Diversity, Seasonal Variation and Antibacterial Activity of Endophytic Fungi Associated With the Genus Jatropha in Mauritius
Received: 04-Dec-2017 / Accepted Date: 18-Jan-2018 / Published Date: 25-Jan-2018 DOI: 10.4172/2155-952X.1000280
Abstract
Background: Endophytic fungi are important components of the forest community and are significantly diverse from one plant to another as well as from one ecosystem to another.
Objective: The current study aimed to characterise, at the molecular level, the diversity and seasonal variation of endophytic fungi from Jatropha plants found in Mauritius and to compare the phytochemicals and the antimicrobial potential of the fungal isolates with the leaves extracts.
Materials and methods: Endophytic and saprophytic fungi were isolated from the leaves of Jatropha plants during summer and winter. The isolated fungi were further classified through molecular characterisation. The isolates were grouped into 76 distinct operational taxonomic units based on the sequence of the internal transcribed spacer regions in the rRNA gene. The colonization frequency and the dominant fungi percentage of these endophytic fungi were calculated. The antimicrobial properties of the extracts from the endophytes were compared to that of the extracts obtained from the leaves of the Jatropha plants.
Results: The overall colonisation rate for the two seasons was 67.42%. Maximum colonisation (27%) was observed in both J. curcas and J. multifida. There was a diverse array of fungi which included 21 common genera. Colonisation frequency of the other genera recovered during this study varied according to the plant from which the isolation was carried out. J. multifida was richer in the genus Phoma, J. curcas were colonised mainly by Neofusicoccum as compared to J. integerrima and J. podagrica, which were colonised by the Corenespora. Climate was also a primary driver of endophytes and saprophytes community diversity and composition. Moreover, the endophytic fungi from the leaves of J. curcas gave highly significant antimicrobial activities effects (p<0.01) against both the tested Gram (+ve) and Gram (-ve) clinical pathogens. Among the four active isolates, Engyodontium was the genus that showed highest antimicrobial activity. Overall, the endophytic fungi from the Jatropha species were more effective than the crude ethyl acetate extract of the leaves. Conclusion: The antimicrobial activity of these endophytic microorganisms could be further exploited and find application in the pharmaceutical industry.
Keywords: Antimicrobial activity; Endophytes; Fungi; Jatropha
Introduction
The development of resistance by pathogenic bacteria to commercial drugs is a problem faced by health services and has become a serious concern worldwide. Several factors such as extensive or inappropriate use of antibiotics have increased numbers of immunocompromised patients. This has led to intensive search for new and effective antimicrobial agents by exploring new niches and habitats. Endophytes are symbiont organisms that colonize the internal tissues of higher plants without causing any symptoms and these microbes can be isolated from plant tissues using strict surface-sterilized methods [1]. Endophytic fungi have been isolated from different kinds of medicinal plants. Fungal endophytes are thought to harbour biologically distinctive natural and bioactive products [2,3]. Numerous antimicrobial metabolites have been described from endophytic fungi pertaining to different structural classes such as alkaloids, peptides, steroids, terpenoids, phenols, quinines and flavonoids [1,4,5]. A considerable number of researchers have investigated the, ecological role, secondary metabolites and bioactivity of the endophytic fungi isolated from various medicinal plants [6-8]. However, it remains unknown if the efficacy attributable to antibacterial properties of the endophyte and the plant from which it has been recovered exhibit same properties.
Moreover, it is expected that plants unique to specific areas are likely to yield a high diversity of endophytes. Mauritius, a sub-tropical volcanic island, is situated in the South-West of the Indian Ocean and covers a total land area of 1860 km2. The topography of Mauritius makes the central plateau more humid (80-85%) and cooler than the other regions. In summer, the island is under the influence of moist, maritime airflow. Temperatures are high; the warmest months lasts from October to May with temperatures generally average about 27°C, with mean daily maxima from 30°C to 38°C. Summers are usually somewhat wetter than winters, with much of the rainfall coming from convectional thunderstorm activity; tropical cyclones also enhance warm-season rainfall. The coldest month is usually quite mild (12°C-20°C), although frosts are uncommon, and winter precipitation is derived primarily from frontal cyclones [9]. Thus, climate coupled with the location of the island favours an ideal niche for an ecologically diverse community of fungal endophytes.
Jatropha is an important plant which has received more attention in recent years for its utilization in biodiesel production as well as for other beneficial uses, such as antimicrobial and pesticidal activity [10,11]. J. curcas is widely grown in Central and South America, Southern Asia, and Central-Southern Peninsular Asia [12,13]. In Mauritius, this plant is mainly distributed all around the island [14]. Previous studies on the endophytes showed that medicinal plants harbour endophytic strains with antimicrobial properties [4,15]. To date, a small number of endophytic fungi have been isolated from J. curcas [14,16]. Moreover, information about the diversity and the antibacterial effects of the secondary metabolites isolated from potential endophytic fungi associated with Jatropha species and their interactions with host is scarce.
