Genetic Variability Study of Yield and Yield Related Traits in Rice (Oryza sativa L.) Genotypes
Received: 12-Jun-2018 / Accepted Date: 26-Jun-2018 / Published Date: 04-Jul-2018 DOI: 10.4172/2329-8863.1000381
Keywords: Genotype; Irrigated rice; Rice; Variability
Introduction
Rice is the most important food grain crop with regard to human nutrition and caloric intake. It is a staple food for more than half of the world population providing one fifth of calorie and 15% protein consumption [1,2]. In Ethiopia, the second populous nation in sub Saharan Africa, rice is the target commodity that has received a due emphasis in promotion of agricultural production. It is considered as alternative technology for farmers for efficient utilization of marginal resources [3]. Ethiopia has considerably vast suitable ecologies for rice production which is unsuitable for production of other food crops [4]. From this, the country has a potential of 3.7 million ha land for irrigated low land rice production [4]. Although the country has a high potential for production rice in irrigated areas, lack of adaptable varieties is one of the major problem. Until now only nine irrigated varieties had been released for irrigated environment [5]. The generation of adaptable varieties largely depends on the availability of desirable genetic variability for important traits [6].
Variability refers to the presence of differences among the individuals of plant population due to their genetic composition and the environment in which they are raised [7,8]. Genetic variability is the basis of plant breeding because any crop improvement depends on the amount and direction of genetic association of the traits in the base population [9,10]. It provides a wide range of genotypes that can be selected to develop new varieties [11]. Development of high yielding varieties requires the existence of genetic variability and genetic variability of agronomic traits is the key component of breeding programs for broadening the gene pool [10,12]. Selection will be effective when there is a significant amount of variability among the breeding materials [7]. The available variability can be measured using genotypic and phenotypic coefficient of variation which used to partition genetic and environmental variance [6,13]. Heritability used to assess the amount genetic improvement that will be transferred to next generation [14]. However, heritability coupled with genetic advance will be more powerful in predicting genetic gain than heritability alone [15,16]. In addition, knowledge about the association of traits with themselves and grain yield is also important for direct and indirect selection of traits which contributes to yield [10,16].
Therefore, the present study was conducted with the objective of assessing the genetic variability, heritability, genetic advance and association between yield and yield components of 64 rice genotypes.
Methodology
The study was conducted in main Season (June to November) of 2016 at Werer Agricultural Research Center (WARC) experimental field. Werer is located 9°27’ N and 40°15’ E in northeastern part of Ethiopia which is about 280 km away from Addis Ababa. The altitude of Werer is 740 m.a.s.l. Fourteen years climatic data on monthly bases showed that the average maximum and minimum temperature in the region is 34°C and 19°C, respectively, and the annual total rainfall in the area is about 571 mm annually. The soil in the region is predominantly vertisol with the porosity and bulk density (0-25 cm depth) of 49.06% and 1.35 gm/cm2, respectively [17].
The experimental design was simple lattice design consisted of 64 (8 × 8) rice genotypes sown in two replications. The seeds (100 kg/ha) were directly drilled in the field in two rows of 30 cm spacing and 3 m row length. Phosphorus fertilizer was applied at sowing in the form of DAP (109 kg/ha). Nitrogen, in the form of Urea (219 kg/ha) was applied in two splits (half at tillering and the remaining panicle initiation). Irrigation water was applied every 4 days interval using furrow irrigation until the crop reached to physiological maturity. All other agronomic management practices were applied as per recommendation. The evaluated genotypes are listed below (Table 1).
