Research Article Open Access
QCM-D Monitoring of Binding-Induced Conformational Change of Calmodulin
Hyun J Kwon* and Brian T Dodge
Department of Engineering and Computer Science, Andrews University, Michigan, USA
- Corresponding Author:
- Kwon HJ
Department of Engineering and Computer Science
HYH 312, Andrews University
Berrien Springs, MI 49104, USA
Tel: 269-471-3890
E-mail: hkwon@andrews.edu
Received Date: July 21, 2015; Accepted Date: September 22, 2015; Published Date: September 24, 2015
Citation: Kwon HJ, Dodge BT (2015) QCM-D Monitoring of Binding-Induced Conformational Change of Calmodulin. Biosens J 4:126. doi:10.4172/2090-4967.1000126
Copyright: © 2015 Kwon HJ, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Biosensors Journal
Abstract
Understanding conformational changes are important when studying a protein such as calmodulin (CaM), which activates various target enzymes and regulates numerous physiological functions. CaM is a highly flexible protein that can transitorily adopt various conformations. A quartz crystal microbalance with dissipation (QCM-D) sensor was used to study binding-induced conformational changes of surface-immobilized CaM. Structural changes of CaM were evaluated using the Voigt’s viscoelastic model with frequency (ΔF) and dissipation change (ΔD). When Apo-CaM layer was incubated in 0.1 mM Ca2+ solution, the layer decreased by approximately 0.56 nm, due to the release of coupled water molecules and conformational change. The application of CaM itself also caused a significantly more compact layer, supporting previous findings that CaM dimerization forms a collapsed structure that exposes a hydrophobic tunnel. The binding characteristics of CaM with peptides derived from proteins in a signal transduction pathway also demonstrated diverse biophysical properties of the CaM complexes. Each peptide showed a unique ΔF/ΔD pattern indicating versatility of CaM configuration to favorably adjust to each target molecule. The study demonstrates that the QCM-D sensor is capable of simultaneously studying binding affinity and plasticity of protein configuration for target binding. The CaM data obtained on hydrated protein layer thickness is complementary to configuration measurements of a single CaM molecule.
Keywords
QCM-D sensor; CaM; Conformation change; Biosensor; XIP; Olfactory
Introduction
Calmodulin (CaM) is a calcium (Ca2+)-binding protein that can bind to and regulate a multitude of different target proteins [1]. CaM consists of 148 amino acid residues and belongs to a class of ubiquitous proteins with similar structure characterized by their distinctive helixloop- helix Ca2+ -binding motif, the so-called EF hand. Like many other Ca2+ -binding proteins, CaM undergoes a conformational change upon binding to Ca2+, which enables it to bind to specific proteins for a specific response [2,3]. This feature is common among the calcium binding protein superfamily. The structure of the Ca2+/CaM complex has been resolved by X-ray crystallography, EIS-MS, 2D-IR [4] and NMR studies [1,5]. CaM is shown to exhibit a dumbbell shape composed of two globular domains linked by a long flexible central helix [6]. CaM exhibits a great diversity of conformational changes when it binds to various binding moieties. The solution structure of a CaM with a 26 residue synthetic peptide comprising of skeletal muscle myosin light chain kinase (skMLCK) has been identified by multidimensional NMR [5]. A recent study reports AFM also could detect conformational change of immobilized CaM from apo-CaM to Ca2+ saturated holo- CaM [7]. Single-molecule force spectroscopy by AFM enabled direct observation of CaM interaction with peptides from skMLCK and CaM-dependent kinase (CaMKK) [8-10]. By applying mechanical force, the result revealed different degrees of cooperativity for CAMbinding peptide sequences. They also suggested that target peptides to only partially Ca2+-saturated CaM may play a crucial role for finetuning the intracellular response to Ca2+ signals [8].
Our focus is to present a simple method to identify and characterize binding-induced conformational change of surface immobilized proteins using a QCM-D sensor. CaM was used because it is a well-studied molecule for conformation change when binding. To demonstrate the versatile dynamics of binding characteristics, it would be ideal to choose peptides from a series of proteins that work in concert within a single cell. As a model system, peptides were derived from proteins present in a single olfactory sensory neuron. CaM plays a pivotal role in olfactory signal transduction cascade by regulating a number of channels, enzymes and receptors, such as olfactory cyclic nucleotide-gated (CNG) channels, ryanodine receptors (RYRs), plasma membrane calcium-ATPase (PMCA), and 3’, 5’-cluclic nucleotide phosphodiesterase (PDE), plasma membrane sodium-calcium exchanger (NCX), etc. [11-13]. These proteins are all well known for interacting with Ca2+/CaM for regulating specific activities [11]. Tentative CaM binding sequences of peptides within those proteins were selected by in silico scan and also from literature [12]. Among them, XIP (exchanger Inhibitory Peptide) region of sodium calcium exchanger (NCX) is a well-known peptide, which we will call peptide X for simplicity. NCX is known to bind to Ca2+/CaM [14] to regulate its ion transport functionality and the peptide X inhibits its own NCX activity when applied in the cytosol [15]. Studying interaction between CaM and peptides that are involved in the single signal transduction cascade would serve as an example to demonstrate versatility of CaM configuration in target binding. Understanding characteristics of binding between Ca2+/CaM and peptides is critical in study of chemosensory signal transduction pathways.
