Research Article Open Access
New Insights into Microbes in the Midgut of Termite Coptotermes formosanus
| Tingting Li, Jijiao Zeng, Deepak Singh and Shulin Chen* | |
| Department of Biological Systems Engineering, Washington State University, Pullman WA, 99164, USA | |
| Corresponding Author : | Shulin Chen Department of Biological Systems Engineering Washington State University, Pullman WA 99164-6120, USA Tel: 509-335-3743 Fax: 509-335-2722 E-mail: chens@wsu.edu |
| Received February 21, 2014; Accepted April 23, 2014; Published April 28, 2014 | |
| Citation: Li T, Zeng J, Singh D, Chen S (2014) New Insights into Microbes in the Midgut of Termite Coptotermes formosanus. J Bioremed Biodeg 5:220. doi:10.4172/2155-6199.1000220 | |
| Copyright: © 2014 Li T, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. | |
Related article at Pubmed Pubmed  Scholar Google Scholar Google |
|
Visit for more related articles at Journal of Bioremediation & Biodegradation
Abstract
Wood-feeding termites have evolved unique capability to effectively digest lignocellulosic material, using it for both energy and nutrition. This ability depends mainly on the mutualistic interaction between symbiotic gut microbiota and the termite itself. This study investigated microorganisms in the midgut of termite Coptotermes formosanus, a segment that has been less studied than the hindgut. Fluorescence in situ hybridization (FISH) was initially used to visualize and identify individual bacteria and archaea in the termite’s midgut. After isolation of microorganisms with six different media, preliminary screening was carried out on plates by testing the capability to oxidize guaiacol as well as decolorize the dye azure B. Two selected strains; B207 and L201 were identified as Streptomyces sp. through 16S rRNA gene sequence analysis. Submerged state fermentation of the strains with softwood biomass as substrate was further performed. The analysis results of attenuated total reflectance fourier transform infrared (ATR-FTIR), chromatography/mass spectrometry (GC/MS) and pyrolysis-gas chromatography/ mass spectrometry (Py-GC/MS) indicated that streptomyces strains B207 and L201 have certain lignocellulose decomposition capabilities.
Biological processes have been recognized as one of the alternative pretreatment technologies. It is well known that natural microorganisms such as white [11,12] and brown-rot fungi [13], secrete complex enzyme systems to degrade lignocelluloses under ambient conditions. For example, Singh et al. [14] found that P. chrysosporium expressed lignin peroxidase and manganese peroxidase at an early growth stage during their growth on wheat straw. The fungus-treated wheat straw had higher S/G ratios [14] and less energy demand for thermal degradation [15]. Another major advantage of biological process is that the enzyme system has selectivity for lignin removal. Understanding of this biological process would provide critical insights into development of new technology for biomass pretreatment and hydrolysis.
Another group of extremely effective wood-degrading organisms is termites. Termites can degrade 65-99% of wood-cellulose and hemicelluloses, as well as 5-83% of the lignin within 24 hours under natural conditions [16-19]. Apparently, the termites have unique facilities to handle lignin-carbohydrate complexes efficiently. A variety of lignocellulolytic enzymes were found in the termite system [20-22]. Despite their small body size, termites harbor an abundant and astonishingly diverse intestinal microbiota, which is one of the most fascinating examples of symbiosis among microbes, and between an animal and microbes [23]. It is widely accepted that lignocellulose digestion in termites is intimately correlated with both host and a highly specific flora of symbiotic microbes [2,5,24-26]. Much work has been performed to understand the exact roles of the host and symbiotic microbiota in the complex processes of lignocellulose degradation and conversion. Warnecke et al. [27] confirmed important symbiotic functions in termite hindgut through metagenomic and functional analysis on carbohydrate hydrolysis, H2 metabolism, CO2-reductive acetogenesis and N2 fixation [27]. Through termite digestome analysis, Tartar et al. (2009) found that in the gut of the lower termite Reticulitermesflavipes, there existed an apparent three-way collaboration among termite, protist and prokaryotic symbionts involving several cellulase, hemicellulase, phenoloxidase and peroxidase genes for lignin degradation/depolymerization [2,28].
