Research Article Open Access
Isolation and Characterization of Microorganisms that Degrade Dimethoate
Shivangi Singh and Priya R Iyer*PG and Research Department of Biotechnology, Women`s Christian College, Chennai, Tamil Nadu, India
- *Corresponding Author:
- Priya R Iyer
PG and Research Department of Biotechnology
Women`s Christian College, Chennai, Tamil Nadu, India
Tel: +91 044 2827 5926
E-mail: brajuraj@yahoo.com
Received Date: August 28, 2017; Accepted Date: September 08, 2017; Published Date: September 11, 2017
Citation: Shivangi S, Priya RI (2017) Isolation and Characterization of Microorganisms that Degrade Dimethoate. J Bioremediat Biodegrad 8:410. doi: 10.4172/2155-6199.1000410
Copyright: © 2017 Shivangi S, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Journal of Bioremediation & Biodegradation
Abstract
Microorganisms have the ability to degrade a large number of pollutants found in the environment. Microorganisms such as bacteria can degrade toxic compounds and chemicals in an environment friendly way. Dimethoate is an organophosphate pesticide. It is a broad-spectrum insecticide. In the present study, dimethoate degrading microorganisms have been isolated from various soil samples. The different isolates that were obtained were characterized by various biochemical tests. Standardization of degradation was performed. Leuco crystal violet reagent was used as an indicator and colorimetric analysis was done at 595 nm. In total, 8 isolates were got. 3 belonged to Bacillus sp. and 3 were identified as Staphylococcus aureus. 2 isolates were identified as Aspergillus sp. HPLC analysis was carried out for the organism that showed the best degradation results.
Keywords
Microorganisms; Bacteria; Degradation; Pollutants
Introduction
Organic synthetic pesticides are used in the environment as a control measure against the pests and the parasites to have a secure agricultural and industrial produce. Despite their usefulness in the increase in the yield of food production, the extensive use of pesticides during production, processing, storage, transport or marketing of agricultural commodities can lead to environmental pollution. Persistence of pesticides in the soil can vary from few hours to many years as in case of Organophosphorous (OP) pesticides. The waste generated during pesticide manufacturing is very complex, containing chemical compounds used for manufacturing and the residuals generated during manufacturing/formulation process [1]. The World Health Organization (WHO) data [2] show that only 2-3% of applied chemical pesticides are effectively used for preventing, controlling and killing pests, while the rest remains in the soil [1]. Therefore, the surface soil containing residual pesticides causes toxicity in the surrounding environment. No segment of the population is completely protected against exposure to pesticides and the potentially serious health effects [2]. Deaths and chronic diseases occurring due to pesticide poisoning are about 1 million per year globally [3]. There was evidence on the risk of cancer of workers occupationally exposed to herbicides or dioxins (since dioxins contaminated the herbicide mixtures used in Vietnam, and of the Vietnamese population [4]). In India, the first case of poisoning was reported in 1958 in Kerala where more than a hundredpeople died due to consumption of wheat contaminated with parathion [5]. India is the tenth largest consumer of pesticides, the first being China and the twelfth largest manufacturer of pesticides globally. Pesticides reach rivers and oceans as runoffs from the fields. More than 90 percent of water and fish samples from all streams contained one, or more often, several pesticides in a study done by US Geological Survey (USGS) on major river basins in the mid 90’s [6]. In a survey in India, 58% of drinking water samples drawn from various hand pumps and wells around Bhopal were contaminated with Organo Chlorine pesticides above the EPA standards [7]. Dimethoate is an organophosphorous pesticide that is used as an insecticide and an aracicide. It has been listed as toxic under the GHS (globally harmonized system of classification and labelling of chemicals) classification. It is toxic if swallowed and in contact with skin. The major objective of the following study is to isolate and characterize microorganisms degrading dimethoate from environmental samples and the standardization of degradation by the organisms. Transformation has been carried out using the plasmid of Bacillus and Staphylococcus. E. coli DH5α was used as the host culture.
Materials and Methods
Collection of sample
The samples used in the study were collected from various industrial areas of Chennai, Tamil nadu, India. The following nine samples were used in the study (Table 1).
| S.No. | Sample | Geographical Area |
|---|---|---|
| 1 | Farmland soil | Avadi, Chennai, Tamil Nadu |
| 2 | Tyre factory soil | Avadi |
| 3 | Home dust | Ashok Nagar, Chennai, Tamil Nadu |
Table 1: Sample collection data.