The quartz crystal microbalance with dissipation (QCM-D) technique is widely used to monitor surface and interface processes [16]. The QCM-D sensor measures the frequency change (ΔF) of oscillating quartz crystal disk at this resonant frequency for adsorbed mass. If the adsorbed film is thin and rigid, the mass uptake can be linearly correlated to the frequency change by the Sauerbrey equation.
Δm=-C/n ΔF
where C is the mass-sensitivity constant (C=17.7 ng Hz-1 cm-2 for a 5 MHz resonance frequency), n is the overtone number, Δm (ng/ cm2) is the adsorbed mass per unit area, and ΔF(Hz) is the frequency shift. However, water is usually coupled with the soft and viscoelastic biomolecule. The QCM-D enables measurement of damping of oscillation or dissipation (ΔD) by periodically on-and-off voltage simultaneously. The dissipation change provides assessment of flexibility/viscoelasticity of the adsorbed mass and structural property [17]. For viscoelastic layer of protein complexes, we employed the Voigt’s viscoelastic model. In the Voigt’s model, both ΔF and ΔD information and multiple overtone measurement were used to estimate thickness of the total mass of the film including both protein and coupled water [18]. Simultaneous measurement of ΔF and ΔD of the QCM-D sensor provides information on structural characteristics of biomolecular interactions. Therefore, the QCM-D technique has been used to detect conformation change of polymer chains [19] and biological interactions [20].
In this study, we utilized a QCM-D sensor to characterize interaction between CaM and its various binding moieties including Ca2+ ions, CaM-binding proteins and peptides. A considerable body of evidence indicates that CaM can assume multiple modes of interactions with targets. This paper demonstrates that bindinginduced conformational changes of surface immobilized CaM can be observed using the QCM-D technique.
Equipment and materials
The measurements were made with a QCM-D sensor (Biolin Scientific Holding AB, Sweden) apparatus. An AT-cut quartz crystal with a fundamental resonant frequency of 5 MHz was from Biolin Scientific Holding AB (Q-sense AB). 16-Mercaptohexadecanoic acid (16 MHA), 11-Hydroxy-1-undecanethiol (11-HUT), (1-Mercapto- 11-undecyl) hexa (ethylene glycol) (MUHEG), and biotin-terminated tri (ethylene glycol) hexadecanethiol (BAT) was purchased from Asemblon, Inc. Bovine calmodulin, biotin-calmodulin, and calcineurin were purchased from EMD Chemicals, Inc. Peptide XIP was provided by Dr. Margolis (University of Maryland School of Medicine) and all other synthetic peptides were generated from Biosynthesis, Inc. Peptide sequences are listed in Table 1. All other chemicals were purchased from Sigma-Aldrich, Inc.
| Peptide name | Origin | Sequence | Dissociation constant, Kd (ΔM) |
|---|---|---|---|
| Peptide P | Phosphodiesterase 1c (PDE1c) | KSIVHAVQAGIFVERMYRRT | 6 ±1 |
| Peptide X | XIP of Na/Ca exchanger | RRLLFYKYVYKRYRAGKQRGM | 0.02 ± 0.01 |
| Peptide C | Cyclic nucleotide gated channel beta 1 (CNGB1 ) | LQELVKLFKERTEKVKEKLI | 2 ± 1 |
| Peptide R | Ryanodine receptor 1(RYR1 ) | SRYGLLIKAFSMTAAETARRTREFR | 9 ± 2 |
Table 1: The origins, sequences, and dissociation constants of the peptides used for binding to Ca2+/CaM.
Sensor surface preparation
For QCM disks, the gold surface was cleaned for 10 minutes with ultraviolet-ozone (UVO) cleaning, immersed for 5 minutes in 5:1:1 miliQ water: 16% ammonium hydroxide: 30% hydrogen peroxide solution at 75°C and then thoroughly rinsed in miliQ water. The QCM disks were then dried with a N2 stream and were used for the next surface preparation steps. In an effort to reduce experimental errors and to ensure consistency, only freshly prepared QCM disks were used for all experiments.
To prepare biotinylated QCM surface, clean QCM disks were immersed in a self-assembled monolayer (SAM) solution overnight. The SAM solution consisted of a 90:10 mixture of (1-Mercapto-11- undecyl) hexa (ethylene glycol) (MUHEG) and biotin-terminated tri (ethylene glycol) hexadecanethiol (BAT). After rinsing with 200 proof ethanol and drying with a N2 stream, the biotin-SAM prepared QCMs were ready and kept in a room temperature until use.