| ENT | Designation | Pedigree | Origin | ENT | Designation | Pedigree | Origin |
|---|---|---|---|---|---|---|---|
| 1 | AMOL 1 | TAICHUNG NATIVE 1/TAROM FIROZKANDEH | IRAN | 33 | WEED TOLERANCE RICE 1 | - | CHINA |
| 2 | B11338F-TB-26 | TB177E-TB-28-D-3/B10384E-MR-1-8-3//IR600-80-23///TB177E-TB-28-D-3/BIO38E-KN-362//BL245 | INDONESIA | 34 | IRRI 134 | - | IRRI |
| 3 | CB04-110 | ASD 18/AS.94140 | PAKISTAN | 35 | 08FAN6 | - | CHINA |
| 4 | MTU-1115 | BPT 5204/DP 13 | INDIA | 36 | LH1 | - | CHINA |
| 5 | AA 1 R 1 | - | INDIA | 37 | NERICA-4 (check) | CG14/WAB56 | WARDA |
| 6 | B11998-MR-1-10-TB-2-MR-1 | FATMAWATI*2/KLEMAS/////BP140F-MR-1-3/PUCUK////BP140F-MR-1*3/TUKAD UNDA | INDONESIA | 38 | IR 65 | - | IRRI |
| 7 | CR 2340-5 | NDR 9830018/IR31238-350-321//IR 41054-102-2-3-2 | INDIA | 39 | IRRI 122 | - | IRRI |
| 8 | CT 17323-1-1-2-2-2-2-M | CT 16629-22/IR25586-45-1-2//EPAGRI 108 | CIAT | 40 | KCD 1 | - | CHINA |
| 9 | CT 19558-2-17-1-2-1-1-M | CT 18228-(8)-F2//POOLBCF/FL00593-6P-1-3P-M | CIAT | 41 | ESMET101 | - | - |
| 10 | PANT DHAN 19 | BG 132/UPRI 95-14 | INDIA | 42 | ESMET126 | - | - |
| 11 | PR 27843-2B-20-P123 | PR26668-29-2/PSB RC 34 | PHILIPPINES | 43 | ESMET122 | - | - |
| 12 | WAS 173-B-B-10-6-2 | Sahel 201/4456 | SENEGAL | 44 | ESMET 115 | - | - |
| 13 | OM 4555 | IR 28/OM 2395 | VIETNAM | 45 | ESMET 120 | - | - |
| 14 | IR 62 558-SRN-17-2-1-3 | - | IRRI | 46 | ESMET102 | - | - |
| 15 | IR 82-489-568-2-1 | IR 72967-12-2-3/PR 31090-33-2-1 | IRRI | 47 | ESMET103 | - | - |
| 16 | ROKAN | IR58025 A/IR53942-69-3-1-1-1R | INDONESIA | 48 | ESMET114 | - | - |
| 17 | CT 16659-8-2CT-1-5-2-1-M | PERLA*2/ORYZA RUFIPOGON//CT 11026-3-9-1T-2P-3P-1 | CIAT | 49 | ESMET119 | - | - |
| 18 | RADAHA 7 | MASULI/JANAKI | NEPAL | 50 | ESMET107 | - | - |
| 19 | ADT(R)47 | ADT 43/JEERAGAS AMBA | INDIA | 51 | ESMET121 | - | - |
| 20 | CIBDGO | S487 B-75/2*IR196 61 -131-3-13//2*IR64 | INDONESIA | 52 | GSR IR 1-17-D6-Y1-D1 | - | IRRI |
| 21 | BAKTULSHI | - | BANGLADESH | 53 | GSR IR 1-12-Y4-D1-Y1 | - | IRRI |
| 22 | BR 7232-6-2-3 | - | BANGLADESH | 54 | GSR IR 1-5-Y4-S1-Y1 | - | IRRI |
| 23 | BRRI DHAN 48 | - | BANGLADESH | 55 | GSR IR 1-5-D7-Y3-S1 | - | IRRI |
| 24 | DARIAL | - | BANGLADESH | 56 | GSR IR 1-5-Y3-Y1-D1 | - | IRRI |
| 25 | JATTA | - | BANGLADESH | 57 | GSR IR 1-5-S14-S2-Y1 | - | IRRI |
| 26 | RJT 74 (ACC 53688) | - | INDIA | 58 | GSR IR 1-8-S6-S3-Y2 | - | IRRI |
| 27 | TAK RATIA (ACC 76415) | - | PAKISTAN | 59 | GSR IR 1-11-D7-S1-S1 | - | IRRI |
| 28 | TOOR THULLA (ACC 76420) | TOOR THULLA | PAKISTAN | 60 | GSR IR 1-15-D7-S4-S1 | - | IRRI |
| 29 | WAB96-1-1 | ITA257/YS121 | AFRICA RICE | 61 | GSR IR 1-5-S14-S2-Y2 | - | IRRI |
| 30 | IR 26 | - | IRRI | 62 | GSR IR 1-9-D12-D1-SU1 | - | IRRI |
| 31 | 926 | - | 63 | GSR IR 1-8-S12-Y2-D1 | - | IRRI | |
| 32 | IRRI 109 | - | IRRI | 64 | GSR IR 1-5-Y7-Y2-SU1 | - | IRRI |
Table 1: List of genotypes evaluated and their origin.