Objectives
The aim of this study was to couple all the variables that have proved to be significant on the fungal population inhabiting plants tissues of Jatropha. The endophytic fungi in four Jatropha species, J.curcas, J. multifida, J. podagrica and J. integerrima were isolated, characterised and identified. The seasonal variation among the endophytic fungi population and host affiliation of the isolates to their respective host were reported. The fungal endophytes were screened for their primary phytochemicals and were tested for their antimicrobial properties. A comparison of effectiveness of the antimicrobial properties of the fungal isolates and the extracts of leaves obtained from the four Jatropha species was made.
Materials and Methods
Plants and fungi samples
J. curcas L. (Acc. No. MAUR 26484), J. integerrima Jack (red) (Acc. No. MAUR 26486), J. multifida L. (Acc. No. MAU 26487) and J. podagrica Hook (Acc. No. MAUR 26485) were chosen for this study. The isolation from healthy mature and dead leaves was carried out. The plants species were randomly selected from the North, South, East, West and Central regions of the island. The collections of samples were carried out during both winter and summer months. Leaves were randomly collected immediately washed in running distilled water for 15 min. From each leaf, 10 disks of 5 mm diameter were cut and subjected to the surface sterilization procedure. This includes soaking in 95% ethanol for 1 min, then immersed in a solution of sodium hypochlorite (3%) for 1 min and finally immersed in 95% ethanol for 30 s. Samples were then washed, dried, and plated on potato extract agar supplemented with Rose Bengal. Plates were incubated at 25°C under a cycle of 12 h light/ dark for 1-3 weeks and hyphal tips any fungal colonies were transferred onto potato dextrose agar (PDA) plates.
DNA extraction, amplification, phylogenetic analysis
DNA was extracted from fresh fungal culture. Mycelia were directly scraped off from fungal cultures grown on PDA plates for 5-10 days. The mycelia were transferred into 1.5 mL centrifuge tube and mixed with 0.2 g sterile white quartz sand. 600 mL preheated CTAB buffer (2% v/w CTAB, 100 mM Tris-HCl, 1.4 M NaCl, 20 mM EDTA, pH 8.0) were added to the centrifuge tube and with a plastic pestle the mixture were grinded manually. The mixture was then incubated at 60°C water bath for 30 min. Then 600 mL phenol/chloroform (1:1) was added into each tube and inverse-mixed. The mixture was then centrifuged at 13,000 rpm for 30 min at room temperature. The procedure was repeated twice. Two volumes of 100% cold ethanol were added, and the tube was inverted gently to mix and stored overnight at 20°C for precipitate DNA [17].
Primer pairs ITS4 and ITS5 were used to amplify the 5.8S gene and flanking ITS1 and ITS2 regions. Amplification was performed in a 50 μL reaction volume containing 5 μL of 10x Mg free PCR buffer, 3 μL of 25 mM MgCl2, 4 μL of 2.5 mM deoxyribonucleotide triphosphates (dNTPs), 1.5 μL of 10 μM primers (ITS4 and ITS5) and 3 μL of DNA template, 0.3 μL of 2.5 units of Taq DNA polymerase. The thermal cycle of 3 min initial denaturation at 95°C, followed by 30 cycles of 1 min denaturation at 95°C, 50 s primer annealing at 52°C, 1 min extension at 72°C, and a final 10 min extension at 72°C was used. The PCR products were examined by electrophoresis in 1.5% (w/v) agarose gel with ethidium bromide (10 mg/mL) and checked for size and purity. Purified PCR products were then directly sequenced in an automated sequencer at Inqaba, South Africa
Phylogenetic analysis
The sequences of the fungi were aligned and adjusted manual using MEGA® 6.0. Phylogenetic trees were constructed using maximum parsimony method (MP) using PAUP® Phylogenetic Software version 4.10. The unconstrained topologies of the equally parsimonious trees were compared using the Kishino-Hasegawa test of PAUP®. Additionally, analysis was also constructed using Bayesian inference. The best evolutionary models were selected using the software Mr ModelTest® v. 2.3 based on the values of the Akaike information criterion (AIC). GenBank accession numbers for your nucleotide sequences: MF164486- MF164558.
Diversity and statistical analysis
The composition, coverage, richness, relative colonisation frequency and abundance of the fungal communities were compared among type of isolates by location and season using diversity indices such as Shannon- Wiener, Chao1, t-tests and analysis of variances were performed to evaluate the distribution. The antimicrobial assays data were expressed as mean ± standard deviation with one/two-way ANOVA at 5% and LSD test compared the differences between the means.