To prepare the QCM surface for amino coupling, clean QCM disks were immersed overnight in mixtures of 95:5, 90:10, 75:25, and 50:50 of 1 mM 16-Mercaptohexadecanoic acid (MHA) and 11-Hydroxy-1- undecanethiol (HUT). The disks were rinsed with 200 proof ethanol and dried with a N2 stream. For binding experiments, CaM was immobilized with 0.2 M EDC/NHS crosslinking agents, followed by ethanol amine (1M, pH=8.3) blocking.
QCM-D Measurement
The QCM-D sensor allows simultaneous measurement of several harmonics at fundamental frequencies of 15, 25, and 35 MHz, corresponding to the overtones of n=3, 5, and 7 respectively. The frequency and dissipation change presented in this study were based on overtone of n=3. All the measurements were collected in a precisely temperature (23.0 ± 0.01°C) controlled environment. The flow rate was set as 100 μl/min [21]. The measurement noise in measuring the frequency and dissipation were ±0.02 Hz and ±0.01×10-6, respectively. All the experiments were conducted after obtaining a stable baseline for at least 10 minutes. The baseline part was truncated in the figures to focus on the binding events.
CaM binding experiments were carried out as follows: (1) establish a stable baseline in a zero Ca2+ HEPES running buffer (in mM: NaCl 150, HEPES 10, EGTA 1 pH 7.4) for at least 10 minutes; (2) apply 1mM neutravidin solution to biotin-SAM prepared QCM surface until the frequency change reaches -45~50 Hz and wash with the running buffer; (3) apply biotinylated CaM solution for frequency change of -6~7 Hz and wash with the running buffer. After these steps, CaM was immobilized on the sensor surface via biotin-neutravidin linkages. Finally, various samples of CaM binding moieties were applied. For Ca2+ saturation experiments, HEPES buffer with 0.1 mM Ca2+ (in mM: NaCl 150, CaCl2 0.1, HEPES 10, pH 7.4) was used.
SPR experiments
The SPR measurement was made with a BIACORE 3000 (Biacore AB, Uppsala, Sweden) system. The CM5 chip was activated with a 1:1 mixture of 0.2 M NHS and EDC in water. Synthetic peptides (1~5 μM in 10 mM sodium acetate pH 5.0) were then immobilized on the surface, resulting in the SPR signals of ~700 RU. Unreacted sites were blocked with 1 M ethanolamine (pH 8.5). For a negative control, a control flow cell was simultaneously activated using NHS/EDC solution and blocked without any peptides. CaM was applied at various concentrations of 0, 15.6, 31.25,..., 1000 nM in a 0.1mM Ca2+ HEPES buffer under a continuous flow of 5 μl/min.
Data analysis
All QCM-D data were modeled using a viscoelastic Voigt model using QTools 3 software (Biolin Scientific AB, Sweden). Three overtones (n=3, 5, 7) were used for the modeling and the protein film was modeled as a homogenous single layer [17]. The dissociation constant (Kd) for binding experiments was calculated with nonlinear regression of one site specific binding of saturation data using the Prism version 5 (GraphPad Software, Inc., LaJolla, CA). Kd value from Biacore sensor were evaluated using Biacorevalution version 4.1 software (Biacore) which used a 1:1 Langmuir binding model. All the data are representatives of three separate experiments and are presented with a mean ± S.D.
Results and Discussion
Comparison of CaM deposition methods
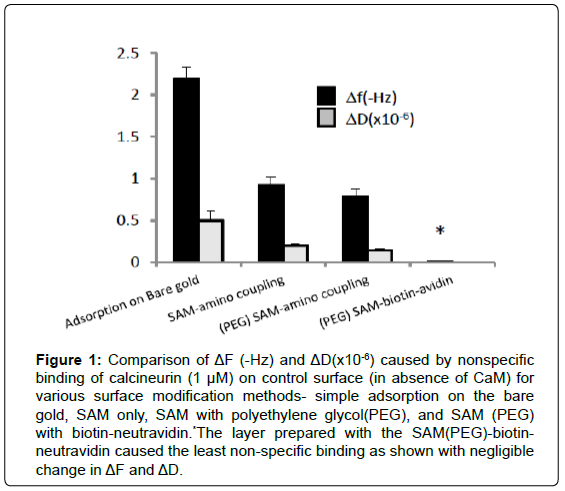
The QCM sensor surface was modified by several methods including direct adsorption, self-assembled monolayers (SAM) for amino coupling, and SAM with neutravidin-avidin capture. All these methods are widely used; however, the need of stable and specific baseline is particularly important for the low signal-to-noise measurements. Gradual increase of baseline frequency (Hz) over time (10 minutes), which occurred due to leaching of immobilized proteins, was used as a measure of baseline stability. Greater change in the baseline indicated instability of the immobilized protein layer. Nonspecific binding was identified by applying calcineurin (CN) sample to the control cell that was prepared without CaM. Although the QCM-D sensor intrinsically reduces nonspecific binding due to its mechanical oscillation and the vacant sites were blocked with ethanol amine, significant non-specific adsorption occurred with direct adsorption and amino coupling methods (Figure 1). On the other hand, the neutravidin-biotin capture on the biotin-functionalized thiols (mixture of 10% of BAT and 90% of MUHEG) provided the best platform for CaM immobilization. In this setting, biotinylated proteins were immobilized on the Neutravidin, which was bound to biotin-functionalized SAM at 10% of the total surface area to avoid steric hindrance and ensure controlled orientation of proteins. In addition, hydrophilic poly ethylene glycol (PEG) chains within the SAM were used to deter non-specific adsorption of proteins. The biotin-neutravidin capture method provided the most stable baseline with less than 0.1 Hz change in 10 minutes and minimal non-specific binding as shown in Figure 1. This result assures that the binding signals of subsequent experiments are from a very stable baseline and specific binding only.