Data for 7 agronomic and phonological traits were taken according to the standard evaluation system for rice. Data were collected for days to emergence (DTE), days to heading (DHE), days to maturity (DMA), plant height (PLH), number of effective tillers (ET), panicle length (Pl) and grain yield (GY). Observations were recorded on the plot basis for DTE, DHE, DMA and GY. For GY the yield harvested on plot basis and adjusted to 14% moisture content before weighting. All the remaining traits were recorded on the plant basis by randomly taking 5 plants in each plot.
Analysis of Variance was conducted using SAS 9.0. Genotypic variance (σ2g), phenotypic variance (σ2p), genotypic coefficient of variation (GCV), phenotypic coefficient variation (PCV), heritability in broad sense (h2b), genetic advance (GA), genetic advance as percentage of means (GAM) were estimated [18,19]. Phenotypic correlations, principal component analysis and cluster component analysis were done using SAS 9.0.
Genotypic variance (σ2g)=(MSg-MSe)/r
Phenotypic Variance (σ2p)=σ2g+(σ2e)/r
Genotypic coefficient of variation (GCV)= 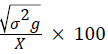
Phenotypic coefficient of variation (PCV)= 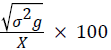
Heritability (h2b)=(σ2g/σ2p) × 100
Genetic Advance (GA)=(k) × (σp) × (h2b)
Genetic Advance as percentage of Mean (GAM)=(GA/X) × 100
Where, MSg=mean square of genotype, MSe=mean square of error, r=number of replication, σ2e=environmental variance, X=grand mean, k=intensity of selection at 5% (2.06).
Results and Discussion
Analysis of variance (ANOVA)
The analysis of variance showed there was a very highly significant difference among the genotypes evaluated for all traits except for panicle length (Table 2). Similar results were obtained in studies of Sumanth et al. [7], Oniya et al. [13], Jambhulkar and Bose [20] and Mural, Sasalawad and Hittalmani [21]. This indicates the genotypes are highly variable especially those traits which showed significant difference. Thus, the possibility genetic improvement through selection is highly promising. The study by Idris and Mohammed [12] and Bekele et al. [22] reported similar results. High variability of breeding materials will increase the probability of producing desirable recombinants in successive generations.
| TRAIT | REP (1) | BLOCK (14) | GENOTYPE (63) | ERROR (49) | CV | MEAN |
|---|---|---|---|---|---|---|
| DTE | 18.63*** | 0.28ns | 1.07*** | 0.29 | 4.73 | 11.45 |
| DHE | 1507.83*** | 2.53ns | 177.39*** | 1.73 | 1.32 | 100.19 |
| DMA | 18.86** | 13.92ns | 173.25*** | 10.02 | 2.52 | 125.69 |
| PLH | 1099.27*** | 0.74ns | 189.51*** | 0.45 | 1.09 | 61.53 |
| ET | 22.68*** | 0.01** | 0.57*** | 0.004 | 1.42 | 4.63 |
| PL | 1.2ns | 4.24ns | 2.97ns | 2.76 | 8.51 | 19.54 |
| YLD | 2417759*** | 22646.7*** | 268561.4*** | 4419.1 | 2.32 | 2886.16 |
Table 2: Analysis of variance for different traits of 64 rice genotypes. Where, DTE: days to emergence, DHE: days to heading, DMA: days to maturity, PLH: plant height, ET: number of effective tillers, PL: panicle length, YLD: grain yield, ns: non-significant, *, ** and ***: significant at 5%, 1% and 0.01% respectively.
Estimate of genetic parameters
The highest variability (genotypic and phenotypic) was exhibited in grain yield (268561.4 and 266351.9) followed by plant height (189.51 and 189.28), days to heading (177.39 and 176.52) and days to maturity (173.25 and 168.24). High variability in grain yield and plant height was also reported by Sumanth et al. [7].