Screening of the fungi secondary metabolites and antimicrobial activity assay
The methods described by Rampadarath et al. [11] were used for the screening of secondary metabolites such as flavonoids, alkaloids, steroids, tannins, coumarins and phenols based on a series of test tube tests. Crude ethyl acetate extracts were obtained using the decoction method. 20 g of mycellia and Jatropha species leaves were macerated in 40 mL of the solvent systems: ethyl acetate for 48 h. All the inoculates were standardized by dilution with sterile nutrient broth (0.1 mL inoculum in 9.9 mL Muller Hinton broth) to an absorbance of 0.4-0.6 at 600 nm. The test microorganisms included in this study were six Gram (+ve) Bacillus cereus ATCC 11778, Campylobacter jejuni NCTC 11351, Clostridium perfringens ATCC 13124, Listeria monocytogenes ATCC 33090, Staphylococcus aureus ATCC 29213 , Staphylococcus epidermis ATCC 12228 and seven Gram(-ve) Enterobacter cloacae ATCC 13044, Enterococcus faecalis ATCC 29212, Escherichia coli ATCC 25922 , Klebsiella oxycota ATCC 43086, Proteus mirabilis strain NCTC 11938, Pseudomonas aeruginosa ATCC 27853 and Serratia marcescens ATCC 14756.
Results
Phylogenetic analysis
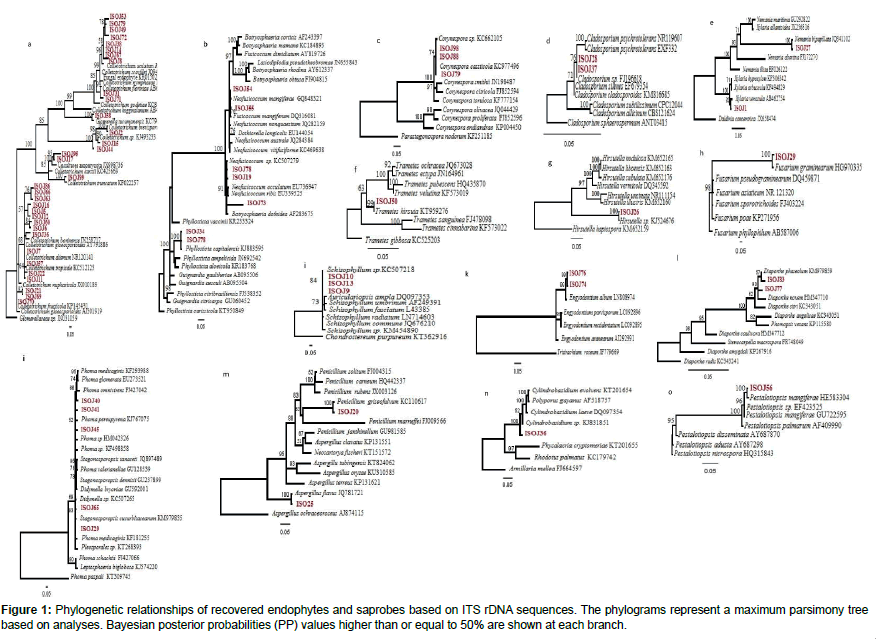
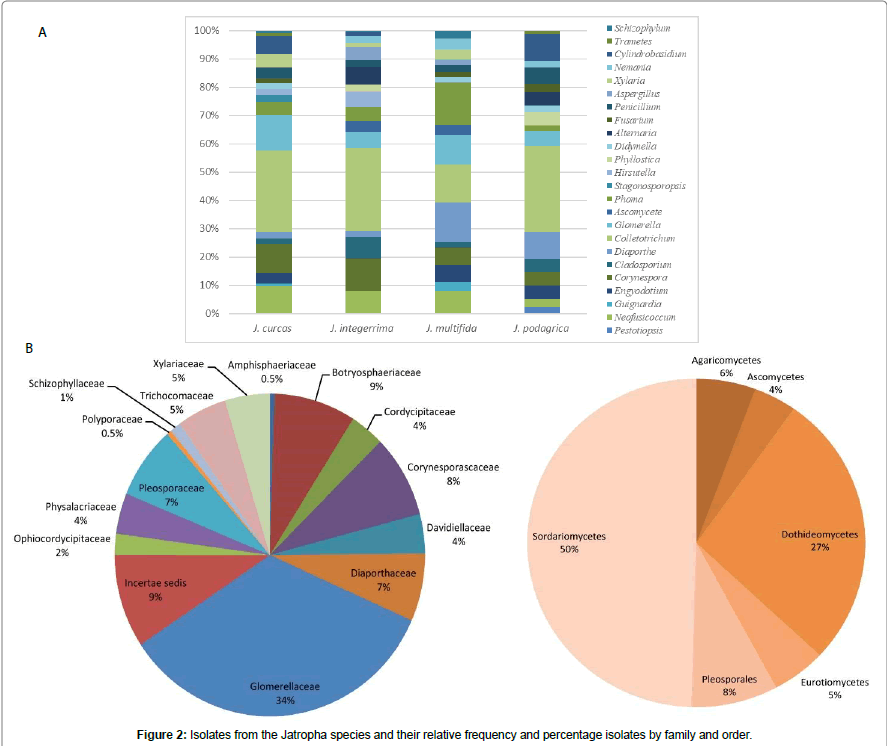
The trees illustrate the results of maximum likelihood (ML) analysis of ITSrDNA data; supporting values are ML bootstrap values (before slash; values ≥ 60%) and Bayesian posterior probabilities values ≥ 50% are shown (Figure 1). Based on the morphological characteristics, the 76 isolates were characterised molecularly. The break down % of the isolated endophytes and saprophytes into families classification were 34% Glomerellaceae, 9% Botryosphaericea and Incertae sedis, 8% Corynesporascaceae, 7% Diaporthaceae and Pleosporacea, 5% Trichocomaceae and Xylariaceae, 4% Cordycipitaceae, Davidiellaceae and Physalacriaceae, the remaining % to Amphisphaeriaceae, Ophiocordycipitaceae, Schizophyllaceae and Polyporaceae (Figures 2a and 2b). The overall colonisation rate for the two seasons was 67.42%, with maximum colonisation (27%) observed in both J. curcas and J. multifida. There was a diverse array of fungi which included 21 common genera (Figure 2a) and 3 uncommon endophytic and saprophytic genera. The fungal species abundance distribution was found to be highly skewed with many frequent species and incidental ones. Among the genera, Colletotrichum was the dominant one with relative frequency of 62 isolates isolated from the leaves of J. curcas and J. integerrima. Additionally, the relative abundance of each common genus was also different between the four Jatropha species. The colonisation frequency of the genera recovered during this study varied according to the plant from which the isolation was carried out. For example, J. multifida was richer in the genus Phoma, J. curcas were colonised mainly by Neofusicoccum as compared to J. integerrima and J. podagrica, which were colonised by the Corenespora genus.
