocompatibility are continuously under development. These improvements provide new opportunities for creating ultra-sensitive, smart sensors that are capable of real-time detection of foodborne pathogens.
Nanomaterials that enhance transduction have recently shown to significantly improve performance of biosensors. Most research has focused on materials that improve electron or photon mobility (e.g. carbon nanotubes, graphene sheets, nanometals, metal oxides and other photocatalysts) [16-18]. A current trend in electrochemical sensing is to create transducer-composites based on carbon nanomaterials and noble nanometals. This approach takes advantage of the synergistic effects on electrical properties that result from combining these types of materials, to obtain devices with superb sensitivity, instantaneous response time, and extremely low detection limit [19].
A number of bio-based hydrogels have recently been developed that can enhance sensor longevity without creating toxic byproducts. Hydrogels have high strength/stiffness, optical transparency, biocompatibility, biodegradability, and highly porous structural network [20-22]. For example, hydrogels composed of chitosan, cellulose, or poly-N-isopropylacrylamide (PNIPAAM) can be used to encapsulate macromolecules designed to specifically bind biomarkers associated with pathogens. In addition to immobilization of recognition agents, hydrogels based on weakly ionizable polysaccharides show pHresponsive phase transition [23]. This stimulus-response behavior can be exploited in biosensors for controlled release of small molecules and scaffolding of enzymes.
Conclusions
The long time required for testing the microbiological safety and quality of food products using standard microbiological methods is a concerning problem for the food industry and public health. Biosensor technologies that target bacterial biomarkers are promising alternatives for rapid screening of harmful bacteria in food samples. Incorporation of nanomaterials with unique electrical and photonic properties, as well as biomaterials with high biocompatibility are the most effective strategies for developing biosensors with ultrafast response time and high stability.
References
- Centers for Disease Control and Prevention. (2011). CDC Estimates of Foodborne Illness in the United States. Retrieved February 25, 2016, from http://www.cdc.gov/foodborneburden/2011-foodborne-estimates.html
- U.S. Food and Drug Administration. (2015). Food: Compilance and Enforcement. Retrieved February 25, 2016, from http://www.fda.gov/Food/ComplianceEnforcement/default.htm
- Denoya C, Colgan S, du Moulin GC. Alternative Microbiological Methods in the Pharmaceutical Industry: The Need for a New Microbiology Curriculum. Am. Pharm. Rev 9: 10-18.
- Zhao X, Lin CW, Wang J, Oh DH (2014) Advances in Rapid Detection Methods for Foodborne Pathogens. J Microbiol Biotechnol 24: 297-312.
- Vanegas DC, Gomes C, McLamore ES (2014) Xanthine Oxidase Biosensor for Monitoring Meat Spoilage. Proc. SPIE 9107.
- Wcisło M1, Compagnone D, Trojanowicz M (2007) Enantioselective screen-printed amperometric biosensor for the determination of D-amino acids. Bioelectrochemistry 71: 91-98.
- Henao-Escobar W, del Torno-de Román L, Domínguez-Renedo O, Alonso-Lomillo MA, Arcos-Martínez MJ (2016) Dual enzymatic biosensor for simultaneous amperometric determination of histamine and putrescine. Food Chem 190: 818-823.
- Ferracci G, Marconi S, Mazuet C, Jover E, Blanchard MP, et al. (2011) A label-free biosensor assay for botulinum neurotoxin B in food and human serum. Anal Biochem 410: 281-288.
- Villalonga R, Díez P, Eguílaz M, Martínez P, Pingarrón JM (2012) Supramolecular immobilization of xanthine oxidase on electropolymerized matrix of functionalized hybrid gold nanoparticles/single-walled carbon nanotubes for the preparation of electrochemical biosensors. AC S Appl Mater Interfaces 4: 4312-4319.
- Ko JW, Woo JM, Jinhong A, Cheon JH, Lim JH, et al. (2011) Multi-order dynamic range DNA sensor using a gold decorated SWCNT random network. ACS Nano 5: 4365-4372.
- Lata S, Batra B, Kumar P, Pundir CS (2013) Construction of an amperometric D-amino acid biosensor based on D-amino acid oxidase/carboxylated mutliwalled carbon nanotube/copper nanoparticles/polyalinine modified gold electrode. Anal Biochem 437: 1-9.
- Wang X, Dong J, Ming H, Ai S (2013) Sensing of glycoprotein via a biomimetic sensor based on molecularly imprinted polymers and graphene-Au nanoparticles. Analyst 138: 1219-1225.
- Cao S, Zhang L, Chai Y, Yuan R (2013) Electrochemistry of cholesterol biosensor based on a novel Pt-Pd bimetallic nanoparticle decorated graphene catalyst. Talanta 167-172.
- Garcia-Galan A, Berenguer-Murcia A, Fernandez-Lafuente R, Rodrigues RC (2011)Potential Different Enzyme Immobilization Estrategies to Improve Enzyme Performance. Advanced Synthesis and Catalysis.353: 2885-2904.
- Hanefeld U, Gardossi L, Magner E (2009) Understanding enzyme immobilisation. Chem Soc Rev 38: 453-468.
- Tiwari JN, Vij V, Kemp KC, Kim KS (2016) Engineered Carbon-Nanomaterial-Based Electrochemical Sensors for Biomolecules. ACS Nano 10: 46-80.
- Kumar S, Ahlawat W, Kumar R, Dilbaghi N (2015) Graphene, carbon nanotubes, zinc oxide and gold as elite nanomaterials for fabrication of biosensors for healthcare. Biosens Bioelectron 70: 498-503.
- Hayat A, Catanante G, Marty JL (2014) Current Trends in Nanomaterial-Based Amperometric Biosensors. Sensors (Basel) 14: 23439-23461.
- Vanegas DC, Taguchi M, Chaturvedi P, Burrs S, Tan M (2014)A comparative study of carbon-platinum hybrid nanostructure architecture for amperometric biosensing. Analyst 139: 660-667.
- Klemm D, Kramer F, Moritz S, Lindström T, Ankerfors M, et al. (2011) Nanocelluloses: a new family of nature-based materials. Angew Chem Int Ed Engl 50: 5438-5466.
- Kalia S, Dufresne A, Cherian BM, Kaith BS, Averous L, et al. (2011) Cellulose-Based Bio- and Nanocomposites: A Review. international Journal of Polymer Science: 837875.
- Burrs SL, Vanegas DC, Bhargava M, Mechulan N, Hendershot, P, et al. (2014) A Comparative study of graphene-hydrogel hybrid bionanocomposites for biosensing. Analyst 140: 1466-1476.
- Gil ES, Hudson SM (2004) Stimuli-reponsive polymers and their bioconjugates. Prog. Polym Sci 29: 1173-1222.
Relevant Topics
- Amperometric Biosensors
- Biomedical Sensor
- Bioreceptors
- Biosensors Application
- Biosensors Companies and Market Analysis
- Biotransducer
- Chemical Sensors
- Colorimetric Biosensors
- DNA Biosensors
- Electrochemical Biosensors
- Glucose Biosensors
- Graphene Biosensors
- Imaging Sensors
- Microbial Biosensors
- Nucleic Acid Interactions
- Optical Biosensor
- Piezo Electric Sensor
- Potentiometric Biosensors
- Surface Attachment of the Biological Elements
- Surface Plasmon Resonance
- Transducers
Recommended Journals
Article Tools
Article Usage
- Total views: 12796
- [From(publication date):
June-2016 - Aug 28, 2025] - Breakdown by view type
- HTML page views : 11452
- PDF downloads : 1344