According to Siva Subramanian and Menon [23] GCV and PCV more than 20% considered as high, whereas values less than 10% are considered to be low and values between 10% and 20% being considered to be moderate. According to this, most the traits have high to intermediate GCV and PCV (Table 3). This indicated that these traits could be improved for breeding high yielding rice varieties through selection and hybridization [7]. Grain yield had the highest environmental variance. However, the environmental variance in plant height (PLH) and days to heading (DHE) was very low. Similar findings were obtained by Rashid et al. [1], Pratap et al. [6] and Idris and Mohammed [12] who studied the genetic variability of different rice genotypes for yield and yield components.
| Mean | Max | Min | GV | PV | EV | GCV | PCV | H2 | GA | GAM | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| DTE | 11.45 | 13.00 | 9.00 | 0.925 | 1.07 | 0.145 | 8.39 | 9.03 | 86.44 | 1.84 | 16.08 |
| DHE | 100.19 | 118.00 | 72.00 | 176.525 | 177.39 | 0.865 | 13.26 | 13.29 | 99.51 | 27.3 | 27.25 |
| DMA | 125.69 | 145.00 | 101.00 | 168.24 | 173.25 | 5.01 | 10.31 | 10.47 | 97.1 | 26.33 | 20.95 |
| PLH | 61.53 | 87.82 | 47.25 | 189.285 | 189.51 | 0.225 | 22.35 | 22.37 | 99.88 | 28.32 | 46.03 |
| ET | 4.63 | 6.23 | 3.52 | 0.568 | 0.57 | 0.002 | 16.27 | 16.31 | 99.64 | 1.55 | 33.47 |
| YLD | 2886.16 | 5690.41 | 797.64 | 266351.9 | 268561.4 | 2209.55 | 17.88 | 17.95 | 99.17 | 1058.76 | 36.68 |
| PL | 19.54 | 22.91 | 17.38 | 1.59 | 2.97 | 1.38 | 6.45 | 8.81 | 53.53 | 1.9 | 9.726 |
Table 3: Mean, range and genetic parameters for different traits of 64 rice genotypes. Where: DTE: days to emergence, DHE: days to heading, DMA: days to maturity, PLH: plant height, ET: number of effective tillers, PL: panicle length, YLD: grain yield, Max: maximum, Min: minimum, GV, PV and EV: genotypic, phenotypic and environmental variance, GCV and PCV: Genotypic and Phenotypic coefficient of variation, H2: heritability in broad sense, GA: expected genetic advance at 5% of selection, GAM: Genetic advance as a percentage of mean.
The genetic variation result showed that phenotypic coefficient of variation (PCV) was relatively higher than genotypic efficient of variation (GCV). This result revealed that the influence of environment on phenotypic expression of each trait [12]. But the difference between genotypic and phenotypic coefficient of variation is very little for all studied trait except for days to emergence (DTE) which was relatively higher (Table 3). This shows the influence of environment in expression most of the traits is very small [9,10]. A little higher difference between PCV and GCV for days to emergence (DTE) indicates the influence of environment on DTE was relatively higher than other traits. This result was similar to studies of Rashid et al. [1], Sumanth et al. [7], Oniya et al. [13] and Tefera, Sentayehu and Leta [24].
The broad sense heritability in the studied traits ranged from 53.53% to 99.9% (Table 3).
According to Johnson, Robinson and Comstock [18] broad sense heritability classified as low (<30%), medium (30% to 60%) and high (>60%). This shows most of the traits studied can be easily improved through selection. The medium heritability in panicle length (53.53%) showed the more influence of environment on this trait. Therefore, direct selection for this trait is not effective. Since heritability do not always indicate genetic gain, heritability coupled with genetic advance is more effective for selection [18]. Genetic advance indicates the expected progress as the result of selection [6]. It used to estimate the type of gene action in polygenetic traits [18]. Genetic advance as percent of mean classified as low (<10%), moderate (10%-20%) and high (>20%) [18]. In this study, it ranges from 10% for panicle length to 112% for grain yield. A small amount of genetic advance in panicle length was also reported in the study of Rashid et al. [1]. High heritability and genetic advance were seen in most of the traits including grain yield (Table 3) which indicated these traits were less influenced by the environment, governed by additive gene action and can easily be selected through phenotypic selection [7]. Days to emergence (DTE) has high heritability and moderate genetic advance, indicating that this character is governed by both additive and nonadditive gene action. This showed there is a possibility of direct selection for this character. The remaining trait panicle length (PL) has medium heritability and low genetic advance. This shows this character is totally governed by non-additive gene action. So, heterosis breeding could be used for such kind of traits [16,24,25].