Various evidences suggest that lignin structure was modified in termites and that the modification process begins in the foregut and continues in the midgut [2,17,27,29]. However, the source of the enzymes or chemicals responsible for this modification still need further research. It is believed that the enzymes from the host largely contribute to the digestive process of lignin. Coy et al. [30] provided evidence that lactases in the gut of Reticulitermesflavipes were produced in the salivary gland, secreted into the foregut and capable of modifying soluble lignin [30]. Compared to the hindgut segment, few reports have addressed the composition and function of the microbes present before the hindgut, except for the mixed segment, present only in higher termites, where it is situated between the midgut and the first proctodeal segment [31]. The lower termite gut is even considered absent occurring of intestinal microbiota in the segments before the hindgut. This lack of information prevents a comprehensive understanding of the symbiosis between gut bacteria and their termite host regarding lignin degradation.
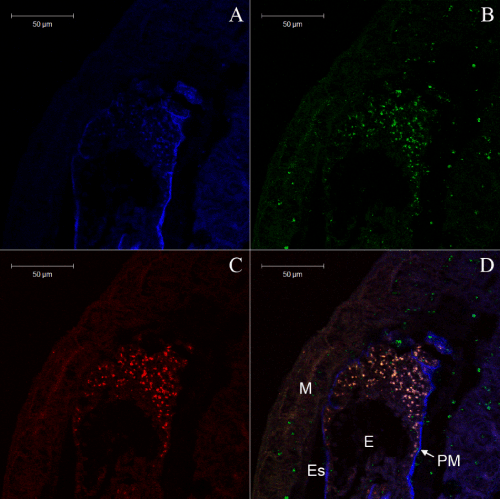
We report in this paper a study using culture and culture-independent approaches on the midgut microbe ecosystem. Fluorescence in situ hybridization (FISH) technique in association with confocal laser scanning microscopy (CLSM) was used to define the microbial communities and display the distribution and diversity of microbial flora in the termite midgut. Two isolates from the termite midgut were identified as the Streptomyces genus by 16S rRNA and named B207 and L201. Their functionality on softwood was determined by Fourier transform infrared spectroscopy (ATR-FTIR), gas chromatography/mass spectrometry (GC/MS) and pyrolysis-gas chromatography/mass spectrometry (Py-GC/MS). The results of this study indicated that the existing microbial community in the termite midgut could have important roles in lignocellulose decomposition.
Symbiotic bacteria associated with the ectoperitrophic space and epithelium in the midgut of insects is considered uncommon, not only because of the protective ability of the peritrophic membrane, but also due to the enzymes/chemicals present in this region that could reduce their ability to survive. These microorganisms might be responsible for supplementing nutrients for these insects, producing amino acids from nitrates and carbohydrates [50]. The presence of microorganisms in the midgut, especially in the ectoperitrophic space and midgut epithelium, suggests their participation in food digestion as symbiotic organisms, representing a new localizing strategy and a new possibility for exploiting food sources found in the environment. However, how bacteria bridge the peritrophic membrane is still not clear.
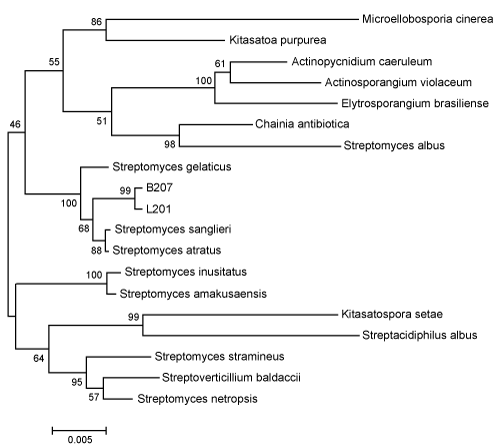
Based on their 16S rDNA gene sequences, the strains B207 and L201 had the highest similarities to streptomyces sanglieri (99.378%) and streptomyces atratus (99.378%). The phylogenetic relationship of strains B207 and L201 to other organisms included in family Streptomycetaceae suggested that strains B207 and L201 were members of the genus streptomyces (Figure 3). Previously, streptomycetes have been isolated from the guts of termites and other wood feeding insects, which provides further evidence that these microbes could be an integral part of the lignocellulosic digestion in the wood feeding termite [51-53].