Figure 1: Comparison of ΔF (-Hz) and ΔD(x10-6) caused by nonspecific binding of calcineurin (1 μM) on control surface (in absence of CaM) for various surface modification methods- simple adsorption on the bare gold, SAM only, SAM with polyethylene glycol(PEG), and SAM (PEG) with biotin-neutravidin.*The layer prepared with the SAM(PEG)-biotinneutravidin caused the least non-specific binding as shown with negligible change in ΔF and ΔD.
CaM conformation change due to Ca2+ binding
To study binding-induced structural change of CaM, we monitored Ca2+ induced modification of CaM structure with the QCM-D sensor. Since CaM is immobilized onto the neutravidin-biotin complex and SAM, it is important to test whether these layers undergo any alteration to the Ca2+ concentration change. The thickness of neutravidin decreased by 0.18 ± 0.013 nm when saturating Ca2+ buffer was applied. This change was reflected in the CaM layer thickness calculation for subsequent experiments.
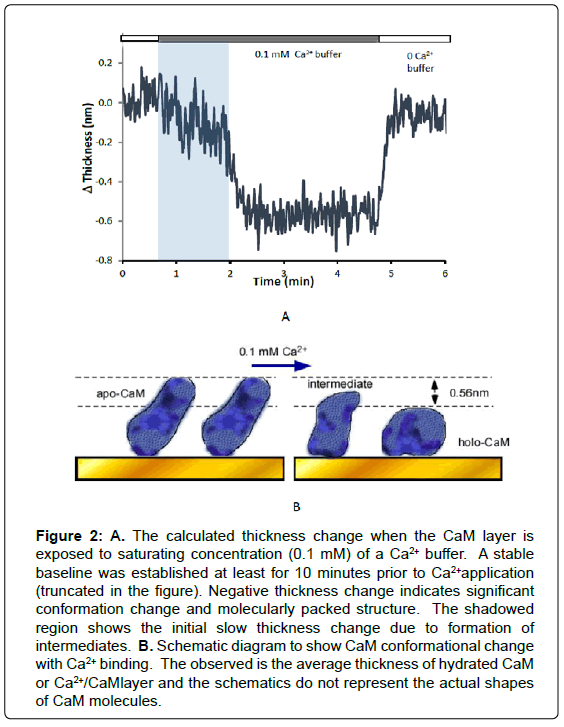
When the apo-CaM layer was exposed to 0.1 mM Ca2+, increased frequency and decreased dissipation were observed. This means both thickness of immobilized CaM layer was and flexibility were decreased. The viscoelastic film thickness of Ca2+/CaM layer was estimated to decrease by 0.56 ± 0.12 nm (n=9) according to the Voigt model (Figure 2). Re-application of zero Ca2+ buffer (quenched with EGTA) reversed this phenomenon, recovering the Apo-CaM layer thickness (Figure 2A). It is well known that CaM undergoes a pronounced conformational change that exposes a hydrophobic pocket and forms so called “bilobed” Ca2+/CaM structure [22]. The reduced thickness in the Ca2+/ CaM layer can be explained that, a large hydrophobic surface that is buried in apo-CaM becomes exposed to the solvent and thus a part of coupled water is released [3].
Figure 2:A. The calculated thickness change when the CaM layer is exposed to saturating concentration (0.1 mM) of a Ca2+ buffer. A stable baseline was established at least for 10 minutes prior to Ca2+application (truncated in the figure). Negative thickness change indicates significant conformation change and molecularly packed structure. The shadowed region shows the initial slow thickness change due to formation of intermediates. B. Schematic diagram to show CaM conformational change with Ca2+ binding. The observed is the average thickness of hydrated CaM or Ca2+/CaMlayer and the schematics do not represent the actual shapes of CaM molecules.