Phenotypic correlation
The phenotypic correlation results showed the presence of positive and significant correlation between yield and most traits except plant height and panicle length (Table 4). The correlation between plant height and yield is negative and highly significant. This shows traits other than plant height and panicle length can be used for direct selection. Similar results were found in the experiment of Aditya and Bhartiya [10] and Idris and Mohammed [12]. The negative correlation grain yield with plant height did not agree with the work Aditya and Bhartiya [10], Idris and Mohammed [12], Jambhulkar and Bose [20], Kar et al. [26]. Although it was not strong and significant, negative correlation between these two traits was found in the study of Pratap et al. [6] and Ogunbayo et al. [16].
| DTE | DHE | DMA | PLH | ET | PL | YLD | |
|---|---|---|---|---|---|---|---|
| DTE | 0.67*** | 0.25* | -0.71*** | 0.72*** | -0.08ns | 0.60*** | |
| DHE | 0.51*** | -0.95*** | 0.95*** | -0.05ns | 0.85*** | ||
| DMA | -0.46*** | 0.43** | 0.02ns | 0.38** | |||
| PLH | -0.98*** | 0.13ns | -0.91*** | ||||
| ET | -0.11ns | 0.89*** | |||||
| PL | -0.21ns | ||||||
| YLD |
Table 4: Phenotypic Correlation among yield and yield components of 64 rice genotypes. Where: DTE: days to emergence, DHE: days to heading, DMA: days to maturity, PLH: plant height, ET: number of effective tillers, PL: panicle length, YLD: grain yield.
Principal component analysis
Principal component analysis was carried out using 64 genotypes and 7 traits. Principal component analysis reflects the importance of larger contributor to the total variation at each axis of differentiation [27]. Eigenvalues are used to determine how many factors to retain [28]. According to Gatten's lower bound principle eigenvalue less than one should be ignored [29]. In the analysis, 10 principal components were extracted with eigen value ranged from 0.88 to 2.97 (Table 5).
| Variable | PC1 | PC2 | PC3 |
|---|---|---|---|
| DTE | 0.35 | 0.07 | -0.02 |
| DHE | 0.38 | -0.34 | 0.31 |
| DMA | 0.31 | -0.45 | 0.26 |
| PLH | -0.36 | 0.33 | 0.15 |
| ET | 0.24 | 0.48 | 0.06 |
| PL | -0.04 | -0.13 | 0.49 |
| YLD | -0.05 | 0.21 | 0.53 |
| Eigen Value | 2.97 | 2.05 | 1.46 |
| Variation (%) | 29.79 | 20.53 | 14.63 |
| Cumulative (%) | 29.79 | 50.31 | 64.94 |
Table 5: Eigenvectors and eigenvalues of the first five principal components of 7 traits. Where: DTE: days to emergence, DHE: days to heading, DMA: days to maturity, PLH: plant height, ET: number of effective tillers, PL: panicle length, YLD: grain yield, ns: nonsignificant, PC: principal component.
In this study, the first three components had eigen value greater than one. The first three principal components explained 64% of the total variation. The first principal component individually contributed for 29.79% of the total variation. The second and third components explained 20.53% and 14.63% of the variation observed in eigenvector analysis (Table 5).
According to Chalal and Gosal [30] characters with larger absolute value closer to unity with the first principal components influence clustering more than those with lower absolute value closer to zero. In the present study, the total variability in the genotypes is due to cumulative effect of a number of traits rather than the small contribution of each character. The positive and negative sign loading shows the presence of positive and negative correlation between the variable and the component [28].
In the first principal component traits such as DTE, DHE, DMA and ET contributed greater for total variation due to their high loading. PLH and ET were important traits in the second principal component which contributed 20.53% of the total variation. In the third principal component, PL and YLD were the greatest contributor for the variation. Similarly, Tuhina-khatun et al. [9] explained 74% of the total variability using the first four principal components among 43 upland rice genotypes based on 22 morphological traits. Similar results were found in the work of Worede et al. [31] and Lasalita-Zapico, Namocatcat and Cariño-Turner [32] where 86% and 82% total variation explained by the first five principal components respectively.
Generally, traits included in the first principal components especially traits with high loading scores are important separating the genotypes [9]. Principal components analysis showed high level of variability and diversity in studied rice genotypes.