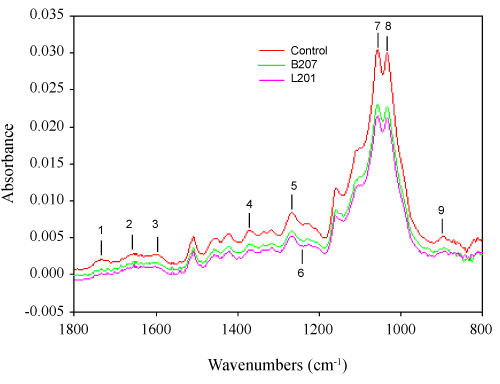
Additionally, we have determined the small fragment accumulation in acidified culture supernatants. Ethyl acetate extracted compounds were analyzed through GC/MS. Table 5 shows the patterns of ethyl acetate extracted compounds from the culture supernatants. Both inoculated and uninoculated control supernatants contained significant amounts of these compounds, but with different fragment patterns. In the control, almost all the fragments were derived from carbohydrates compounds, such as lactic acid, succinic acid and pyroglutamic acid, but were almost totally absent in the inoculated supernatants due to utilization as carbon source by streptomyces. In contrast, oxalic acid, propanedioic acid, and glutaric acid increased 10 fold or appeared as principal components in the inoculated samples. Oxalic acid is secreted by wood-degrading fungi, and it is thought to depolymerize cellulose and hemicellulose through non-enzymatic mechanisms. As such it has been proven to be a strong catalyst for hemicellulose hydrolysis [60], which is consistent with our FTIR results and component analysis. Furthermore, lignin-derived compounds were also found in the culture supernatants. The principal substituted phenolic acids that would be expected to arise from streptomyces attack on hydroxyphenyl- and guaiacyl propane units in lignin include 2-hydroxyphenylacetic acid, 4-hydroxybenzoic acid, vanillic acid, 4-hydroxyphenylacetic acid, 2-hydroxy-3-(4-hydroxyphenyl) propanoic acid, and ferulic acid. Substituted benzoic acids 4-hydroxybenzoic acid and vanillic acid have been reported as lignin degradation intermediates in fungi [61]. The accumulation of benzoic acid and vanillic acid is derived from the cleavage of Ca and Cß of the aliphatic side chain in the G unit [62]. The appearance of ferulic acid, found to be linked at different positions on the arabinose residues in the arabinoxylans through ester bonds [63], indicated hydrolysis of these ester bonds, which are considered to be critical to hemicellulose degradation; the hydrolysis of which was also supported by the FTIR results. The presence of low-molecular-weight aromatics in the controls was probably due to leaching over the extended incubation period. Overall, Py-GC/MS and GC/MS revealed that the strains B207 and L207, isolated from the midgut of termites, deconstructed softwood through degradation of hemicellulose and conversion of the condensed G unit into soluble lignin derivatives.
References
- Arakawa G, Watanabe H, Yamasaki H, Maekawa H, Tokuda G (2009) Purification and molecular cloning of xylanases from the wood-feeding termite, Coptotermes formosanusShiraki. BiosciBiotechnolBiochem73:710-718.
- Scharf ME, Tartar A (2008) Termite digestomes as sources for novel lignocellulases. Biofuels, Bioproducts and Biorefining2:540-552.
- Yang B, Wyman CE (2004) Effect of xylan and lignin removal by batch and flowthrough pretreatment on the enzymatic digestibility of corn stover cellulose. BiotechnolBioeng86:88-98.
- Ohgren K, Bura R, Saddler J, Zacchi G (2007) Effect of hemicellulose and lignin removal on enzymatic hydrolysis of steam pretreated corn stover. BioresourTechnol98:2503-2510.
- Breznak JA, Brune A (1994) Role of Microorganisms in the Digestion of Lignocellulose by Termites. Ann Rev Entomol39:453-487.
- Wong D (2009) Structure and Action Mechanism of Ligninolytic Enzymes. ApplBiochemBiotechnol157:174-209.
- Anderson W, Akin D (2008) Structural and chemical properties of grass lignocelluloses related to conversion for biofuels. J IndMicrobiolBiotechnol35:355-366.
- Harmsen P, Huijgen W, Bermudez L, Bakker R (2010) Literature review of physical and chemical pretreatment processes for lignocellulosic biomass. Wageningen: Wageningen UR, Food & Biobased Research.
