Research Article Open Access
Deproteinization of Distillery Yeast Biomass Waste by Protease-producingBacillus megaterium PB 4
| Rajasundari Kandaiah* and Murugesan Ramasamy | |
| Department of Agricultural Microbiology, TamilNadu Agricultural University, Coimbatore-641 003, Tamilnadu, India | |
| Corresponding Author : | Rajasundari Kandaiah Department of Agricultural Microbiology Tamil Nadu Agricultural University Coimbatore-641 003 Tamilnadu, India Tel: 0422 661 1200 E-mail: rajasundarikandaiah@ gmail.com |
| Received: September 28, 2015; Accepted: October 28, 2015; Published: October 30, 2015 | |
| Citation: Kandaiah R, Ramasamy M (2015) Deproteinization of Distillery Yeast Biomass Waste by Protease-producing Bacillus megaterium PB 4. J Bioremed Biodeg 6:319. doi:10.4172/2155-6199.1000319 | |
| Copyright: © 2015 Kandaiah R, et al. This is an open-a ccess article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. | |
| Related article at Pubmed, Scholar Google | |
Visit for more related articles at Journal of Bioremediation & Biodegradation
Abstract
The distillery yeast biomass (DYB), a waste by-product from distilleries is one of the under-utilized protein sources to be exploited for the production of protein hydrolysates. In this study, a neutral protease produced from Bacillus megaterium PB4 isolated from agro wastes enriched soil exhibited significant deproteinization of distillery yeast biomass to yield protein hydrolysates. Among three protease producing strains isolated, the maximum protease production of 120.3 ± 1.4 U mL-1 was exhibited by PB4 grown in Luria-Bertani (LB) broth supplemented with casein at pH 7.0 and 30°C. The minimal media supplemented with various carbon and nitrogen sources, glucose and peptone significantly improved the protease production in PB4. The effect of metal ions such as Zn2+, Mg2+, Mn2+, Cu2+ at 0.5 and 1.0 mM final concentration in the media on protease production indicated that Zn2+ (1.0 mM) was favorable in enhancing the protease production. The Km and Vmax of PB4 protease for casein substrate was 0.6 U ml-1 and 217.3 μM min-1 mg-1 protein, respectively. The deproteinization rate for distillery yeast biomass was 84 and 76.4% using culture and crude enzyme extracted from PB4, respectively. To the best of our knowledge, deproteinization of distillery yeast biomass using proteolytic bacterial isolate has never been demonstrated before. The experimental results in the present study suggest that Bacillus megaterium PB4, can be utilized for the eco-friendly, economical production of protein hydrolysates and may be considered as a potential candidate in various biotechnological applications.
| Keywords |
| Bacillus megaterium; Proteases; Distillery yeast biomass; Deproteinization |
| Introduction |
| Global demand for human and animal feed is ever increasing and will continue in future. Protein-rich food is considered as essential for the growth and development of animals including humans. Generally in plants, protein/amino acid deficiency results in decreased resistance against plant pathogens and pests which in turn affects the growth and yield [1]. On the other hand in animals, protein deficiency causes retarded growth, decreased appetite, weight loss, poor growth, depressed reproductive performance, and reduced milk production in farm animals and egg production in broilers [2]. Bridging the gap in protein-deficiency in plant and animal nutrition, protein rich wastes and by-products from the agro industries on further recycling and reprocessing results in the production of higher value products that can also be utilized as feed supplements [3,4]. Even without pre-treatments, a wide variety of protein rich by-products from agro industries are currently being supplemented as feed in the animal nutrition but in negligible quantity [5]. However, the direct ingestion of bulk protein rich products resulted in negative effect on the growth and development animals and human beings. Occasionally the consumption of protein rich material requires prolonged digestion that results in indigestion and aversion. Consequently the need for finding alternative protein rich feed sources has gained its momentum. |