Cluster analysis
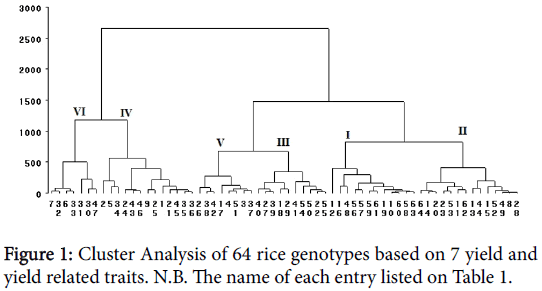
In the present study sixty-four genotypes are grouped in six clusters based of the data taken (Table 7). Among the clusters, cluster IV comprised the largest genotypes (14 genotypes) followed by cluster II with 13 genotypes and cluster I with 12 genotypes. Cluster III, V and VI consisted of 10, 8 and 7 genotypes respectively (Figure 1). Days to emergence is almost similar across all clusters but Cluster VI, III and II are the latest in terms of days to heading and maturity (Table 6). Relatively the late maturing genotypes were found in cluster IV. The longest one among the six clusters was cluster IV. Effective tiller was a bit higher in cluster II but panicle length was almost similar across the clusters. Clear difference in grain yield was observed in the six clusters. Genotypes found in cluster VI was the highest yielder (5304 kg/ha) followed by cluster IV (4189 kg/ha) and Cluster III (3278 kg/ha). Crossing between promising genotypes of cluster VI and II may provide superior desirable segregates for developing high yielding genotypes. Also considering genotypes in cluster IV will provide genotypes with better height for breeding in addition to high yield.
| Cluster | No. of Genotypes | DTE | DHE | DMA | PLH | ET | PL | YLD |
|---|---|---|---|---|---|---|---|---|
| I | 12 | 11.61 | 99.63 | 126.28 | 56.72 | 4.22 | 19.29 | 1138.90 |
| II | 14 | 11.52 | 100.43 | 127.51 | 59.45 | 5.02 | 19.25 | 1911.97 |
| III | 10 | 11.53 | 100.88 | 127.91 | 60.73 | 4.72 | 19.95 | 2645.05 |
| IV | 13 | 11.03 | 99.23 | 119.83 | 70.55 | 4.56 | 19.52 | 4189.86 |
| V | 8 | 11.77 | 97.54 | 124.54 | 60.06 | 4.77 | 19.69 | 3278.68 |
| VI | 7 | 11.30 | 104.51 | 130.08 | 60.01 | 4.43 | 19.91 | 5304.56 |
Table 6: Mean value of 6 clusters in 7 traits of 64 rice genotypes. Where: DTE: days to emergence, DHE: days to heading, DMA: days to maturity, PLH: plant height, ET: number of effective tillers, PL: panicle length, YLD: grain yield.
| Cluster | No. of Genotype | Name of genotypes |
|---|---|---|
| I | 12 | PANT DHAN 19, WAS 173-B-B-10-6-2, ROKAN, ADT(R)47, ESMET114, GSR IR 1-5-Y3-Y1-D1, GSR IR 1-5-S14-S2-Y1, GSR IR 1-8-S6-S3-Y2, GSR IR 1-11-D7-S1-S1, GSR IR 1-15-D7-S4-S1, GSR IR 1-5-S14-S2-Y2, GSR IR 1-8-S12-Y2-D1 |
| II | 14 | CT 17323-1-1-2-2-2-2-M, PR 27843-2B-20-P123, OM 4555, IR 62 558-SRN-17-2-1-3, IR 82-489-568-2-1, CIBDGO, BRRI DHAN 48, TOOR THULLA (ACC 76420), ESMET 115, ESMET119, GSR IR 1-17-D6-Y1-D1, GSR IR 1-12-Y4-D1-Y1, GSR IR 1-9-D12-D1-SU1, GSR IR 1-5-Y7-Y2-SU1 |
| III | 10 | RADAHA 7, BR 7232-6-2-3, TAK RATIA (ACC 76415), WAB96-1-1, IRRI 122, KCD 1, ESMET101, ESMET107, GSR IR 1-5-Y4-S1-Y1, GSR IR 1-5-D7-Y3-S1 |
| IV | 13 | AMOL 1, B11338F-TB-26, AA 1 R 1, CT 19558-2-17-1-2-1-1-M, BAKTULSHI, DARIAL, JATTA, IRRI 134, 08FAN6, LH1, ESMET122, ESMET 120, ESMET102 |
| V | 8 | CB04-110, MTU-1115, CT 16659-8-2CT-1-5-2-1-M, RJT 74 (ACC 53688), LOCAL CK(NERICA-4), IR 65, ESMET126, ESMET121 |
| VI | 7 | B11998-MR-1-10-TB-2-MR-1, CR 2340-5, IR 26, 926, IRRI 109, WEED TOLERANCE RICE 1, ESMET103 |
Table 7: Clustering Pattern of 64 rice genotypes based on 7 yield and yield related traits.