Seasonal and regional variation in the endophytes and saprophytes communities
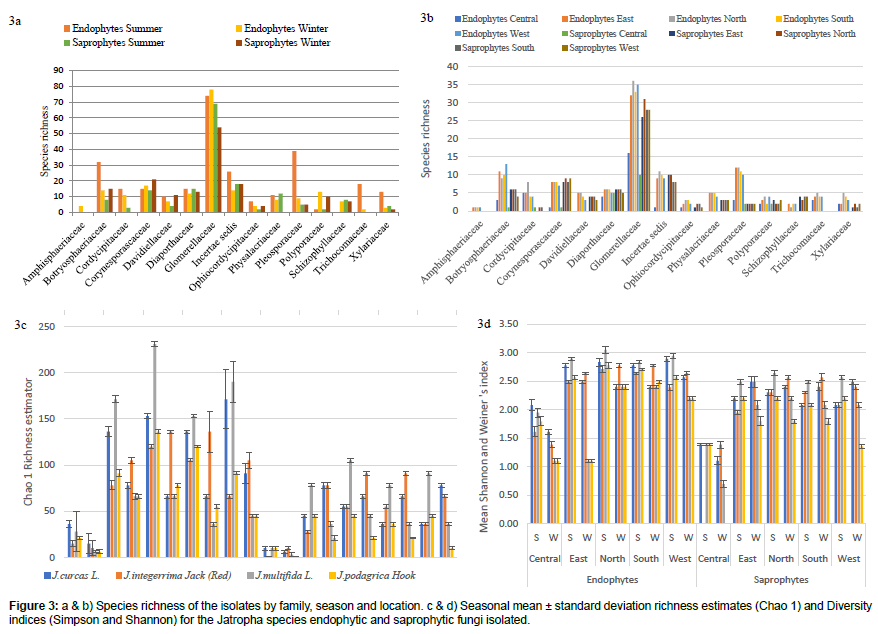
The fungal species richness and species composition were differed significantly (p<0.05) between types of isolates, season and locations. The abundance and composition for endophytes (60%) were higher compared to the number of saprophytes isolated from the four Jatropha species and the five locations during both winter and summer. Moreover, the highest endophytic (twenty one) and saprophytic (fourteen) OTUs obtained was in the north during summer from the J. multifida. Except for J. integerrima, the other three Jatropha species were rich in both types of fungi, endophytic and saprophytic during summer than winter months. Also, the saprophytes were lowest in the central location for the Jatropha species during both seasons (Figure 3a).
Composition of the saprophytes and endophytes families during summer and winter
The fungal communities associated with season were compared (Figure 3b). A significant seasonal difference (p<0.05) in colonisation among the families was obtained as a total of 275 Glomerellaceae isolates were recovered of which 74 were summer endophytes, 78 winter endophytes, 69 summer saprophytes and 54 winter saprophytes. In addition, 69 isolates were isolated from Botryosphaeriaceae family with more summer endophytes (thirty two) and 67 from Corynesporascaceae with more winter saprophytes (twenty one) (Figures 3a and 3b). Variation in the representation of the individuals in the community was also observed, as 15 families were recovered at the 5 locations around Mauritius.
Diversity indices provide important information about rare and common species in a community (Figures 3c and 3d). There were differences in fungal diversity, species abundance and species richness of the Jatropha species between the different locations during summer and winter. The maximum Shannon-Weiner diversity index was found for summer endophytes in the north (3.045) followed by again summer endophytes in the west part of the island (2.944) isolated for J. multifida and least in the central part of the island for summer saprophytes colonised from J. integerrima. Shannon-Weiner index as well as species richness Chao 1 was found to be higher (231) in the north of Mauritius from J. multifida in summer season (Figures 3c and 3d). The greatest species richness per sampling was recovered in the north.