- Hendriks ATWM, Zeeman G (2009) Pretreatments to enhance the digestibility of lignocellulosic biomass. BioresourTechnol100:10-18.
- Taherzadeh MJ, Karimi K (2008) Pretreatment of lignocellulosic wastes to improve ethanol and biogas production: a review. Int J MolSci9:1621-1651.
- Hatakka AI (1983) Pretreatment of wheat straw by white-rot fungi for enzymicsaccharification of cellulose. ApplMicrobiolBiotechnol18:350-357.
- Lee JW, Gwak KS, Park JY, Park MJ, Choi DH, et al. (2007) Biological pretreatment of softwood Pinusdensiflora by three white rot fungi. J Microbiol45:485-491.
- Dey S, Maiti TK, Bhattacharyya BC (1994) Production of some extracellular enzymes by a lignin peroxidase-producing brown rot fungus, Polyporusostreiformis, and its comparative abilities for lignin degradation and dye decolorization. Appl Environ Microbiol60:4216-4218.
- Singh D, Zeng J, Laskar DD, Deobald L, Hiscox WC, et al. (2011) Investigation of wheat straw biodegradation by Phanerochaetechrysosporium. Biomass and Bioenergy 35:1030-1040.
- Zeng J, Singh D, Chen S (2011) Biological pretreatment of wheat straw by Phanerochaetechrysosporium supplemented with inorganic salts. BioresourTechnol102:3206-3214.
- Brune A (2007) Microbiology: woodworker's digest. Nature450:487-488.
- Ke J, Sun JZ, Nguyen HD, Singh D, Lee KC, et al. (2010) In-situ oxygen profiling and lignin modification in guts of wood-feeding termites. Insect Science17:277-290.
- Ohkuma M (2003) Termite symbiotic systems: efficient bio-recycling of lignocellulose. ApplMicrobiolBiotechnol61:1-9.
- Chaffron S, von Mering C (2007) Termites in the woodwork. Genome Biol8:229.
- Chandrasekharaiah M, Thulasi A, Bagath M, Kumar DP, Santosh SS, et al. (2011) Molecular cloning, expression and characterization of a novel feruloyl esterase enzyme from the symbionts of termite(Coptotermes formosanus) gut. BMB Rep44:52-57.
- Lo N, Tokuda G, Watanabe H (2011) Evolution and Function of Endogenous Termite Cellulases: Biology of Termites: a Modern Synthesis.(Bignell DE, Roisin Y, Lo N edn), Springer, Netherlands, 51-67.
- Zhang D, Lax AR, Bland JM, Allen AB (2011) Characterization of a new endogenous endo-[beta]-,4-glucanase of Formosan subterranean termite (Coptotermes formosanus). Insect BiochemMolBiol41:211-218.
- Hongoh Y, Ohkuma M, Kudo T (2003) Molecular analysis of bacterial microbiota in the gut of the termite Reticulitermessperatus (Isoptera; Rhinotermitidae). FEMS MicrobiolEcol44:231-242.
- Watanabe H, Nakamura M, Tokuda G, Yamaoka I, Scrivener AM, et al. (1997) Site of secretion and properties of endogenous endo-beta-,4-glucanase components from Reticulitermessperatus (Kolbe), a Japanese subterranean termite. Insect BiochemMolBiol27:305-313.
- Yang H, Peng JX, Liu KY, Hong HZ (2006) Diversity and function of symbiotic microbes in the gut of lower termites. Wei Sheng Wu XueBao46:496-499.
- Kudo T (2009) Termite-microbe symbiotic system and its efficient degradation of lignocellulose. BiosciBiotechnolBiochem73:2561-2567.
- Warnecke F, Luginbuhl P, Ivanova N, Ghassemian M, Richardson TH, et al. (2007) Metagenomic and functional analysis of hindgut microbiota of a wood-feeding higher termite. Nature450:560-565.
- Tartar A, Wheeler MM, Zhou X, Coy MR, Boucias DG, et al. (2009) Parallel metatranscriptome analyses of host and symbiont gene expression in the gut of the termite Reticulitermesflavipes. Biotechnol Biofuels 2:25.
- Geib SM, Filley TR, Hatcher PG, Hoover K, Carlson JE, et al. (2008) Lignin degradation in wood-feeding insects. ProcNatlAcadSci105:12932.