Conclusion
The overall result showed the presence of adequate variability in the genotypes studied. This variation could be effectively manipulated using appropriate breeding techniques and program to develop improved varieties. High estimate of heritability and genetic advance were observed in most of the traits, indicating the predominance of additive gene action and the possibility of direct selection through these traits. The phenotypic correlation and principal component analysis showed most of the traits evaluated are important for selection of high yielding genotypes and contributing their share for the wider genetic variability of the genotypes. Although some of the results did not agree with the previous findings. Therefore, it is advisable to repeat the experiment by adding more important traits in representative irrigated areas of the country, especially in irrigated low lands.
Acknowledgements
We acknowledge the national rice research program for funding the experiment. We also acknowledge cereal breeding caste team research technicians for data collection and other supports.
References
- Rashid MM, Nuruzzaman M, Hassan L, Begum SN (2017) Genetic variability analysis for various yield attributing traits in rice genotypes. Journal of the Bangladesh Agricultural University 15: 15-19.
- Sathisha TN (2013) Genetic Variation Among Traditional Landraces of Rics with Specific Reference to Nutritional Quality (Doctoral Dissertation, UAS, Dharwad).
- Gebrekidan H, Seyoum M (2006) Effects of mineral N and P fertilizers on yield and yield components of flooded lowland rice on vertisols of Fogera Plain, Ethiopia. Journal of Agriculture and Rural Development in the Tropics and Subtropics (JARTS) 107: 161-176.
- Ministry of Agriculture and Rural Development (MOARD) (2010) National Rice research and development strategy of Ethiopia. Addis Ababa: MOARD.
- Hsu YC, Tseng MC, Wu YP, Lin MY, Wei FJ, et al. (2014) Genetic factors responsible for eating and cooking qualities of rice grains in a recombinant inbred population of an inter-subspecific cross. Molecular Breeding 34: 655-673
- Pratap N, Singh PK, Shekhar R, Soni SK, Mall AK (2014) Genetic variability, character association and diversity analyses for economic traits in rice (Oryza sativa L.). SAARC Journal of Agriculture 10: 83-94.
- Sumanth V, Suresh BG, Ram BJ, Srujana G (2017) Estimation of genetic variability, heritability and genetic advance for grain yield components in rice (Oryza sativa L.). Journal of Pharmacognosy and Phytochemistry 6: 1437-1439.
- Falconer DS (1996) Introduction to quantitative genetics. New York: Longman.
- Tuhina-Khatun M, Hanafi MM, Rafii Yusop M, Wong MY, Salleh FM, et al. (2015) Genetic variation, heritability, and diversity analysis of upland rice (Oryza sativa L.) genotypes based on quantitative traits. BioMed research international.
- Aditya JP, Bhartiya A (2013) Genetic variability, correlation and path analysis for quantitative characters in rainfed upland rice of Uttarakhand Hills. Journal of Rice Research 6: 24-34.
- Pandey P, Anurag PJ, Tiwari DK, Yadav SK, Kumar B (2009) Genetic variability, diversity and association of quantitative traits with grain yield in rice (Oryza sativa L.). Journal of Bio-Science 17: 77-82.
- North S (2013) Estimation of genetic variability and correlation for grain yield components in rice (Oryza sativa L.). Global Journal of Plant Ecophysiology 3: 1-6.
- Onyia VN, Okechukwu EC, Atugwu AI, Akpan NM (2017) Genetic Variability Studies on Twelve Genotypes of Rice (Oryza sativa L.) for Growth and Yield Performance in South Eastern Nigeria. Notulae Scientia Biologicae.
- Dutta P, Dutta PN, Borua PK (2013) Morphological traits as selection indices in rice: A statistical view. Universal Journal of Agricultural Research 1: 85-96.