| Protein hydrolysates, the amino acid rich component extracted from protein rich substrates can be very effective and efficient in facilitating the absorption of amino acids in the blood system ultimately improving the overall physiological performances of animals. The protein hydrolysates are previously reported to be used as feed supplement in poultry [6], piggery [7], and aquaculture [8] and also in direct application to plants [9]. Protein hydrolysates are derived by the proteolytic cleavage or deproteinization of protein rich substances. Deproteinization can be achieved by means of both chemical and biological methods. The chemical method of deproteinization is considered as an efficient way for separating the amino acids from protein-rich wastes. However this method of deproteinization results in the discharge of effluents which are highly acidic and alkaline [10]. Hence, biological deproteinization is considered as the efficient way of separating protein with the aid of proteases are widely used. Proteases are ubiquitous enzymes found in plants, animals and microorganisms. Plant-derived proteases such as papain, bromelain and keratinases and animal-derived protein such trypsin, chymotrypsin, pepsin and rennins are well-known for their applications [11,12]. Applications involving protease enzymes derived from microorganisms are preferred over plant and animal derived proteases since microbial proteases possesses desirable characteristics for biotechnological application [13,14]. Microbes effectively utilize protein-rich substrates as the source of carbon and energy which indirectly facilitates the production of proteases which in turn cleaves protein into amino acids [15]. Moreover genetic manipulation of protease encoding gene(s) in microorganisms also improves the catalytic properties of proteases suitable for much wider applications [16]. |
| Proteases are the class of enzymes that catalyze hydrolytic reactions in which protein molecules are degraded to smaller peptides and amino acids [17]. Proteases are used extensively in the pharmaceutical, leather, food and detergent industries. According to their activity in various pH ranges, proteases are classified as acidic, neutral and alkaline. Acidic proteases are produced by Mucor sp. Pencillium sp. and Aspergillus sp. Neutral proteases are produced by B. cereus, B. megaterium, B. stearothermophilus, B. thuringiensis, B. pumilus, B. polymyxa, B. licheniformis and B. amyloliquefaciens and also by Pseudomonas aeruginosa and Streptomyces griseus [15] and on the other hand, alkaline proteases are reported in Bacillus spp. Pseudomonas, Xanthomonas maltophila, Vibrio alginolyticus etc., [18]. Over past few decades, the isolation and application of protease producing microorganism and the enzymes were reported. |
| There is still paucity in research including the isolation of protease producing microorganisms from protein-rich waste material dumped soil. Hence, the objective of the present study was aimed in (I) isolation, characterization and screening of protease producing bacteria from various sources such as from dump sites of poultry waste, meat waste, fish waste, and distillery yeast biomass dump yard, (II) identification of efficient protease-producing bacterial strain by 16s rRNA gene sequencing, (III) proteolytic kinetics by crude enzyme isolated from B. megaterium PB4, and (IV) deproteinization of distillery yeast biomass by protease producing B. megaterium PB4. |
| Materials and Methods |
| Materials |
| Protein rich agro wastes namely distillery yeast biomass used in the experiment was purchased from, M/S Sakthi Sugars, Appakudal, Bhavani Taluk, Erode District, Tamilnadu, India and Amaravathi Co-Operative Sugar Mills Ltd, Krishnapuram, Udumalpet Taluk, Coimbatore District, Tamilnadu, India. All the reagents and chemicals used in this study were of analytical grade, obtained from Sigma Aldrich. |
| Isolation of proteolytic microorganisms |