- Coy MR, Salem TZ, Denton JS, Kovaleva ES, Liu Z, et al. (2010) Phenol-oxidizing laccases from the termite gut. Insect BiochemMolBiol40:723-732.
- Tokuda G, Yamaoka I, Noda H (2000) Localization of Symbiotic Clostridia in the Mixed Segment of the Termite Nasutitermestakasagoensis (Shiraki). Appl Environ Microbiol66:2199-2207.
- Kikuchi Y, Meng XY, Fukatsu T (2005) Gut symbiotic bacteria of the genus Burkholderia in the broad-headed bugs Riptortusclavatus and Leptocorisachinensis (Heteroptera: Alydidae). ApplEnviron Microbiol71:4035.
- Yanagita K, Manome A, Meng XY, Hanada S, Kanagawa T, et al. (2003) Flow cytometric sorting, phylogenetic analysis and in situ detection of Oscillospiraguillermondii, a large, morphologically conspicuous but uncultured ruminal bacterium. Int J SystEvolMicrobiol 53: 1609-1614.
- Cho MJ, Kim YH, Shin K, Kim YK, Kim YS, et al. Symbiotic adaptation of bacteria in the gut of Reticulitermessperatus: Low endo-[beta]-, 4-glucanase activity. BiochemBiophysRes Commun 395: 432-435.
- Okino L, Machado K, Fabris C, Bononi V (2000) Ligninolytic activity of tropical rainforest basidiomycetes. World J MicrobiolBiotechnol16:889-893.
- Archibald FS (1992) A new assay for lignin-type peroxidases employing the dye azure B. Appl Environ Microbiol58:3110-3116.
- Arora DS, Gill PK (2001) Comparison of two assay procedures for lignin peroxidase. Enzyme MicrobTechnol28:602-605.
- Morisaki K, Fushimi T, Kaneko S, Kusakabe I, Kobayashi H (2001) Screening for phenoloxidases from edible mushrooms. BiosciBiotechnolBiochem65:2334-2336.
- Mercer DK, Iqbal M, Miller P, McCarthy AJ (1996) Screening actinomycetes for extracellular peroxidase activity. Appl Environ Microbiol62:2186-2190.
- Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, et al. (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389-3402.
- ASTM (1996) Standard Test Method for Ethanol-Toluene Solubility of Wood.
- Zimbardi F, Viggiano D, Nanna F, Demichele M, Cuna D, et al. (1999) Steam explosion of straw in batch and continuous systems. ApplBiochemBiotechnol77:117-125.
- LabbeN, Rials TG, Kelley SS, Cheng ZM, Kim JY, et al. (2005) FT-IR imaging and pyrolysis-molecular beam mass spectrometry: new tools to investigate wood tissues. Wood SciTechnol39:61-76.
- Oh SY, Yoo DI, Shin Y, Kim HC, Kim HY, et al. (2005) Crystalline structure analysis of cellulose treated with sodium hydroxide and carbon dioxide by means of X-ray diffraction and FTIR spectroscopy. Carbohydr Res340:2376-2391.
- Pandey KK, Pitman AJ (2003) FTIR studies of the changes in wood chemistry following decay by brown-rot and white-rot fungi. IntBiodeterBiodegr52:151-160.
- Terra WR, Ferreira C (1994) Insect digestive enzymes: properties, compartmentalization and function. Kidlington, ROYAUME-UNI: Elsevier.
- Terra WR (2001) The origin and functions of the insect peritrophic membrane and peritrophic gel. Arch Insect BiochemPhysiol47:47-61.
- Lehane MJ, Billingsley P (1996) Biology of the insect midgut. Chapman & Hall.
- Breznak JA, Pankratz HS (1977) In situ morphology of the gut microbiota of wood-eating termites [Reticulitermesflavipes (Kollar) and Coptotermes formosanusShiraki]. Appl Environ Microbiol33:406-426.
- Caetano FH, Bution ML, Zara FJ (2009) First report of endocytobionts in the digestive tract of ponerine ants. Micron40:194-197.
- Pasti MB, Pometto AL III, Nuti MP, Crawford DL (1990) Lignin-solubilizing ability of actinomycetes isolated from termite (Termitidae) gut. Appl Environ Microbiol56:2213-2218.