Phytochemical constituents of the crude ethyl acetate of Jatropha species endophytes and mature leaves extracts
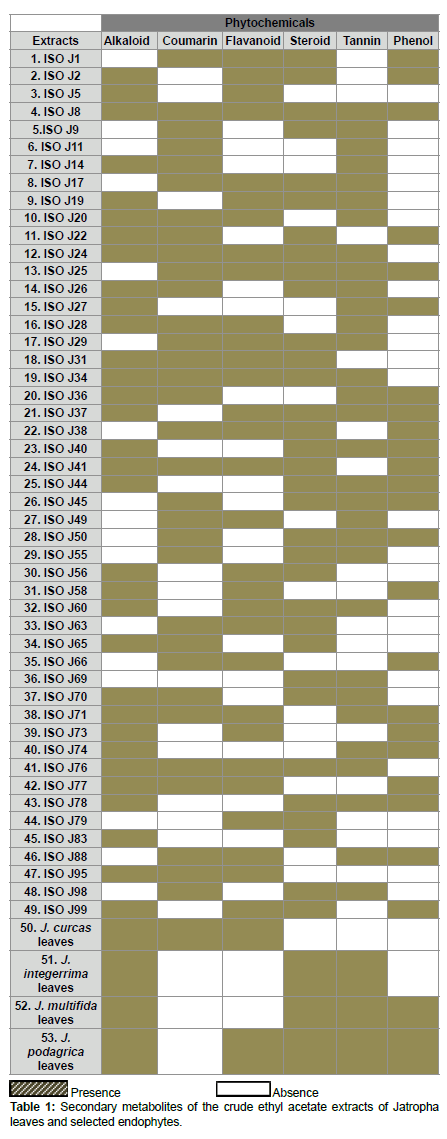
The qualitative preliminary test tube analysis of the different Jatropha species mature leaves and endophytes extracts were tested positive for most of the secondary metabolites - alkaloid, coumarin, flavonoid, steroid, tannin and phenol. 61% of the endophytes extracts revealed the presence of more than three secondary bioactive metabolites. Among the 49 endophytes, more than 72% of them contained the phytochemicals - alkaloid, coumarin, flavonoid, steroid and tannin and 47% had phenol (Table 1).
Antimicrobial activities of the crude ethyl acetate extracts of endophytic fungi and leaves of Jatropha species
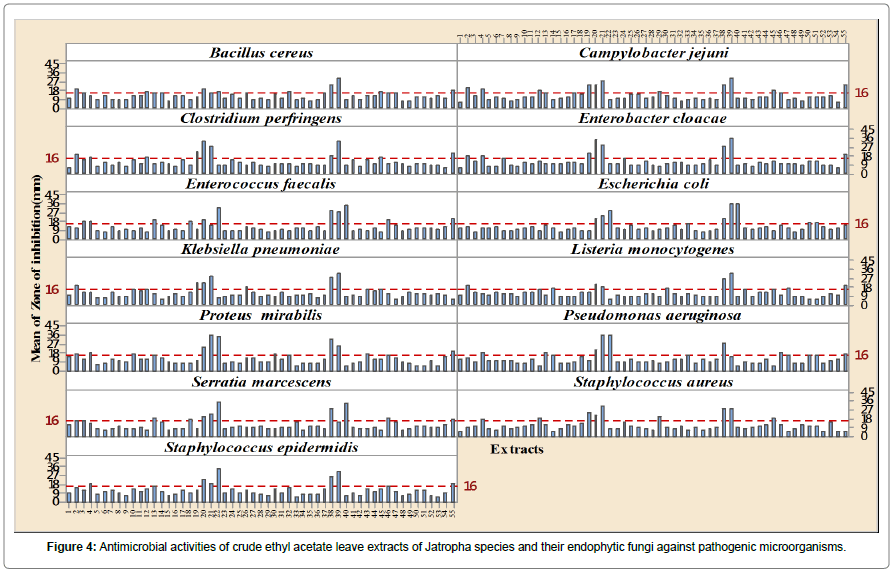
The endophytic fungi from the leaves of J. curcas gave highly significant antimicrobial activities effects (p<0.01) against both the tested Gram (+ve) and Gram (-ve) clinical pathogens and endophytes ISO J34, ISO J36, ISO J73 and ISO J74 were the most effective ones (Figure 4 and Table 2). The widest spectrum of antimicrobial activity was demonstrated by three endophytes ISO J34, ISO J73 and ISO J74, and the latter were very effective on eleven of the thirteen pathogenic microorganisms tested with mean IZ mm of above 16 mm. Among the four active isolates, Engyodontium was the genus that showed highest antimicrobial activity. Most of the pathogens were highly susceptible to the crude ethyl acetate extract of Endophyte ISO J74 with mean IZ (mm) for the Gram (+ve) varying from 29.20 ± 1.23 mm (Staphylococcus aureus) to 33.60 ± 2.30 mm (Clostridium perfringens) and for Gram (-ve) from 35.80 ± 0.45 mm (Escherichia coli) to 25.40 ± 4.72 mm (Proteus mirabilis) (Figure 4) Overall antimicrobial activities of the crude ethyl acetate extracts of the endophytic fungi from the Jatropha species were more effective than the crude ethyl acetate extract of the leaves of J. curcas and both the negative and positive control, tetracycline. Ethyl acetate proved to be a good polar solvent.