Pesticide used in the study
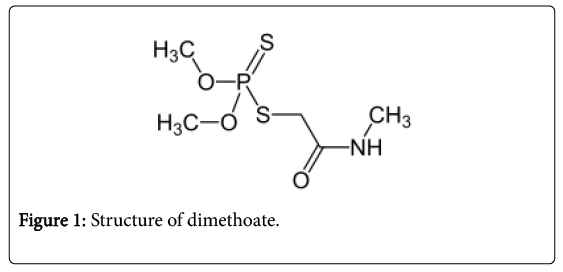
The pesticide used in the study was dimethoate. It is an organophosphorous pesticide. The pesticide was purchased from a local pesticide seller in T.nagar, Chennai (Figure 1).
Isolation of microbial consortia
Primary screening was done to isolate the microorganisms that are able to tolerate dimethoate pesticide and use it as a source of energy. For this purpose minimal salt medium was used. The samples were sprinkled in minimal salt medium broth in test tubes in different concentrations of pesticide starting from 0.5 to 4%. Growth of organisms was observed under the microscope when gram staining of broth was performed. The organisms were transferred from broth to MSM plates with corresponding pesticide concentration using spread plate method and allowed to grow for a few days. Growth was observed in plates with 0.5 to 2% pesticide concentration. Organisms growing in 2% concentration were taken for further studies.
Biochemical characterization
Biochemical tests were performed for all the isolates using Bergey’s Manuel of Systemic Bacteriology for biochemical characterization.
Standardization of degradation
Degradation tests were carried out at varying concentration, pH, temperatures and static and non-static cultures. The varying concentrations of pesticides used were 0.5%, 1%, 2%, 3% and 4%. The different pH used were 2, 4, 6 and 8. Three different temperatures were used. They were 14°C, 34°C and 45°C. Colorimetric analysis was done at 595 nm to check the amount of pesticide degraded.
Transformation studies
The plasmids were isolated from bacterial strains and they were transformed into host culture. E.coli DH5α strain was used as the host culture. For isolation of plasmid DNA alkaline lysis method was used. Competent cells were prepared using 0.1% calcium chloride solution. Blue white screening method was used for checking transformation which involved the use of ampicillin, X-Gal and IPTG.
Result
Isolation of microbial consortia
In total 6 isolates of bacteria were obtained from the 3 different environmental samples. Five isolates were gram positive rods or cocci. The bacteria got from ice cream factory effluent were gram negative cocci. All the bacterial cultures were pure cultures.
Biochemical characterization
The biochemical tests showed that four isolates were Bacills sp. while two isolates were Staphylococcus sp. (Table 2).
| Biochemical test | Farmland soil | Tyre factory soil | Home dust | Home dust | Home dust | Home dust |
|---|---|---|---|---|---|---|
| SS1 | SS2 | SS3 a | SS3 b | SS3 c | SS3 d | |
| Oxidase test | + | + | + | + | + | + |
| Catalase test | + | + | + | + | + | + |
| Citrate utilization test | + | + | + | + | + | + |
| Methyl red test | + | + | + | + | + | + |
| Vogesproskauer test | + | + | + | + | + | + |
| Triple sugar ion test | + | + | + | + | + | + |
| Urease test | - | + | - | - | - | - |
| Indole test | + | + | + | + | + | + |
| Coagulase test | - | - | - | + | + | + |
| Identification | Bacillus sp. | Bacillus sp. | Bacillus sp. | Staphylococcus aureus | Bacillus aureus | Staphylococcus aureus |
Table 2: Biochemical test results.
From tyre factory sample, 2 types of fungi were obtained. Under microscopic observation, they were both found to be Aspergillus sp.
Standardisation of degradation
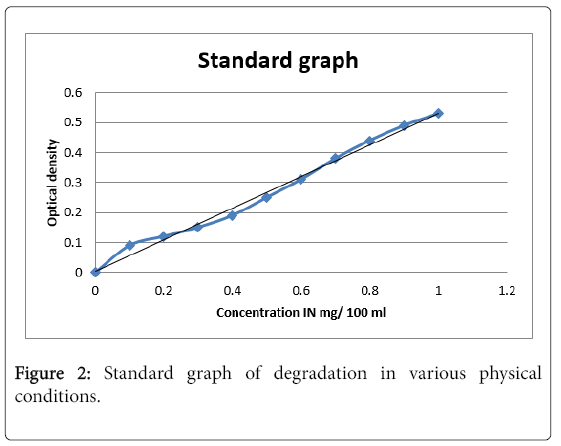
Standard graph was drawn to study degradation in various physical conditions (Figure 2). The linear regression equation of the standard graph obtained in the study was y=0.005+0.5264x.