| Soil samples collected from various agro waste dumpsites were used for the isolation of protease producing bacteria. Protease producing bacteria was isolated by enrichment technique in Luria- Bertani (LB) broth supplemented with 1.0% (w/v) casein [19] by spread plate method. The plates were kept for incubation under aerobic condition in an incubation chamber at 30 ± 2°C for 2 days to determine the proteolytic activity. Based on the highest protease production, the bacterial isolate Bacillus megaterium PB4 was selected among three isolates and used for this study. The organisms were stored at –20°C in LB broth containing 20% (v/v) glycerol. The bacteria were subcultured twice in fresh LB medium before it was used. |
| Screening and characterization |
| For the morphological classification, bacteria were cultured on LB agar plates. Morphological characterization was carried out using light microscope [20]. The proteolytic bacterial cells were fixed and viewed under the scanning electron microscope (Philips 501) as per the procedure stated by Wagner and Stewart [21]. The isolate was subjected to various tests as per standard protocol for biochemical testing described elsewhere [22]. The ability to produce protease activities we used minimal medium agar plates containing skim milk 1.0%. Plates were incubated at 30 ± 2 °C for 2 days, and clearing halos were determined. |
| PCR amplification of the 16s rRNA gene and sequence analysis |
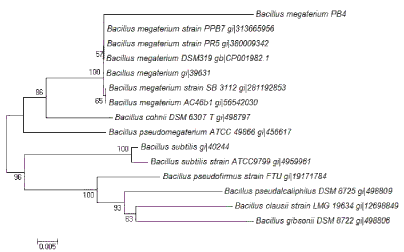
| The genomic DNA of PB4 which exhibited higher K release from the insoluble minerals was isolated using the standard protocol of hexadecyl-trimethyl ammonium bromide (CTAB) method given by Melody [23] with slight modifications. Full-length 16S rDNA (1500 bp) were amplified from the PB4 isolates by PCR using the bacterial universal primers 27F (5′-AGA GTT TGA TCC TGG CTC AG-3′) and 1492R (5′- GGT TAC CTT GTT ACG ACT T -3′) [24]. Sequencing reactions were performed using ABI prism terminator cycle sequencing ready reaction kit and electrophoresis of the products were carried out on an Applied Biosystems (Model 3100) automated sequencer. The cultures were identified at species level by performing a similarity search against the GenBank database (website: http://www.ncbi.nih.gov/BLAST). The phylogenetic tree was constructed with sequences of 16S rDNA from different eubacteria which was obtained from NCBI. Sequences were aligned using the CLUSTAL X software [25]. The evolutionary distances were calculated by neighbour-joining method [26] using MEGA 6.0 software [27]. |
| Effects of pH, temperature, carbon, nitrogen and metal ions on protease production |
| Protease activity from B. megaterium PB4 in M9 minimal salt medium at various pH (4.0-9.0), temperature (20-60°C), carbon sources (glucose, fructose, lactose, maltose, and cellulose), nitrogen sources (peptone, yeast extract, tryptone, beef extract and casein), 1 mM metal ions (Na+, K+, Zn2+, Mg2+, Mn2+, Cu2+) using casein 1.0% (w/v) as a substrate. Protease activity was determined after 24 h of incubation at 37°C. Heat killed B. megaterium PB4 cells served as control. All the experiments were conducted in triplicate unless otherwise stated. |
| Proteolytic kinetics |
| Proteolytic activity of crude enzyme was assayed using casein as substrate [28]. The crude enzyme from B. megaterium PB4 was extracted by growing the cells in 1000 ml of TSY broth (pH 7.0). The log phase culture was centrifuged at 4000 rpm at 4°C for 30 minutes. The cell pellet was washed twice with 0.1 M phosphate buffered saline (PBS) (pH 7.0) and resuspended in 40 ml buffer. The protease inhibitor, phenyl methyl sulfonyl fluoride (PMSF) was added to the resuspended cells to a final concentration of 1.0 mM. The suspended cells were disrupted by sonication (Branson Digital Sonifier®, USA) for 2 mins with 30 seconds interval in 20% amplitude. The lysate was centrifuged for 12,000 rpm for 30 mins at 4°C and the supernatant was used as crude enzyme. The protein concentration was measured by Bradford assay. Caesin at various concentration was incubated with the crude enzyme at 37°C