Diagnostic of Invasive Pulmonary Aspergillosis Using Nested PCR and Culture
Received: 19-Mar-2018 / Accepted Date: 23-Mar-2018 / Published Date: 30-Mar-2018
Abstract
A total of 120 pulmonary samples collected from 100 patients who were at high risk for invasive aspergillosis were examined by mycological test and nested PCR specific of A. fumigatus and A. flavus. Nested PCR was done using two sets of primers alp11/alp12 and alp13/alp14 coding for alkaline protease. Nineteen seven of these patients had negative PCR and culture results, two patients had nested PCR+/culture- PCR products revealed the presence of A. flavus. Thus only one patient had culture+/PCR-, morphologic identification characterized an A. niger. From 100 patients, only 3 had probable invasive pulmonary aspergillosis. Nested PCR and fungal culture correlate to 97% of case. Our data suggest that this nested PCR method may confirm and can improve the diagnosis of IPA in highrisk patients.
Keywords: Nested PCR; Mycology test; Invasive pulmonary aspergillosis
Introduction
Invasive aspergillosis (IA) is an increasingly important cause of morbidity and mortality, particularly in cancer patients but also in other vulnerable groups [1,2]. A. fumigatus remains the most frequent cause of IA. However, at least 20 other species including A. flavus , A. terreus, A. niger , A. nidulans, A. versicolor and A. ustus have been reported to cause human infection [3-5].
An early diagnosis and prompt initiation of appropriate antifungal therapy are imperative and essential for a favorable clinical outcome. However, an early diagnosis is usually difficult to establish, due to the limited value of microscopy, mycological cultures and conventional radiology [3,6,7].
In recent years, detection of nucleic acid by PCR has emerged as a promising tool for early diagnosis of [8,9]. Despite a variety of “inhouse” specific primers and conditions reported in published studies [5,8,10,11].
In the present study, we reported the usefulness of nested PCR methods for the detection of DNA specific for A. fumigatus and A. flavus in pulmonary samples. Then, nested PCR results were compared with mycology diagnosis.
Materials and Methods
During 3 years, a total of 120 pulmonary samples “BLA and PTP” were prospectively collected from 100 patients who were hospitalized in intensive care units of military hospital from Tunisia “HMPIT”.
Aspergillus culture
Cultures were performed by current standard accepted procedures. Samples were centrifuged at 3000 g for 10 min. The sediment was inoculated into Sabouraud agar, incubated for 5 days at 27°C and examined daily.
Identification of Aspergillus species was done on the basis of cultural and morphological characteristics. Macroscopic features like colony color, texture and margins, as well as microscopic such as size of conidia and conidiophores and their arrangements were examined for species differentiation.
DNA extraction
Extraction of DNA was done using in house protocol. Firstly, 250 μl of samples were combined with the same volume of lyses buffer containing "10 mM Tris-HCl (pH 8.0), 10 mM EDTA, 0.15M NaCl, 2% sodium dodecyl sulfate (SDS). 10 μl of proteinase K (Invitrogen) has been added and the mixture was incubated for 1 hr at 65°C. Then, proteinase K was inactivated by heating the mixture to 100°C for 15 min. An equal volume of phenol-chloroform has been added and the mixture was centrifuged for 30 min at 17000 g at +4°C. The supernatant was collected and transferred to a new tube. A double volume of absolute ethanol, stored at -20°C was added to precipitate DNA. After centrifugation (30 min at 17000 g/4°C), ethanol was removed and the pellet was washed with an equal volume of ethanol 70%. After centrifugation (30 min at 17000 g) the supernatant was removed by pipette and the residue was evaporated. The DNA pellet was dried for 1 hour and resuspended in 70 μl of elution buffer. Concentration and purity of DNA extracted are usually measured at OD260 and by 260 nm/280 nm absorbance ratio.
DNA extraction using charge switch® DNA blood
The second protocol consist to extract DNA using the charge switch DNA blood kit® "invitrogen". It is based to the use of magnetic beads which provide a surface charge dependent on pH and buffers were used to facilitate purification of nucleic acid. At acidic pH, the magnetic beads have a positive charge pull DNA molecules and Proteins are not fixed, they are eliminated. Nucleic acids are elected at the end following the increase of the pH. Magnetic beads were neutralized and the DNA elute with 150 μl of elute buffer.