It is notable that transition from apo-CaM to holo-CaM appears as two stages, a slower change followed by a steep thickness change as indicated in Figure 2. This first stage consists of average 17 % of entire thickness changes (n=10) during saturation of apo-CaM with Ca2+. The observed “two stage” transition can be attributed to transition from the structure at low level of Ca2+ to that of saturating level of Ca2+ [23]. As suggested in the single molecule force study by AFM, partially saturated CaM is favorable for certain binding moieties [8]. The slow change shows that a distinctive configurational change occurs at the lower level of Ca2+, and CaM fully folds to become holo-CaM at the saturating Ca2+ concentration. However, in the reverse experiment where EGTA added zero Ca2+ solution was reapplied to the Ca2+ saturated CaM layer, the reverse transition from holo-CaM to apo-CaM did not show any distinctive transition to intermediate forms.
The QCM-D observation of the CaM structure change for Ca2+ binding is coherent with the NMR observation. In the NMR study, a comparison of the structures of the apo- and Ca2+ saturated CaM showed that Ca2+ binding causes major rearrangements of the secondary structure elements with changes in inter-residue distances and exposure of the hydrophobic interior of the four-helix bundle [24]. In the recent AFM study on structure of a surface immobilized single CaM, application of Ca2+ caused thickness change of 0.39 ± 0.13 nm from CaM (1.87 ± 0.19 nm) to Apo-CaM (2.26 ± 0.21 nm) [7].
It should be noted that the thickness change information obtained from this study was averaged value on hydrated and surface immobilized ensemble of CaM molecules before and after Ca2+ application. The data provided valuable information on the conformational change with a relatively easy and quick experimental setting. The result suggests that the QCM-D study can be a complementary method to study conformation change of Ca2+ binding proteins.
CaM binding characteristics with protein partners
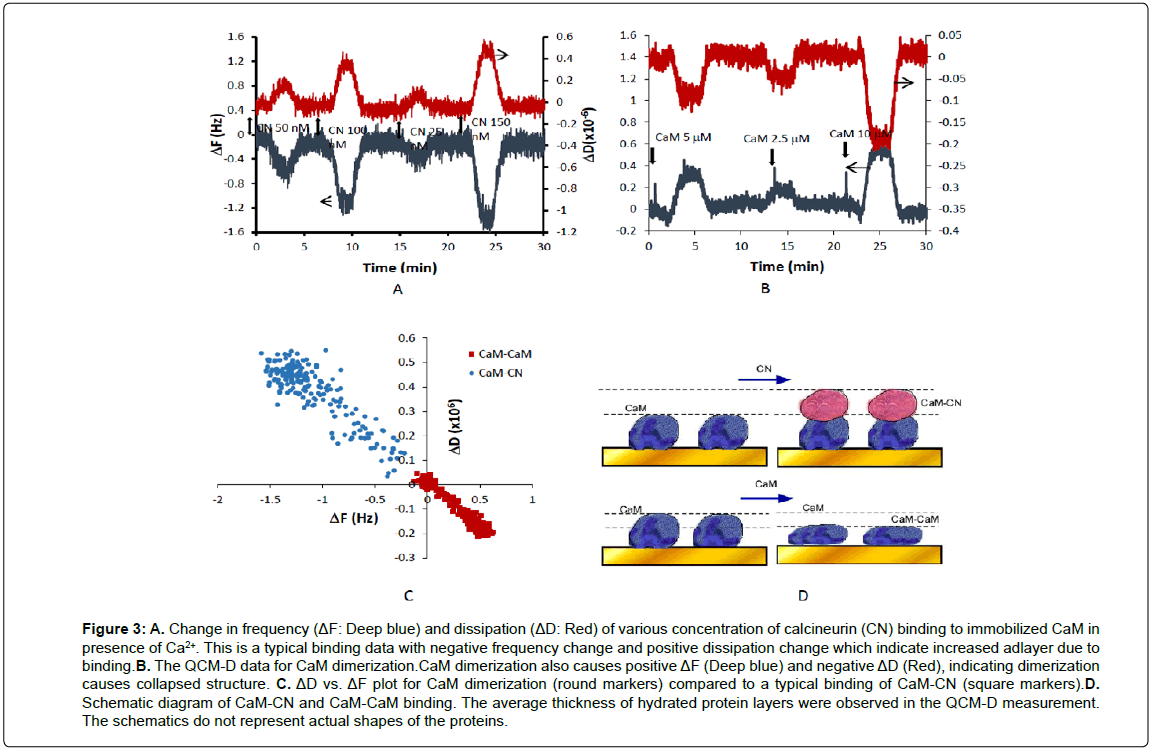
A well-known protein binding partner, calcineurin (CN, ~20 kDa) which has a similar size to the CaM was applied in presence of Ca2+ on the CaM immobilized surface. Adsorption of calcineurin resulted in negative ΔF and positive ΔD, which was interpreted as added thickness and increased flexibility of the layer. This is a stereotypical binding for majority of protein binding events that do not involve significant conformational change. Using the QCM-D sensor, CN was detected at the level of 50 nM with the sample volume of 200 μL (Figure 3A). The affinity constants were estimated to be Kd=8.8 ±1.1 nM using the equilibrium signals, which are consistent with the previously reported values [25]. This experiment was performed to demonstrate CaM binding that does not cause significant conformational change. Most of protein binding causes increase in thickness and flexibility due to higher water content and less close-packed nature of these films. When this experiment was repeated in absence of Ca2+, the signal was insignificant confirming Ca2+ dependence of CN binding to CaM [26].