| Endophytic fungi | Maximum IZ (mm diameter) | Gram (+ve) and Gram (-ve) bacteria |
|---|---|---|
| ISO J34: Phyllosticta capitalensis | 34.00 ± 0.00 | Enterococcus cloacae ATCC 13044 |
| ISO J36: Colletotrichum boninense | 35.60 ± 0.00 | Proteus mirabilis NCTC 12453 |
| ISO J73: Botryosphaeria dothidea | 31.60 ± 0.00 | Serratia marcescens ATCC 14756 |
| 25.00 ± 1.23 | Enterococcus cloacae ATCC 13044 | |
| ISO J74: Engyodontium album | 35.80 ± 0.00 | Escherichia coli ATCC 25922 |
| 29.20 ± 1.23 | Staphylococcus aureus ATCC 29213 | |
| 33.60 ± 2.30 | Clostridium perfringens ATCC 13124 | |
| 35.80 ± 0.45 | Escherichia coli ATCC 25922 | |
| 25.40 ± 4.72 | Proteus mirabilis NCTC 1245 |
Table 2: Summary of endophytic fungi with highest zone of inhibition (IZ) against Gram (+ve) and Gram (-ve) bacteria.
Discussion
Phylogenetic diversity
The fungi recovered in the study belonged to the Glomerallaceae family. The trees were congruent in topology and were supported with >50% of 1000 bootstrap replicates. The family comprised of members from of both endophytes and saprophytes. Glomerella and its anamorph Colletotrichum species are pathogens of plants distributed worldwide and have also frequently been isolated as endophytic fungi. In the study of Kumar and Kaushik [14] six species from J. curcas leaves were reported as Colletotrichum truncatum. The isolate Colletotrichum truncatum EF10 yielded oil and the fatty acid similar to the oil isolated from Jatropha seed oil. Our phylogeny confirms that endophyte ISO J58 is Colletotrichum truncatum as it formed a terminal cluster with strong bootstrap support of 100% Bayesian support. The phylogenetic relationship of endophytic fungus and the plant species therefore indicates that the isolates may have an important role in the plant’s ability to produce oil.
Endophytic and saprotrophic loculoascomycetes
Phylogenetic analysis and sequence similarity comparison of the ITS–5.8S gene sequences showed that some of the taxa isolated from the Jatropha plants were from Loculoascomycetes (Pleosporales and Botryosphaeriaceae). Both families have been formerly reported as endophytic and pathogenic [18]. The phylogeny confirms a close relationship between endophytic fungi ISO J34 and saprophyte ISO J78 forming a monophyletic clade with Phyllosticta capitalensis (teleomorph: Guignardia) genus. This study reports for the first time Phyllosticta species from Jatropha. Despite the close resemblance in the DNA sequences of the isolates and the phylogenetic affiliation among the two isolates, the time at which they were recovered were different. It is possible that the endophyte ISO J34 changed its ecological strategies and adopted a saprophytic life style as often predicted in previous studies [19]. Corynespora is hyphomycetous identified as isolates ISO J98, ISO J88 and ISO J79. The endophytic and saprotrophic phenotypes of Corynespora cassiicola were identical in both morphology and sequences. Corynespora cassiicola that was recorded as ISO J98 and ISO J79 from living tissues of Jatropha was also recovered as ISO J88 during later decomposition stages of the leaves.
Cladistic analyses in this study also showed that of Xylaria species can coexist within leaf tissues of Jatropha in both their anamorphic and teleomorphic forms. Saprotroph ISO J27 was identified as a Nemania bipapillata and endophyte ISO J1 as Xylaria venosula. Among these two isolates, there were only 6 nucleotide differences in the ITS and 5.8S regions (4 bp insertions in Endophyte ISO J1 and 2 transitions from C to T in Saprobe ISO J27). Members of the Xylariaceae are well known for their role in wood decomposition hence confirming the presence of ISO J27. It is highly probable that endophytic Xylaria species turn to their teleomorphs stage on fallen leaves.
Host specificity and richness
A diverse fungal community of different taxa were found living the four species of Jatropha growing all over Mauritius. Fungal genera such as Diaorpthe, Colletotrichum, Aspergillus, Phoma Fusarium, Penicillium and Xylaria obtained in this study have been described as host generalists endophytes and the most frequent isolates of tropical plants [20]. However, endophytic species such as Hirsutella, Trametes, Schizophyllum and Pestalotiopsis were among the less dominant endophytic species and host-specific fungi in the Jatropha species. The current results also show that more than one species of Colletotrichum can inhabit the plants [8]. The fungal community were dominated by Ascomycetes and more specifically, by the classes Sordariomycetes and Dothideomycetes, which represented 50% and 27% of total fungi, isolated. Fungi expressing their pathogenic behaviour in one host but acting as endophytic in other host species are not uncommon [21]. The presence of some well-known pathogens, such as Colletotrichum sp., Fusarium sp. and Neofusicoccum sp. and Phoma sp. were identified.