- Harazono K, Yamashita N, Shinzato N, Watanabe Y, Fukatsu T, et al. (2003) Isolation and characterization of aromatics-degrading microorganisms from the gut of the lower termite Coptotermes formosanus. BiosciBiotechnolBiochem67:889-892.
- Watanabe Y, Shinzato N, Fukatsu T (2003) Isolation of actinomycetes from termites' guts. BiosciBiotechnolBiochem67:1797-1801.
- Ohgren K, Bura R, Saddler J, Zacchi G (2007) Effect of hemicellulose and lignin removal on enzymatic hydrolysis of steam pretreated corn stover. BioresourTechnol98:2503-2510.
- Pérez J, Muñoz D, de la R, Martínez J (2002) Biodegradation and biological treatments of cellulose, hemicellulose and lignin: an overview. IntMicrobiol5:53-63.
- Ramachandra M, Crawford DL, Hertel G (1988) Characterization of an extracellular lignin peroxidase of the lignocellulolyticactinomycete Streptomyces viridosporus. Appl Environ Microbiol54:3057-3063.
- Hatakka A (2005) Biodegradation of Lignin: Wiley-VCH Verlag GmbH & Co. KGaA.
- Crawford DL,Pometto AL III, Crawford RL (1983) Lignin Degradation by Streptomyces viridosporus: Isolation and Characterization of a New Polymeric Lignin Degradation Intermediate. Appl Environ Microbiol45:898-904.
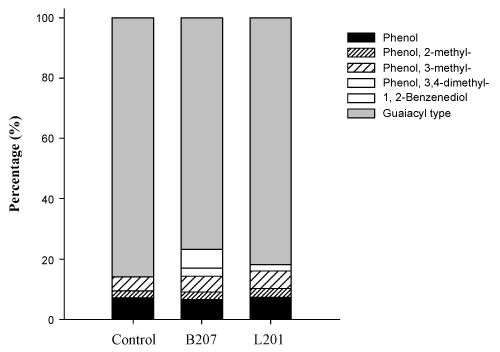
- Camarero S, Bocchini P, Galletti GC, Martínez AT (1999) Pyrolysis-gas chromatography/Mass spectrometry analysis of phenolic and etherified units in natural and industrial lignins. Rapid Commun Mass Sp13:630-636.
- Lee JW, Rodrigues RCLB, Kim HJ, Choi I-G, Jeffries TW (2010) The roles of xylan and lignin in oxalic acid pretreated corncob during separate enzymatic hydrolysis and ethanol fermentation. BioresourTechnol101:4379-4385.
- Kirk TK, Higuchi T, Chang H (1980) Program US-JCS Lignin biodegradation: microbiology, chemistry, and potential applications: CRC Press.
- Hammel KE, Cullen D (2008) Role of fungal peroxidases in biological ligninolysis. CurrOpin Plant Biol11:349-355.
- Hua D, Ma C, Song L, Lin S, Zhang Z, et al. (2007) Enhanced vanillin production from ferulic acid using adsorbent resin. ApplMicrobiolBiotechnol74:783-790.
--
Tables and Figures at a glance
| Table 1 | Table 2 | Table 3 | Table 4 | Table 5 |
Figures at a glance
 |
 |
 |
 |
 |
| Figure 1 | Figure 2 | Figure 3 | Figure 4 | Figure 5 |
Relevant Topics
- Anaerobic Biodegradation
- Biodegradable Balloons
- Biodegradable Confetti
- Biodegradable Diapers
- Biodegradable Plastics
- Biodegradable Sunscreen
- Biodegradation
- Bioremediation Bacteria
- Bioremediation Oil Spills
- Bioremediation Plants
- Bioremediation Products
- Ex Situ Bioremediation
- Heavy Metal Bioremediation
- In Situ Bioremediation
- Mycoremediation
- Non Biodegradable
- Phytoremediation
- Sewage Water Treatment
- Soil Bioremediation
- Types of Upwelling
- Waste Degredation
- Xenobiotics
Recommended Journals
Article Tools
Article Usage
- Total views: 15490
- [From(publication date):
June-2014 - Aug 31, 2025] - Breakdown by view type
- HTML page views : 10676
- PDF downloads : 4814