Figure 3: A. Change in frequency (ΔF: Deep blue) and dissipation (ΔD: Red) of various concentration of calcineurin (CN) binding to immobilized CaM in presence of Ca2+. This is a typical binding data with negative frequency change and positive dissipation change which indicate increased adlayer due to binding.B. The QCM-D data for CaM dimerization.CaM dimerization also causes positive ΔF (Deep blue) and negative ΔD (Red), indicating dimerization causes collapsed structure. C. ΔD vs. ΔF plot for CaM dimerization (round markers) compared to a typical binding of CaM-CN (square markers).D. Schematic diagram of CaM-CN and CaM-CaM binding. The average thickness of hydrated protein layers were observed in the QCM-D measurement. The schematics do not represent actual shapes of the proteins.
CaM dimerizes readily and this phenomenon plays an important role in their regulation of target enzymes and channels [27]. In order to test whether the dimerization causes any conformational change, various concentration of CaM was applied to the sensor surface where biotinylated-CaM was immobilized. The interaction caused the thickness of the film decreases (positive ΔF and negative ΔD). Binding of equal size CaM onto CaM itself did not cause increase of film but rather lead to reduce the film thickness. The binding is characterized by fast association and fast dissociation kinetics, supporting the previous findings that CaM dimerizes through non-covalent binding such as electrostatic interactions within dimers [27]. The binding affinity was observed to be Kd=25 ±1.2 nM based on the equilibrium data of thickness, which is consistent with reported data [27]. The dimerization was observed regardless of presence of Ca2+ ions in the solution.
Combination of ΔD and ΔF data can provide insights into understanding the differences in dynamic binding behaviors of CaM and its binding partners [17]. If a protein binding causes extended conformation, it would be positioned in the left and upper quadrant in the ΔD vs. ΔF plot. On the other hand, if the binding causes more compact structure, it would be positioned in the right and lower quadrant. In Figure 3C, ΔD and ΔF plots of CaM-CN and CaM dimers were compared. The plot for CaM-CN showed a typical binding characteristics of negative ΔF and positive ΔD due to the augmented layer thickness, increased flexibility, loose binding, and higher degree of hydration in the layer. In contrast, CaM dimerization showed positive ΔF and negative ΔD. This indicates formation of well-structured complex with a high binding strength and less water entrapment due to dehydration (Figure 3D). This suggestion is supported by a number of reports that CaM dimerization is formed between isolated calcium binding loops of EF hand proteins [27] and creates a hydrophobic tunnel, forming a collapsed structure.
Ca2+/CaM binding to various peptides
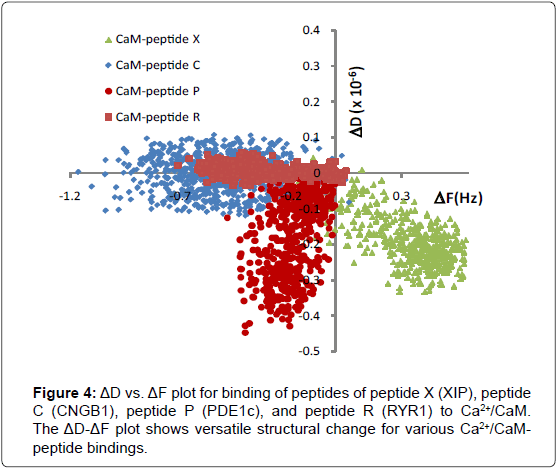
To determine versatile adaptation of a CaM structure, characteristics of binding between various synthetic peptides and CaM was studied. Synthetic peptide sequences were selected from CaM binding proteins in a sensory neuronal cell (see Table 1 for the peptide sequences) and each peptide was applied to surface immobilized Ca2+/ CaM layer separately. Since peptides are significantly smaller (10-13% in amino acid length) than CaM, the observed thickness change of the film is mostly attributed to that of a CaM layer. Combinational ΔD and ΔF plot captured the distinct viscoelastic properties of peptides and Ca2+/CaM binding (Figure 4). The dissociation constant of each binding is summarized in Table 1. Among all the peptides we tested, Peptide X binding to Ca2+/CaM resulted in the most dramatic change of a CaM structure (positive ΔF and negative ΔD). In ΔD-ΔF plot, CaM binding to Peptide X is positioned in the right and bottom quadrant as shown in Figure 4. This indicates that dehydrated, structured, and more compact CaM layer was formed when Peptide X was bound to CaM. This is again a salient example of observing conformational change where CaM conformation changes to wrap around the Peptide X and the hydrophobic chains releases entrapped water, resulting in smaller mass and energy loss.