Seasonal variation
Plant-inhabiting endophytes belong to a diverse and active group of plant associated fungi that form part of plants in different environments. Climate was the primary driver of endophyte and saprophytes community diversity and composition in this study [22]. However, some species turnovers were not affected by the climate but the role of the fungi acting as endophyte or saprophytes were determined by the climate change [23]. Fluctuation of endophytic number during winter and summer season was detected, [24]. It is also noteworthy that there was only a small pattern in the summer samples and those patterns were 100% recorded in the winter samples. This may lead to one explanation of this situation that the endophyte population depends on the physiological status of the host plant, which, in turn, is partly related to the seasonal weather variation, though only two seasonal samples were collected in this experiment.
Secondary metabolites
Research on active secondary metabolites produced by microorganisms continues to be a significant area of attention in order to cope with the increasing demand for the treatment of human diseases. Over the last several decades, natural products have contributed highly in drug discovery. The antifungal activity of endophytic fungi isolated from J. curcas plants have been reported in previous studies carried out by Kumar and Kaushik [14], while the plants present in Mauritius have not gained much research attention. Therefore, the prevalence of the fungal community isolated as endophytes and saprophytes were screened for their phytochemicals and inhibitory action against bacterial growth. According to Hao et al. [25], endophytes of plants participate in the metabolic pathways of medicinal plants. The results were further compared to the leaves extracts of the plant species. The qualitative preliminary test tube analysis of the different Jatropha species were tested positive for most of the secondary metabolites, 61% of the endophytes extracts revealed the presence of more than three secondary bioactive metabolites. It was also noted that only one endophyte ISO J8, clustering with strong bootstrap support of 98% Bayesian support with Colletotrichum acutatum had all the secondary metabolites screened. Although Colletotrichum acutatum is responsible for its devastation on economically important crop plants in temperate and sub tropics [26], in the present study the fungal isolate was recovered as endophyte and the secondary metabolite recovered from the fungus bared a similar resemblance to the leaves from which it was isolated.
Antibacterial properties
The endophytic fungi from the leaves of J. curcas gave highly significant antimicrobial activities effects (p<0.01) against both the tested Gram (+ve) and Gram (-ve) clinical pathogens and endophytes. The widest spectrum of antimicrobial activity was demonstrated by these three endophytes ISO J34, ISO J73 and ISO J74 and the latter were very effective on eleven of the thirteen pathogenic microorganisms selected. Among the four active isolates, Engyodontium was the genus that showed better strengths of antimicrobial activity. The antimicrobial activities confirmed that crude ethyl acetate extracts of the endophytic fungi from the Jatropha species were more effective than the crude ethyl acetate extract of the leaves of J. curcas from which it was isolated.
Conclusion
The aim of this study was to evaluate the abundance, diversity, host affiliations, and local distributions of endophytic and saprophytic fungi associated with the four Jatropha species present in Mauritius. Often, abiotic factors (e.g. geographic distance, climate, seasonal and spatial variations, microclimates, disturbances) are described as drivers of fungal diversity in tropical environments, we can therefore conclude on the same observation in subtropical environment also. The fungi isolated as both endophytes and saprophytes demonstrated a better antimicrobial property than the leaves of their host. It was also that recorded the presence of a fungal isolate with the ability to produce oil with the same properties as its host. The results confirmed that endophytic fungi from Jatropha plants are sources of novel and antimicrobial compounds. The study paves way for a new direction for research, where endophytes can be further explored rather than their host. Hence it would be informative to further confirm whether the endophytes are more capable than their host.
Acknowledgement
The research was supported by the University of Mauritius under the Grant 0082.
References
- Strobel G, Daisy B (2003) Bioprospecting for microbial endophytes and their natural products. Microbiol Mol Biol Rev 67: 491-502
- Cho KM, Hong SY, Lee SM, Kim YH, Kahng GG, et al. (2007) Endophytic bacterial communities in ginseng and their antifungal activity against pathogens. Microb Ecol 54: 341-351.
- Jayasiri SC, Hyde KD, Ariyawansa HA, Bhat J, Buyck B, et al. (2015) The faces of fungi database: Fungal names linked with morphology, phylogeny and human impacts. Fungal Divers 74: 3-18.
- Malhadas C, Malheiro R, Pereira JA, de Pinho PG, Baptista P (2017) Antimicrobial activity of endophytic fungi from olive tree leaves. World J Microbiol Biotechnol 33: 46.
- Shankar NB, Shashikala J, Krishnamurthy YL (2008) Diversity of fungal endophytes in shrubby medicinal plants of Malnad region, Western Ghats, Southern India. Fungal Ecol 1: 89-93.