Degradation test at varying concentrations
The OD (Optical Density) values for each of the samples were extrapolated on the standard graph in order to get the concentrations in mg/100 ml. Out of the five, organism 4 (Staphylococcus aureus) gave the best degradation results. Organism 2 (Bacillus sp.) also gave good degradation results comparable with that of organism 4 (Staphylococcus aureus). Organism 5 (Staphylococcus aureus) showed the least degradation activity. The amount of residual dimethoate after 7 days were 0, 4, 17, 28 and 29 µg/mL; the rate constants were 0.775, 0.305, 0.225, 0.167 and 0.127 each per day, and the efficiency of dimethoate degradation were 100%, 96%, 83%,72% and 71%, for Bacillus licheniformis, Pseudomonas aeruginosa, Aeromonashydrophila, Proteus mirabilis and Bacillus pumilus respectively [8].
Degradation at varying pH
At pH 2, very good results of degradation are shown by Organism 1 (Bacillus sp.), organism 2 (Bacillus sp.) and organism 3 (Staphylococcus aureus) while organisms 3 and 5 (Staphylococcus aureus) show least degradation.
At pH – 4, moderate degradation is observed in organism 1 (Bacillus sp.) and organism 5 (Staphyloccus aureus). Least degradation is observed in organism 5 (Staphylococcus aureus). The other 2 showed good degradation activity.
At pH – 6, maximum degradation is observed in organism 4 (Staphylococcus aureus) and 1(Bacillus sp). However, maximum activity is seen within the first 24 hours and then it seems to have slowed down.
At, pH – 8, organisms 1 (Bacillus sp.), 3 and 5 (Staphylococcus aureus) show good degradation.
Both Bacillus sp. and Staphylococcus aureus show better activity in an alkaline pH.
Degradation at varying temperature
Out of the five, organism 4 (Staphylococcus aureus) gave the best degradation results. All the organisms showed best degradation at 40°C.
Degradation test in static and non-static broth culture
Staphylococcus aureus was selected to check its degradation in two different conditions, shaking and non-shaking broth culture, since it gave best degradation results in varying concentrations, pH and temperature. The degradation is slightly more in shaking than in non-shaking flask.
Transformation
Plasmid isolation:The presence of plasmid was detected in bacteria from Ambattur and Vedanthangal lake water. Alkaline lysis method was used for the extraction of the plasmids. The plasmid was run on agarose gel along with ladder to see the approximate size of the plasmid. The size of the plasmids varied from 1 kb to 3 kb.
Transformation:Transformation was done using DH 5α as host culture and transformed colonies with plasmids from the two different organisms were transferred to different plates and left for incubation for 24 hours.After incubation, growth of blue and white colonies was observed.
The white colonies indicate the transformed colonies. The blue colonies indicate the untransformed colonies. The transformation mixture was plated onto LB plates with pesticide concentration of 2%. It was found that competent cells of DH 5α were successfully transformed.
High performance liquid chromatography
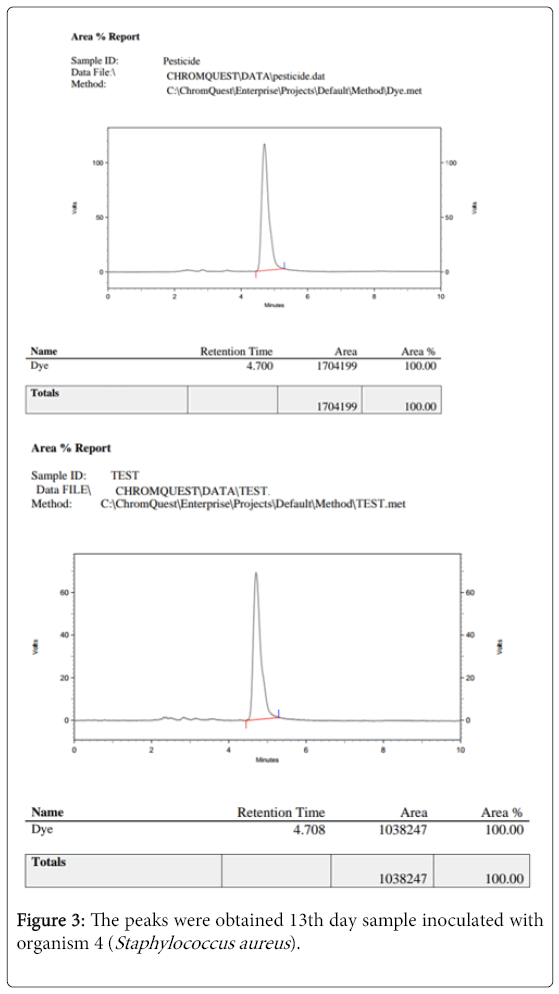
The 13th day sample inoculated with organism 4 (Staphylococcus aureus) was analysed by HPLC test (figure 3).