Binding experiments of CaM and Peptide X was also performed in Biacore setting to confirm the binding affinity. In the Biacore setting, CaM was immobilized via amino coupling on a CM5 chip and Peptide X was applied on the sensor surface [14]. The binding affinity was estimated to be around ~20 nM confirming the QCM-D data. However, structural information of the binding was not obtained in the Biacore setting. This result confirms that QCM-D enabled accurate affinity measurement while providing additional information on structural rearrangement.
Peptide C- Ca2+/CaM binding caused negative ΔF due to increased thickness, however, little change in dissipation, indicating that the overall flexibility of the layer decreased. Interaction with Peptide R also showed the same non-classical conformation change of negative ΔF and neutral ΔD [13]. These can be interpreted as the binding forms a dehydrated and packed layer and the degree of dehydration is less extensive than Peptide X- Ca2+/CaM binding. Peptide P- Ca2+/CaM binding caused both negative frequency and dissipation change. This indicates that the binding leads to increased thickness and decreased flexibility forming a rigid ad layer.
In all data, ΔD/ΔF is constant over different concentration range (0-20 μM of peptides) which indicates that the viscoelastic properties of peptides-CaM were independent of the concentration range tested in these experiments. Taken together, the ΔD-ΔF plot shows dynamic structural change upon the binding of peptides to the CaM. All the peptides caused more compact structure but at different degrees. This result supports the versatility of CaM molecule which reacts with multitudes of proteins to regulate various cellular functions.
Conclusions
The QCM-D allows measurement of changes in viscoelastic, watercoupled protein layer thickness and viscoelasticity. Most of binding increases the film thickness, however, when protein folding exposes more hydrophobic region for binding, the entrapped water molecules are released leading to decreased thickness change. This makes the QCM-D sensor a great tool to quickly assess conformational change for various target bindings for CaM or other Ca2+ binding proteins. The QCM-D measurement provides averaged film thickness and viscoelasticity information on ensemble of hydrated protein molecules. These data can be complementary to structural information obtained from single molecular measurements [8,10]. The QCM-D measurement also allows binding affinity screening for unknown binding targets and simultaneously collecting conformational change in an easy and quick experimental setting. In this study, the QCM-D measurement provided experimental verification on ability of CaM to achieve a large range of diverse conformations at three levels of binding moieties –Ca2+ ions, peptides and proteins. Peptides were exclusively selected from an olfactory sensory neuron to demonstrate diversity and versatility of target binding even in type of neuronal cell.
We suggest that QCM-D measurement is a simple yet effective way of revealing diverse conformational changes to gain better insights into structural plasticity and target binding of Ca2+ binding proteins.
Acknowledgment
This work was supported by the faculty research grant (FRG; Kwon) from Andrews University. Authors thank Dr. Margolis (University of Maryland School of Medicine) for supply of some of peptides used for this study.
References
- Milos M, Comte M, Schaer JJ, Cox JA (1989) Evidence for four capital and six auxiliary cation-binding sites on calmodulin: Divalent cation interactions monitored by direct binding and microcalorimetry. J InorgBiochem 36: 11-25.
- Ikura M (1996) Calcium binding and conformational response in ef-hand proteins. Trends in Biochemical Sciences 21: 14-17.
- Wriggers W, Mehler E, Pitici F, Weinstein H, Schulten K (1998) Structure and dynamics of calmodulin in solution. Biophysical Journal 74: 1622-1639.
- Sasakura D, Nunomura W, Takakuwa Y (2012) Dynamic secondary structural changes in ca(2+)-saturated calmodulin upon interaction with the antagonist, w-7. BiochemBiophys Res Commun 423: 360-365.
- Ikura M, Clore GM, Gronenborn AM, Zhu G, Klee CB, et al. (1992) Solution structure of a calmodulin-target peptide complex by multidimensional NMR. Science 256: 632-638.
- Falke JJ, Drake SK, Hazard AL, Peersen O (1994) Molecular tuning of ion binding to calciumsignaling proteins. Q Rev Biophys 3: 219-290.
- Trajkovic S, Zhang X, Daunert S, Cai Y (2011) Atomic force microscopy study of the conformational change in immobilized calmodulin. Langmuir 27: 10793-10799.