PCR amplification
Nested PCR was performed according to the protocol described by Marie Pierre Hayett and her team in 1996 [8], using two sets of primers; the outer set consisted of alp11 “5’- AGCACCGACTACATCTAC-3’” and alp12 “5’- GAGATGGTGTTGGTGGC-3’”, which together amplify 740bp and 690pb sequence for A. fumigatus and A. flavus species. The inner primer set consisted of asp1 (alp13 “5’- CTGGCATACAACGCCGCTG-3’” and alp14 “5’- TTGTTGATCGCAACC-3’”, which together amplify 527bp of Aspergillus fumigatus .
The first amplification consisted of 5 μl buffer solution (1x), 0.5 μl of MgCl2 (25 mM), 2 μl of dNTPs (10 mM), 1 μl of each external primer (25 μM), 1.25 U of DNA polymerase (Biomatik) and 1 μl (50 ng/μl) of DNA. PCR was performed under the following conditions: Denaturation at 95°C for 5 min, annealing at 50°C for 1 min, and extension at 72°C for 60 sec for 30 cycles. The last round of PCR was performed at 72°C for 7 min.
The second amplification was accomplished by using 2 μl of the first product as a template. PCR reaction was performed in 50 μl of PCR solution containing 5 μl of buffer solution (1x), 2 μL of MgCl2 (25 mM), 1 μl of dNTPs (10 mM), 1 μl of each primer and 1.25 U of DNA polymerase (biomatic).
The amplification process was performed under the following conditions: 30 cycles of 94°C for 5 min for denaturation, 63°C for 60 sec for annealing and extension at 72°C for 2 min. Finally, last extension was done at 72°C for 7 min. The PCR products were electrophoresis on a 2% agarose gel containing Redsef and photographed with a UV transillumination camera.
Result
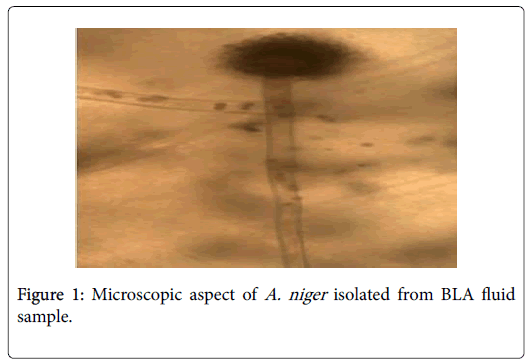
Phenotypic analysis of cultures (macroscopic and microscopic identification) showed that 119 samples were negative and only one was positive. An A. niger was macroscopic and microscopic identified (Figure 1).
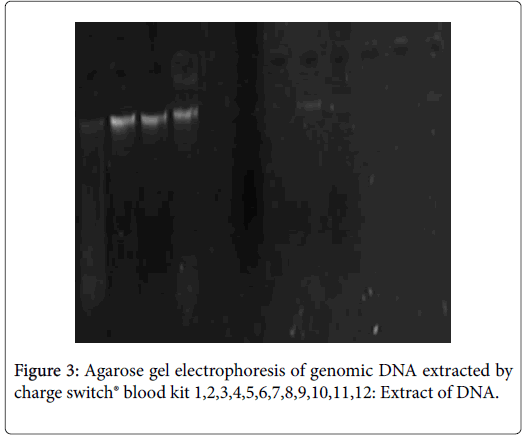
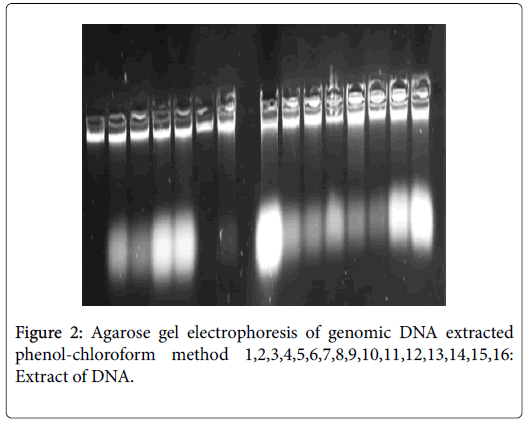
Following lacked standardization of PCR extraction protocol, we have optimized in house protocol using phenol-chloroform, it is matched more efficient than «Charge Switch blood®» protocol. Indeed, DNA extracted by the first protocol has high degree of purities (1.8_2.2) and quantity (1500 and 5000 ng/μl) (Figure 1). In contrast, the second protocol give 100 ng/μl as maximum and purity included between 1 and 1.4 (Figures 2 and 3).