- Nirmaladevi G, Padmavathi G, Kota S, Babu VR (2015) Genetic variability, heritability and correlation coefficients of grain quality characters in rice (Oryza sativa L.). SABRAO J Breed Genet 47: 424-433.
- Ogunbayo SA, Ojo DK, Sanni KA, Akinwale MG, Toulou B, et al. (2014) Genetic variation and heritability of yield and related traits in promising rice genotypes (Oryza sativa L.). Journal of Plant Breeding and Crop Science 6: 153-159.
- Chekol W, Mnalku A (2012) Selected physical and chemical characteristics of soils of the middle awash irrigated farm lands, Ethiopia. Ethiopian Journal of Agricultural Sciences 22: 127-142.
- Johnson HW, Robinson HF, Comstock R (1955) Estimates of Genetic and Environmental Variability in Soybeans 1. Agronomy Journal 47: 314-318.
- Burton GW, Devane EH (1953) Estimating Heritability in Tall Fescue (Festuca Arundinacea) from Replicated Clonal Material 1. Agronomy Journal 45: 478-481.
- Jambhulkar NN, Bose LK (2014) Genetic variability and association of yield attributing traits with grain yield in upland rice. Genetika 46: 831-838.
- Mural RV, Sasalawad R, Hittalmani S (2012) Evaluation of Rice (Oryza sativa L.) Accessions for Growth and Yield Attributes in Aerobic Condition. International Journal of Plant Breeding 6: 147-149.
- Bekele BD, Naveen GK, Rakhi S, Shashidhar HE (2013) Genetic evaluation of recombinant inbred lines of rice (Oryza sativa L.) for grain zinc concentrations, yield related traits and identification of associated SSR markers. Pakistan journal of biological sciences: PJBS 16: 1714-1721.
- Sivasubramanian S, Menon M (1973) Heterosis and inbreeding depression in rice. Madras Agric J 60: 1139.
- Abebe T, Alamerew S, Tulu L (2017) Genetic variability, heritability and genetic advance for yield and its related traits in rainfed lowland rice (Oryza sativa L.) genotypes at Fogera and Pawe, Ethiopia. Adv Crop Sci Technol 5: 1-8.
- Panse VG (1957) Genetics of quantitative characters in relation to plant breeding. Indian J Genet 17: 318-328.
- Kar RK, Mishra TK, Pandey RK, Das SR (2016) Assessment of genetic variability in low land sub-1 introgressed rice genotypes showing tolerance to submergence and stagnant flooding. IOSR J Agric Vet Sci 9: 42-46.
- Sharma JR (1998) Statistical and biometrical techniques in plant breeding. New Age International Pvt Limited.
- Hailegiorgis D, Mesfin M, Gangwar SK (2010) Genetic divergence analysis on some bread wheat. Int J Sci Nat 1: 53-57.
- Kumar S, Vashisht RP, Gupta R, Singh K, Kaushal M (2011) Characterization of European carrot genotypes through principal component analysis and regression analysis. Int J Veg Sci 17: 3-12.
- Chahal GS (2002) Principles and procedures of plant breeding biotechnology and conventional approaches. New Delhi: Narosa Publishing House
- Worede F, Sreewongchai T, Phumichai C, Scripichitt P (2014) Multivariate analysis of genetic diversity among some rice genotypes using morpho-agronomic traits. J Plant Sci 9: 14-24.
- Lasalita-Zapico FC, Namocatcat JA, Cariño-Turner JL (2010) Genetic diversity analysis of traditional upland rice cultivars in Kihan, Malapatan, Sarangani Province, Philippines using morphometric markers, Philipp. J Sci 139: 177-180.
Citation: Girma BT, Kitil MA, Banje DG, Biru HM, Serbessa TB (2018) Genetic Variability Study of Yield and Yield Related Traits in Rice (Oryza sativa L.) Genotypes. Adv Crop Sci Tech 6:381. DOI: 10.4172/2329-8863.1000381
Copyright: © 2018 Girma BT, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Select your language of interest to view the total content in your interested language
Share This Article
Recommended Journals
Open Access Journals
Article Tools
Article Usage
- Total views: 7743
- [From(publication date): 0-2018 - Dec 08, 2025]
- Breakdown by view type
- HTML page views: 6467
- PDF downloads: 1276