for 30 min. The reaction was stopped by the addition of 2.5 ml of 1.2 M trichloroacetic acid. Test and test blank solutions were then filtered through a Whatman No. 1 filter paper. For 2.5 ml of the filtrate 5 ml of 0.4 M Na2CO3 and 0.5 ml of 0.5 N Folin Ciocalteu reagent were added and mixed thoroughly. Tyrosine formed was measured at OD 660nm. One unit of enzyme activity is defined as the amount of enzyme required to liberate 1 μmol of tyrosine per min under the defined assay conditions. The Michaelis-Menten parameters for proteolysis were estimated by initial substrate concentrations and the rate of proteolysis. The kinetic parameters, Vmax and Km were calculated using Lineweaver- Burk plot. |
| Deproteinization of distillery yeast biomass by culture |
| The deproteinization process of each substrate was carried out in 5 L capacity fermentor with the volume of 3 L. Dried and powered distillery yeast biomass (5.0%) was taken and mixed in 3 L of media and sterilized at 121°C for 15 min. Then 10% of the inoculum was transferred into the fermentor aseptically. The fermentor temperature was maintained at room temperature. The drive motor was allowed to rotate continuously for uniform mixing of the medium to augment the better growth of the bacterial culture. The deproteinization process was carried out up to 5 days. Periodically samples were withdrawn aseptically for the measurement of deproteinization rate and growth of PB4. |
| Enzymatic deproteinization of distillery yeast biomass |
| The crude enzyme was used for the deproteinization of distillery yeast biomass. About 10 g of distillery yeast biomass was mixed with water at a ratio of 1:3 (w/v) and the pH was adjusted to 7.0. Enzymatic deproteinization was carried out using different concentration of enzyme/substrate ratios (unit of enzyme/mg of protein) for various lengths of time. The reaction was stopped by heating the mixture at 90°C for 10 minutes to inactivate the enzyme. The mixture was centrifuged for 10 minutes at 8000 g. The pellet resulted from centrifugation was washed with deionized water and dried. The protein content was determined to determine the deproteinization rate. |
| Deproteinization (DP) was expressed as percentages and computed by the following equation as described by [29]. |
| Where, PO and PR are protein concentrations (percent) before and after hydrolysis; while, O and R represent the mass (g) of original sample and hydrolyzed residue in dry weight basis, respectively. |
| Results |
| Isolation and identification of protease producing bacterium |
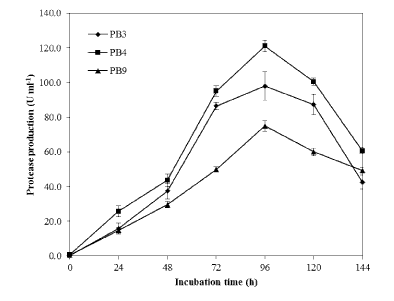
| Three proteolytic bacterial isolates namely PB3, PB4 and PB9 were found to produce clearing zone in LB agar medium containing casein (1%) which varied protease production abilities were isolated (Figure 1). During the subsequent screening experiments, microbial colony formed large and clear halos on the LB-skim milk agar was selected for further experiment. Among the isolates, PB4 was found to produce significant level of protease in the liquid medium (Figure 2) (120.3 ± 1.4 U mL-1). The isolate PB4 was subjected to taxonomic analysis as described in Bergey’s Manual of Systematic Bacteriology [22], and identified as a member of the genus Bacillus. The morphological characteristics of the isolate observed though light microscopy (Figure 3) and scanning electron microscope (SEM). The strain was Gram positive, aerobic, spore forming (spore mean length 1.0 - 1.20 μm; diameter 980 nm, and rod-shaped) (Table 1 and Figure 4). Phylogenetic analysis revealed that the isolate PB4 is affiliated with the genus Bacillus and that it closely related to Bacillus megaterium (Figure 5). |
| Optimization of growth conditions for maximizing protease production |
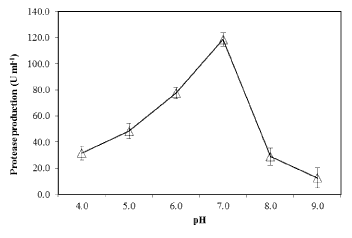
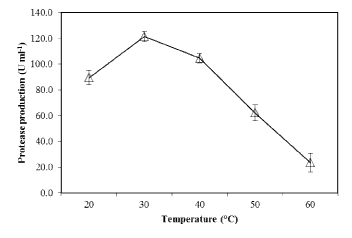