Discussion
Dimethoate has been labeled as hazardo 13th day sample inoculated with organism 4 (Staphylococcus aureus) us under the GHS (globally harmonized system of classification and labelling of chemicals) classification. The present study focuses on the isolation of microorganisms that degrade dimethoate from two different soil samples. The isolated microorganisms have been characterized and identified as Bacillus sp., Staphylococcus sp. and Aspergillus sp. A study on the degradation capabilities of the five different organisms has been carried out and degradation has been standardized for them. Leuco crystal violet reagent has been used to check degradation. It has been observed that the amount of pesticide in each sample is less after 312 hours when compared to 24 hour old samples after inoculation. Organism SS3 c showed the best degradation potential and HPLC (High Performance Liquid Chromatography) analysis was carried out for the sample degraded by the organism. Dimethoate adsorption was measured on eight soil types (pH from 8.0-8.45; OM from 0.73-2.95%; clay content from 5.9-14.9%) in which resulting isotherms followed an L-shape [9]. Some pesticides hydrolyze very rapidly. The hydrolysis rate can be rapid in the pH range of 8 to 9. For every pH point increase, the rate of hydrolysis will increase by approximately 10 times [10]. Within the pH range of 7–11, dimethoate degradation depends on the alkalinity of the medium rather than the time of storage. At pH 4.2 degradation is largely dependent on the length of time the dimethoate solution has been stored [11]. The half-life of dimethoate in clay loam soil was determined to be 10 and 5 days at 10°C and 20°C, respectively. This confirmed a faster decrease in the pesticide due to the increased temperature [9]. HPLC analysis was carried out to check degradation. Bioremediation was carries out on a small scale using varying combinations of the organisms that were isolated in the study. When all the organisms were present, degradation was maximum. The overall aim is to achieve a natural method to eliminate dimethoate from the environment in an environment friendly manner.
Conclusion
The present study suggests that the isolated organisms are able to utilize dimethoate as energy source since no other nutrients were available or were present in limited quantity. Hence, they degrade dimethoate. In the study, Bacillus sp., Staphylococcus aureus and Aspergillus sp. are found to flourish well in presence of dimethoate.
References
- U.S. Environmental Protection Agency and California Department of Pesticide Regulation (2005) Pesticide Product Databases.
- World Health Organization (1990) Public health impact of pesticides used in agriculture.
- Environews Forum (1999) Killer environment. Environ Health Perspect 107: A62-A63.
- Frumkin H (2003) Agent Orange and cancer: an overview for clinicians. CA Cancer J Clin 53: 245-255.
- Karunakaran CO (1958) The Kerala food poisoning. Journal of the Indian Medical Association 31: 204-207.
- Kole RK, Banerjee H, Bhattacharyya A (2001) Monitoring of market fish samples for Endosulfan and Hexachlorocyclohexane residues in and around Calcutta. Bull EnvirronContamToxicol 67: 554-559.
- Kole RK, Bagchi MM (1995) Pesticide residues in the aquatic environment and their possible ecological hazards. J Inland Fish Soc India 27: 79-89.
- Manisha DM, Shyamapada M, Nishith KP (2011) Kinetics of dimethoate degradation in bacterial system. Microbiology Research 2: 2.
- Van Scoy A, Pennell A, Zhang X (2016) Environmental fate and toxicology of dimethoate. In: Reviews of Environmental Contamination and Toxicology, Springer International Publishing, New York, USA, 237: 53-70.
- Deer HM, Beard R (2001) Effect of water pH on the chemical stability of pesticides. AG/Pesticides 14: 1.
- El Beit IO, Wheelock JV, Cotton DE (1978) Factors influencing the degradation of dimethoate in soils and solutions. International Journal of Environmental Studies 11: 253-260.
Relevant Topics
- Anaerobic Biodegradation
- Biodegradable Balloons
- Biodegradable Confetti
- Biodegradable Diapers
- Biodegradable Plastics
- Biodegradable Sunscreen
- Biodegradation
- Bioremediation Bacteria
- Bioremediation Oil Spills
- Bioremediation Plants
- Bioremediation Products
- Ex Situ Bioremediation
- Heavy Metal Bioremediation
- In Situ Bioremediation
- Mycoremediation
- Non Biodegradable
- Phytoremediation
- Sewage Water Treatment
- Soil Bioremediation
- Types of Upwelling
- Waste Degredation
- Xenobiotics
Recommended Journals
Article Tools
Article Usage
- Total views: 6201
- [From(publication date):
September-2017 - Aug 29, 2025] - Breakdown by view type
- HTML page views : 4973
- PDF downloads : 1228