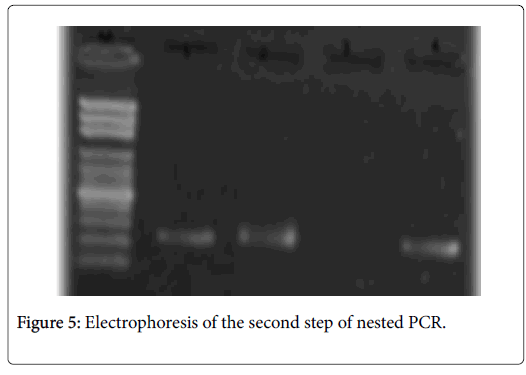
The amplification of the gene encoding of alkaline protease was positive only in two cases. The resulting profile is corresponding to A. flavus; two bands were observed; two bands of 690bp and 275pb were detected after external and internal amplification (Figures 3-5). In our study, we didn’t detected A. fumigatus profile.
Discussion
Invasive pulmonary infection caused by Aspergillus spp. is associated with a very high mortality in immunocompromised patients. IPA usually caused by Aspergillus fumigatus . Therefore, A. flavus was also involved of IPA [8,12]. The diagnosis of IPA is often very tardif. It is based on radiological, biological and clinical arguments. An early diagnosis is critical for a successful treatment [1,4].
Following lacked standardization of PCR extraction and amplification technique to detect Aspergillus DNA from pathological sampling; we have chosen to optimize in house extraction protocol. Extraction with phenol-chloroform appears much efficient than the commercial protocol. The use of phenol-chloroform for protein separation and purification guarantee to obtain significant purity and quantity of DNA. According to the literature, the use of technologybased phenol chloroform provides nucleic acid with high quantity [8].
Microbiological diagnosis of IPA is difficult to establish, as cultivation of Aspergillus has poor sensitivity and may require several days for fungal growth to be evident. Thus, molecular diagnostic techniques such as nucleic acid detection by PCR are emerging as potentially more sensitive and rapid alternatives to conventional techniques for the diagnosis of IPA [2,3,6].
In this report, we describe a rapid, sensitive and reliable molecular technical for the detection and identification of two Aspergillus species. Primers were choosing to detect alkaline protease gene specific of A. fumigatus and A. flavus .
Aspergillus conidia are ubiquitous in the environment, creating the potential for false positive PCR results [8]. Fungal cells or fungal DNA can enter the assay process at numerous points, including at the time of BAL collection, during DNA extraction, or at nested PCR set up. UV irradiation had used to control PCR contamination.
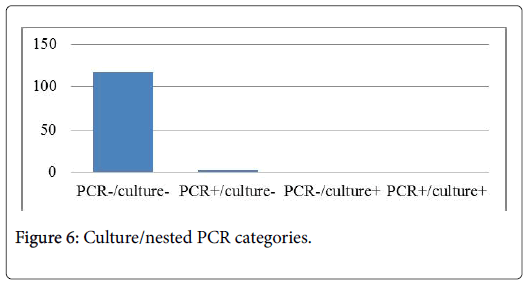
Alkaline protease gene was detecting positive into two BLA, an A. flavus was reveled after electrophoresis. In the other hand, morphologic characters of culture showed that only one BLA was positive, the strain was an A. niger (Figure 6).
Detection of Aspergillus in bronchiolar-alveolar fluid (BLA) is one of the criteria for the probable or proven diagnosis of invasive aspergillosis (AI). We have diagnosed 3 patients with probable IPA and 97 patients were defined as not having IPA.
The positive and negative predictive values were respectively 100% and 98.9% for nested PCR, 100% and 97.9% for mycological test. Nested PCR showed a higher sensitivity (66.66%) than mycological test (33.33%) and heir specificity were 100%. The correlation coefficient (r) calculated between fungal culture and nested PCR in BAL was 0.97. BAL culture was somewhat more sensitive than PCR in detecting IPA (sensitivity 84.6%) and had high specificity (100%). In the other hand nested PCR was more sensitivity [1,3-5,8,10,13].
Indeed, sensitivity has been highly variable because of differences in assay protocols characteristics of patients evaluated, and form of IA diagnosis (proven or probable) [11,13,14].
Conclusion
We were successful detecting DNA specific for Aspergillus flavus in BLA samples from patients with probable IA by using the nested PCR. Nested PCR of BAL fluid is an accurate test to diagnose culturenegative patients with high risk of IPA. It should be included in routine laboratory practice.
Disclosure of Interest
The authors declare that they have no competing interest.
Acknowledgement
We thank very much the staff of parasitology-mycology service “Military Hospital of Tunisia” for their valuable technical assistance.