- Junker JP, Rief M (2009) Single-molecule force spectroscopy distinguishes target binding modes of calmodulin. PNAS 106: 14361-14366.
- Best RB, Hummer G (2009) Unfolding the secrets of calmodulin. Science 323: 593-594.
- Junker JF, Ziegler R, Rief M (2009) Ligand-dependent equilibrium fluctuations of single calmodulin molecules. Science 323: 633-637.
- Ronnett GV, Moon C (2002) G proteins and olfactory signal transduction. Annu Rev Physiol 64: 189-222.
- Trudeau MC, Zagotta WN (2002) Mechanism of calciumcalmodulin inhibition of rod cyclic nucleotide-gated channels. PNAS 99: 8424-8429.
- Elshorst B, Hennig M, Försterling H, Diener A, Maurer M, et al. (1999) Nmr solution structure of a complex of calmodulin with a binding peptide of the ca2+-pump. Biochemistry 38: 12320-12332.
- Kwon HJ, Koo JH, Leinders-zufall T, Zufall F, Margolis FL (2009) Ca extrusion by ncx is compromised in olfactory sensory neurons of OMP mice. PlosOne 4: e4260.
- Blaustein MP, Charpentier TH, Weber DJ (2007) Getting a grip on calcium regulation. PNAS 104: 18349-18350.
- Chen H, Su X, Neoh KG, Choe WS (2006) Qcm-d analysis of binding mechanism of phage particles displaying a constrained heptapeptide with specific affinity to sio2 and tio2. Anal Chem 78: 4872-4879.
- Höök F, Kasemo B, Nylander T, Fant C, Sott K, et al. (2001) Variations in coupled water, viscoelastic properties, and film thickness of a mefp-1 protein film during adsorption and cross-linking:A quartz crystal microbalance with dissipation monitoring, ellipsometry, and surface plasmon resonance study. Anal. Chem. 73: 5796-5804.
- Voinova MV, Rodahl M, Johnson M, Kasemo B (1999) Viscoelastic acoustic reponse of layerd polymer films at fluid-solid interfaces: Continuum mechanics approach. PhysicaScripta 59: 391-396.
- Berglin M, Pinori E, Sellborn A, Andersson M, Hulander M, et al. (2009) Fibrinogen adsorption and conformational change on model polymers: Novel aspects of mutual molecular rearrangement. Langmuir 25: 5602-5608.
- Shur O, Wu J, Cropek DM, Banta S (2011) Monitoring the conformational changes of an intrinsically disordered peptide using a quartz crystal microbalance. Protein Sci 20: 925-930.
- Kwon HJ, Bradfield CK, Dodge BT, Agoki GS (2009) Study of fluid and mass adsorption model in the qcm-d sensor for characterization of biomolecular interaction. Proceedings for COMSOL conference: 6561.
- Bridge MJ (2008) Sensors and effectors. In: Cell signalling biology. Portland Press Limited, London.
- Houdusse A, M MS, Cohen C (1996) A model of ca(2+)-free calmodulin binding to unconventional myosins reveals how calmodulin acts as a regulatory switch. Structure 4: 1475-1490.
- Finn BE, Evenäs J, Drakenberg T, Waltho JP, Thulin E, et al. (1995) Calcium-induced structural changes and domain autonomy in calmodulin. Nat StructBiol 2: 777-783.
- Bornhop DJ, Latham JC, Kussrow A, Markov DA, Jones RD, et al.(2007) Free-solution, label-free molecular interactions studied by back-scattering interferometry. Science 317: 1732-1736.
- Quintana AR, Dan Wang, Forbes JE, Waxham MN (2005) Kinetics of calmodulin binding to calcineurin. Biochemical and Biophysical research communications 334: 674-680.
- Lafitte D, Heck AJ, Hill TJ, Jumel K, Harding SE, et al.(1999) Evidence of noncovalent dimerization of calmodulin. Eur J Biochem 261: 337-344.
Relevant Topics
- Amperometric Biosensors
- Biomedical Sensor
- Bioreceptors
- Biosensors Application
- Biosensors Companies and Market Analysis
- Biotransducer
- Chemical Sensors
- Colorimetric Biosensors
- DNA Biosensors
- Electrochemical Biosensors
- Glucose Biosensors
- Graphene Biosensors
- Imaging Sensors
- Microbial Biosensors
- Nucleic Acid Interactions
- Optical Biosensor
- Piezo Electric Sensor
- Potentiometric Biosensors
- Surface Attachment of the Biological Elements
- Surface Plasmon Resonance
- Transducers
Recommended Journals
Article Tools
Article Usage
- Total views: 16071
- [From(publication date):
specialissue-2015 - Aug 29, 2025] - Breakdown by view type
- HTML page views : 15036
- PDF downloads : 1035