| The effect of pH on PB4’s protease production with casein (1%) as substrate, was examined. At pH 7.0 (at 30°C), the proteolytic bacteria (PB4) exhibited maximum protease production (119.7 U mL-1) (Figure 6). Further increase in pH decreased the protease production. The effect of temperature on the protease production in PB4 was examined at various temperature at pH 7.0. The protease production was found to be unaffected at temperature ranging from 20-40 °C, with an optimum at 30°C (Figure 7), whereas the production was rapidly reduced at higher temperature. |
| Protease producing ability of the strain PB4 was evaluated in batch experiments in mineral salts medium supplemented with various carbon (0.1 and 0.5%), nitrogen (0.1 and 0.5%) and metal ions (0.5 and 1.0 mM) (Table 2). Among carbon sources such as glucose, fructose, lactose, maltose, and cellulose, glucose supplementation at 0.1 and 0.5% resulted in the highest protease production of 103.5 ± 3.6 and 120.3 ± 5.3 U mL-1, respectively. Whereas cellulose resulted in the lowest protease production (8.2 ± 1.7 and 10.1 ± 2.3 U ml-1) at 0.1 and 0.5% when compared to control. On the other hand among the nitrogen sources, peptone facilitated the maximum protease production at both the test concentrations (0.1 and 0.5%) followed by yeast extract and casein supported the lowest protease production over control. Metal ions were also found to influence the protease production in PB4. Higher protease production (141.3 ± 4.7 U mL-1) was observed with the addition of Zn2+ (1.0 mM) followed by Mg2+ and Mn2+. |
| Proteolytic kinetics |
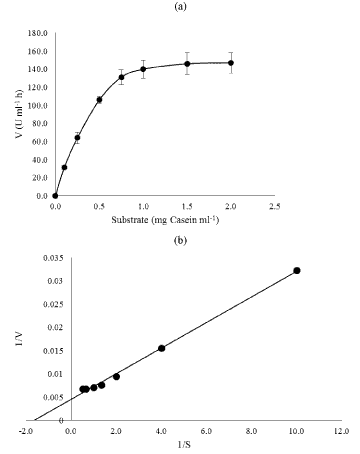
| The kinetic parameters (Km and Vmax) were determined at 37°C and pH 7.0 for casein concentrations ranging between 0.5 and 2.5 mg ml- 1. The Michelis-Menten’s constant (Km) and maximum velocity (Vmax) values of crude enzyme were 0.60 and 217.3 U mL-1, respectively (Figure 8a and 8b). In the present study, the cell free extract of B. megaterium PB4 showed higher affinity to casein was revealed by low Km value. |
| Deproteinization of distillery yeast biomass |
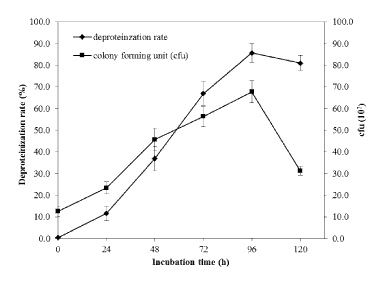
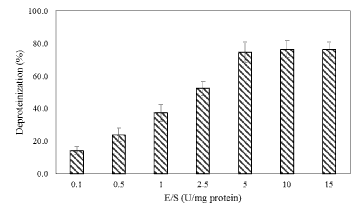
| Biological deproteinization of distillery yeast biomass was carried out in batch fermentation with PB4 culture and crude enzyme extracted from PB4. Using PB4 cell culture the deproteinization efficiency of the distillery yeast biomass was found to be 83.5 ± 2.3% in 96 h (Figure 9). The results clearly indicated that deproteinization rate and the growth of PB4 increased with incubation time. However after 96 h of incubation both the deproteinization rate and growth of PB4 were found to decrease. From the results it is evident that PB4 strain was found to grow well in distillery yeast biomass with corresponding deproteinization indicate that the organism can satisfy its carbon and nitrogen requirement without an external source. On the other hand, deproteinization of distillery yeast biomass at different substrate and enzyme concentration was carried out. Enzyme/Substrate ratio (E/S ratio) of 0.1 and 15 (Unit of enzyme/mg of protein) were used for the comparison of deproteinization rate. In general the deproteinization rate increased with the increase in E/S ratio. As shown in the Figure 10, the deproteinization rate was 14.2 and 76.4% at 0.1 and 15 E/S ratio, respectively. There were no significant increase in the deproteinization rate observed at E/S ratio beyond 5%. |
| Discussion |
| Protein hydrolysis from protein rich agro wastes is considered as a viable and cost effective technique for the production of short chain peptides and free amino acids [30]. Generally chemical and biological methods of deproteinization are widely practiced with former causing several environmental concerns, biological deproteinization using microorganisms and their enzymes proved to be effective and ecofriendly [18]. Among hydrolytic enzymes, protease catalyze the hydrolysis of peptide bond and also involves in a multitude of cellular processes as mediator of protein function. Protease production is an inherent capacity of all microorganisms; and large numbers of bacterial species are known to produce proteases. Among microorganisms, bacterial species belonging to the genus Bacillus have been extensively studied for their ability for the production of both neutral and alkaline proteases [31-33]. Among the bacterial group, interest in Bacilli gained much importance accounts for 35% of total microbial enzyme sale and it was widely used in the detergent formulation since 1960s [34,35]. Besides, protease enzyme isolated from Bacillus sp. used as additives in detergents to facilitate the release of proteinaceous materials in stains such as grime, blood, milk etc., [36]. |