- Gao XX, Zhou H, Xu DY, Yu CH, et al. (2005) High diversity of endophytic fungi from the pharmaceutical plant, Heterosmilax japonica Kunth revealed by cultivation-independent approach. FEMS Microbiol Lett 249: 255-266.
- Keerthi D, Aswati Nair R, Prasath D (2016) Molecular phylogenetics and anti-Pythium activity of endophytes from rhizomes of wild ginger congener, Zingiber zerumbet Smith. World J Microbiol Biotechnol 32: 41.
- Nalini MS, Sunayana N, Prakash HS (2014) Endophytic fungal diversity in medicinal plants of Western Ghats, India. Int J Biodivers, pp: 1-9.
- Sussman RW, Tattersall I (1986) Distribution, abundance and putative ecological strategy of Macaca fascicularis on the Island of Mauritius, southwestern Indian Ocean. Folia Primatologica 46: 28-43.
- Rampadarath S, Puchooa D, Jeewon R (2016) Jatropha curcas L: Phytochemical, antimicrobial and larvicidal properties. Asian Pac J Trop Med 6: 858-865.
- Rampadarath S, Puchooa D, Ranghoo-Sanmukhiya VM (2014) A comparison of polyphenolic content, antioxidant activity and insecticidal properties of Jatropha species and wild Ricinus communis L. found in Mauritius. Asian Pac J Trop Med 7: S384-S390.
- Contran N, Chessa L, Lubino M, Bellavite D, Lobina R, et al. (2016) Potentialities and limits of Jatropha curcas L. as alternative energy source to traditional energy sources in northern Ghana. Energy Sustain Dev 31: 163-169.
- Koh MY, Ghazi TI (2011) A review of biodiesel production from Jatropha curcas L. oil. Renewable and Sustainable Energy Reviews 15: 2240-2251.
- Kumar S, Kaushik N (2013) Endophytic fungi isolated from oil-seed crop Jatropha curcas produces oil and exhibit antifungal activity. PLoS One 8: e56202.
- Gashgari R, Gherbawy Y, Ameen F, Alsharari S (2016) Molecular characterization and analysis of antimicrobial activity of endophytic fungi from medicinal plants in Saudi Arabia. Jundishapur J Microbiol 9: e26157.
- Qin S, Miao Q, Feng WW, Wang Y, Zhu X, et al. (2015) Biodiversity and plant growth promoting traits of culturable endophytic actinobacteria associated with Jatropha curcas L. growing in Panxi dry-hot valley soil. App Soil Ecol 93: 47-55.
- Jeewon R, Liew ECY, Hyde KD (2004) Phylogenetic evaluation of species nomenclature of Pestalotiopsis in relation to host association. Fungal Divers 17:39-55.
- Ganley RJ, Newcombe G (2006) Fungal endophytes in seeds and needles of Pinus monticola. Mycol Res 110: 318-327.
- Promputtha I, Lumyong S, Dhanasekaran V, McKenzie EH, Hyde KD (2007) A phylogenetic evaluation of whether endophytes become saprotrophs at host senescence. Microb ecol 53:579-590.
- Arnold AE, Lutzoni F (2007) Diversity and host range of foliar fungal endophytes: Are tropical leaves biodiversity hotspots? Ecol Soc Am 88.
- Rodriguez RJ, White JF Jr, Arnold AE, Redman RS (2009) Fungal endophytes: Diversity and functional roles. New Phytol 182: 314-330.
- Bonfim JA, Vasconcellos RLF, Baldesin LF, Sieber TN, Cardoso EJBN (2016) Dark septate endophytic fungi of native plants along an altitudinal gradient in the Brazilian Atlantic forest. Fungal Ecol 20: 202-210.
- Higgins KL, Coley PD, Kursar TA, Arnold AE (2011) Culturing and direct PCR suggest prevalent host generalism among diverse fungal endophytes of tropical forest grasses. Mycologia 103: 247-260.
- Giauque H, Hawkes CV (2016) Historical and current climate drive spatial and temporal patterns in fungal endophyte diversity. Fungal Ecol 20: 108-114.
- Hao J, Song F, Huang F, Yang C, Zhang Z, et al. (2007) Production of laccase by a newly isolated deuteromycete fungus Pestalotiopsis sp. and its decolorization of azo dye. J Ind Microbiol Biotechnol 34: 233-240.
- Peres NA, Timmer LW, Adaskaveg JE, Correll JC (2005) Lifestyles of Colletotrichum acutatum. Plant Dis 89: 784-796.
Citation: Rampadarath S, Puchooa D, Jeewon R, Bandhoa K (2018) Diversity, Seasonal Variation and Antibacterial Activity of Endophytic Fungi Associated With the Genus Jatropha in Mauritius. J Biotechnol Biomater 8: 280. DOI: 10.4172/2155-952X.1000280
Copyright: ©2018 Rampadarath S, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Select your language of interest to view the total content in your interested language
Share This Article
Recommended Journals
Open Access Journals
Article Tools
Article Usage
- Total views: 5943
- [From(publication date): 0-2018 - Oct 03, 2025]
- Breakdown by view type
- HTML page views: 4964
- PDF downloads: 979