References
- Avni T, Levy I, Sprecher H, Yahav D, Leibovici L, et al. (2012) Diagnostic accuracy of PCR alone compared to galactomannan in bronchoalveolar lavage fluid for diagnosis of invasive pulmonary aspergillosis: A systematic review. J Clin Microbiol 50: 3652-3658.
- Buess M, Cathomas G, Halter J, Junker L, Grendelmeier P, et al. (2012) Aspergillus-PCR in bronchoalveolar lavage for detection of invasive pulmonary aspergillosis in immunocompromised patients. BMC Infect Dis 12: 237.
- Buchheidt D, Baust C, Skladny H, Ritter J, Suedhoff T, et al. (2001) Detection of Aspergillus species in blood and bronchoalveolar lavage samples from immunocompromised patients by means of 2-step polymerase chain reaction: Clinical results. Clin Infect Dis 33: 428-435.
- Buchheidt D, Baust C, Skladny H, Baldus M, Brauninger S, et al. (2002) Clinical evaluation of a polymerase chain reaction assay to detect Aspergillus species in bronchoalveolar lavage samples of neutropenic patients. Brit J Haemat 116: 803-811.
- Hinrikson HP, Hurst SF, De Aguirre L, Hinrikson HP, Hurst SF, et al. (2005) Molecular methods for the identification of Aspergillus species. Med Mycol 43: 129-137.
- De Aguirre L, Hurst SF, Choi JS, Shin JH, Hinrikson HP, et al. (2004) Rapid differentiation of Aspergillus species from other medically important opportunistic molds and yeasts by PCR-enzyme immunoassay. J Clin Microbiol 42: 3495-3504.
- Francesconi A, Kasai M, Petraitiene R, Petraitis V, Kelaher AM, et al. (2006) Characterization and comparison of galactomannan enzyme immunoassay and quantitative real-time PCR assay for detection of Aspergillus fumigatus in bronchoalveolar lavage fluid from experimental invasive pulmonary aspergillosis. J Clin Microbiol 44: 2475-2480.
- Hayette MP, Vaira D, Susin F, Boland P, Christiaens G, et al. (2001) Detection of Aspergillus species DNA by PCR in bronchoalveolar lavage fluid. J Clin Microbiol 39: 2338-2340.
- Hedhili S, Baccouchi N, Souid H, Jemli B, Gargouri S (2017) Comparison of real-time PCR, nested PCR and galactomannan antigen detection by enzyme-linked immunosorbent assay in sera for diagnosis of invasive aspergillosis. J Med Mycol 27: 487-493.
- Musher B, Fredricks D, Leisenring W, Balajee SA, Smith C, et al. (2004) Aspergillus galactomannan enzyme immunoassay and quantitative PCR for diagnosis of invasive aspergillosis with bronchoalveolar lavage fluid. J Clin Microbiol 42: 5517-5522.
- Reinwald M, Hummel M, Kovalevskaya E, Spiess B, Heinz WJ, et al. (2012) Therapy with antifungals decreases the diagnostic performance of PCR for diagnosing invasive aspergillosis in bronchoalveolar lavage samples of patients with haematological malignancies. J Antimicrob Chemother 67: 2260-2267.
- Jalava RK, Laaksonen S, Issakainen J, Vauras J, Nikoskelainen J, et al. (2003) Semiquantitative detection by real-time PCR of Aspergillus fumigatus in bronchoalveolar lavage fluids and tissue biopsy specimens from patients with invasive aspergillosis. J Clin Microbiol 41: 4304-4311.
- White PL, Bretagne S, Klingspor L, Melchers WJ, McCulloch E, et al. (2010) Aspergillus PCR: One step closer to standardization. J Clin Microbiol 48: 1231-1240.
- Skladny H, Buchheidt D, Baust C, Krieg-Schneider F, Seifarth W, et al. (1999) Specific detection of Aspergillus species in blood and bronchoalveolar lavage samples of immunocompromised patients by two-step PCR. J Clin Microbiol 37: 3865-3871.
Citation: Hedhili S, Souid H, Baccouchi N, Jemli B, Gargouri S (2018) Diagnostic of Invasive Pulmonary Aspergillosis Using Nested PCR and Culture. Arch Sci 2: 115.
Copyright: © 2018 Hedhili S, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Select your language of interest to view the total content in your interested language
Share This Article
Open Access Journals
Article Usage
- Total views: 7795
- [From(publication date): 0-2018 - Dec 22, 2025]
- Breakdown by view type
- HTML page views: 6606
- PDF downloads: 1189