| Generally production of proteases by microorganisms requires optimization of carbon and nitrogen sources in the media which is vital in determining the cost involved in the selection of substrate and enzyme production [37]. In the present study, the effect of carbon and nitrogen sources revealed glucose and peptone favoured the increased protease activity by PB4. It is evident that glucose was not found to be a repressor of protease production as reported by [38] whereas some reports suggests that glucose plays role of catabolic repression in B. licheniformis and B. subtilis, B. horikoshii, B. nesterkonia sp. AL-20 [39-42]. All the other carbons sources had resulted in the reduction of protease production. Particularly the effect of lactose in affecting protease production in Bacillus sp. has been previously reported [43,44]. In Pseudomonas aeruginosa and Pseudoalteromonas sp. strain CP76, carbon sources such as galactose, maltose and lactose were reported to increase the protease production [45,46]. Microorganisms generally require nitrogen source (both organic and inorganic forms) for their metabolism to produce primarily amino acids, nucleic acids, proteins, and cell wall components [47]. For protease production, the requirement for a specific nitrogen supplement differs from organism to organism [4]. Among nitrogen sources, both inorganic and complex sources were reported to be an effective ingredient for increasing the protease production by various microorganisms including Bacillus genus [48-50]. |
| The pH in the growth media plays a vital role in the activity and production of protease because it strongly influences many enzymatic processes and transport of various components across the cell membranes, which in turn support the cell growth and product production [38,51,52]. In some bacterial species it was reported that at neutral pH protease activity was enhanced in Bacillus sp. VITP4, Bacillus sp. PCSIR EA-3, Bacillus cereus [53-55]. Temperature is a critical factor for protease activity that should be controlled and the optimal temperature will vary from one organism to another [14]. Temperature strongly affects the synthesis of protease, either non-specifically by influencing the rates of biochemical reactions or specifically by inducing or repressing their production [56]. Temperature also influences the secretion of extracellular enzymes possibly by changing the physical properties of the cell membrane [47]. At 30 and 37°C, Bacillus cereus and Bacillus licheniformis ATCC12759 showed the maximum neutral protease activity [57]. Bacteria belonging to Bacillus genera such as Bacillus sp. JB-99, Bacillus cereus SIU1 were reported to have a wide range of temperature for maximum protease activity [58,59]. In the current study at 37°C, optimum growth for protease production was observed at which is in concordance with the earlier studies [60-62]. |
| Metal ions plays a vital role in the production of protease for its activity and stability. Proteases from B. megaterium and B circulans were reported to be zinc-containing metalloenzymes [33,63]. However in Brevibacillus brevis the protease secretion were reported to have inhibited by Zn2+ [64]. In the present study, Zn2+ stimulated the protease activity of PB4 which suggests that Zn2+ performed a vital role in maintaining the active conformation [65]. On the other hand Mg2+, Mg2+ and Cu2+ were found to be inhibit the protease activity in PB4. The inhibitory effects of metal ions on protease production and activity is well documented in literature [66]. |
| Most of the enzyme-catalyzed reactions, the Km and Vmax are considered as the important parameters [67]. In the present study, the cell free extract of PB4 exhibited lower Km and higher Vmax with casein as substrate which suggest that the crude enzyme was found to have higher affinity for casein. This results was similar to the kinetic constants documented in B. mojavensis [67]. Similarly in Brevibacterium linens and P. fluorescens RO98, the Km values were found to be 1.3 mg mL-1 and 144.28 μM, respectively with casein as substrate [68,69]. Proteolytic activity of Streptomyces sp. A6 showed highest affinity to chymotrypsin as substrate [70]. P. aeruginosa PseA strain isolated from soil capable of producing novel alkaline protease with Km and Vmax towards casein were found to be 2.7 mg mL-1 and 3 μmol min-1, respectively [71]. In P. aeruginosa MCM B-327, dehairing protease showed 2.78 mg mL-1 and 830 U min-1 of Km and Vmax, respectively with casein as substrate [72]. |
| Utilization of protease in the deproteinization of agro wastes have been previously reported. Generally the agro-waste residues with high amount of organic matter consists of 50-60% of total solid wastes which are being used as an alternative source for production of important compounds as these are valuable raw materials; rich in sources like energy and other nutrients (lignocelluloses, proteins, carbohydrates, lipids etc.,) [73]. Deproteinization of crustacean wastes by protease from Bacillus sp. TKU004, B. licheniformis RP1, B. subtilis Y-108, P. aeruginosa K-187 Streptomyces sp. Bacillus cereus SVl, Bacillus licheniformis MP1 and Zosterisessor ophiocephalus has been previously well studied and reported [74-80]. On the other hand solid and liquid wastes possessing high protein and nutrient generated from brewing industries should be given importance for the production of low cost protein products such as single cell proteins and also as feed additives [10]. In this study, deproteinization of distillery yeast biomass a second major by-product rich in nutrients and protein content [18] by whole cells and crude enzyme of PB4 was recorded. However, the neutral protease produced by B. megaterium PB4 in the present study as demonstrated by the experiments on pH-dependent activity and stability at neutral range. B. megaterium PB4 protease is metalloenzyme as suggested by the facts that the activity was increased in the presence of Zn2+ ions [33]. Although the PB4 protease is still not fully characterized, the development of optimal conditions for deproteinization of other protein-rich agro wastes is currently under investigation. |
| Conclusion |
| The application of microorganisms or proteolytic enzymes for deproteinization of distillery yeast biomass is a current research trend in conversion of wastes into useful biomass. Distillery yeast biomass is an excellent and complex microbial substrate for the growth and protease production by B. megaterium PB4. Since distillery yeast biomass is an agro waste that can be obtained at low cost, its use as a carbon and nitrogen source instead of commercial substrates may considerably reduce the cost of protease production and could promote the growth of new industrial applications. |
References
|
Tables and Figures at a glance
| Table 1 | Table 2 |
Figures at a glance
 |
 |
 |
 |
 |
| Figure 1 | Figure 2 | Figure 3 | Figure 4 | Figure 5 |
 |
 |
 |
 |
 |
| Figure 6 | Figure 7 | Figure 8 | Figure 9 | Figure 10 |
Relevant Topics
- Anaerobic Biodegradation
- Biodegradable Balloons
- Biodegradable Confetti
- Biodegradable Diapers
- Biodegradable Plastics
- Biodegradable Sunscreen
- Biodegradation
- Bioremediation Bacteria
- Bioremediation Oil Spills
- Bioremediation Plants
- Bioremediation Products
- Ex Situ Bioremediation
- Heavy Metal Bioremediation
- In Situ Bioremediation
- Mycoremediation
- Non Biodegradable
- Phytoremediation
- Sewage Water Treatment
- Soil Bioremediation
- Types of Upwelling
- Waste Degredation
- Xenobiotics
Recommended Journals
Article Tools
Article Usage
- Total views: 11660
- [From(publication date):
November-2015 - Aug 29, 2025] - Breakdown by view type
- HTML page views : 10614
- PDF downloads : 1046
