Research Article Open Access
Screening of Different Rice Genotypes against (Pyricularia grisea) Sacc. in Natural Epidemic Condition at Seedling Stage in Chitwan, Nepal
Khanal Sabin1*, Subedi Bijay1, Bhandari Amrit1, Giri Dilli Raman1, Shrestha Bhuwan1, Neupane Priyanka2, Shrestha Sundar Man2 and Gaire Shankar Prasad21Institute of Agriculture and Animal Science, Tribhuwan Univeristy, Chitwan, Nepal
2Department of Plant Pathology, Faculty of Agriculture, Agriculture and Forestry University, Nepal
- Corresponding Author:
- Sabin Khanal
Institute of Agriculture and Animal Science
Tribhuwan University, Nepal
Tel: +9779849745130
E-mail: savvy.khanal33@gmail.com
Received date: July 02, 2016; Accepted date:July 15, 2016; Published date: July 20, 2016
Citation:Sabin K, Bijay S, Amrit B, Raman GD, Bhuwan S, et al. (2016) Screening of Different Rice Genotypes against (Pyricularia grisea) Sacc. in Natural Epidemic Condition at Seedling Stage in Chitwan, Nepal. Adv Crop Sci Tech 4:231. doi:10.4172/2329-8863.1000231
Copyright: © 2016 Sabin K, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Advances in Crop Science and Technology
Abstract
Numerous research has already establish blast as the continuous and devastating threat to rice production in Nepal and on the contrary Nepalese farmers do not have efficient knowledge and understanding about the complexity of disease for the management of the blast epidemic development. The most effective physical tool seems to be provision of resistant genotypes obtained against screening of different rice genotypes: effective management practices against the complexity of blast pathogen. Experiments were conducted for screening 50 rice genotypes under natural epidemic condition against seedling blast (Pyricularia grisea) in Randomized complete block design at Chitwan. Rice grains were sown on July 6, 2015 at field and disease scoring was done on 21, 24, 27 and 30 DAS; Scoring was done based on the standard scale of 0-9 developed by IRRI. Based on the result Taichung-176 and Sankharika showed the highest percentage of incidence and severity of disease. Sabitri, however, was found to be most resistant among genotypes with the lowest percentage of incidence and severity during observation.
Keywords
Rice blast; Pyriculariagrisea;Sabitri; Blast susceptible
Introduction
Blast is caused by Pyriculariagrisea. It occurs in nearly all rice growing areas of the world. It is considered the most serious disease in both temperate and tropical rainfedenviroments. With increasing nitrogenous fertilizer and higher plant density, blast is known to be devastating [1].Blast was first recorded in china in 1637. The causal organism was named Pyriculariaoryzaeby Cavara in italy in 1891 and was renamed by Rossman 1990 to Pyriculariagrisea [2].
Rice is truly a crop of global importance. Almost half the world’s population, particularly in east and south east asia, depends on rice as the major source of nutritional calories [3]. Every year it is estimated that rice blast destroy food more than enough to eat for 60 million people and 50% of the rice yield is lost in the field by the occurrence of blast [4] Rice is the most prestigious food crop of Nepal. It is grown in a diverse environment ranging from tropical plains to foot of the mountain and higher elevation (3050 masl) in Chhumchure, Jumla. Nepal is considered as one of the origin center of rice. It is one of the most important cereal crops in Nepal. Rice is grown in 1440 thousand ha and the productivity is 2.56 t/ha. It contributes nearly 20 per cent to the agricultural gross domestic product. Nepal has released fifty five rice varieties with full package of growing practices in the last 40 years. The coverage by improved varieties is 85 percent of the total rice cultivated land. Popularly cultivated improved varieties are Radha-4, Radha-12, Masuli, Sabitri, CH-45, Bindeswori in terai, Khumal-4, Khumal-11, Taichung-176, chaining in mid-hills and Chandanaath-3 in high hills (NARC 2014). Radha-12, sabitri, janaki possess higher level of resistance [5]. Seedlings of high yielding masuli were affected in late june in saradanagar, Rampur, kiranganj, mangalpur and ratanagar area of the chitwan district [6]. Radha-12 had 7 fold less neck blast than masuli whereas other genotypes showed less neck blast than masuli [5].
Rice blast genetic analysis confirmed gene for gene interaction that control cultivar specificity in fungal plant interactions. Nuclear and mitochondrial genomes molecular analyses suggest that M. grisea pathogen remain in nature as different types of genetically distinct asexually reproducing population [7]. An understanding of the molecular mechanism that govern host specificity should aid in the development of new strategies for control of rice blast [8]. Mechanism controlling host species specificity differ in basic compatibility factor that allows pathogen to infect particular species. PWL2 host species specificity gene has properties analogous to classical avirulence genes, which function to prevent infection of certain cultivars of particular host species. The PWL2 gene encodes a glycine-rich, hydrophilic protein with a putative secretion signal sequence [8]. Blast, caused by Pyricularia grisea Sacc has been a continuous threat to rice production in Nepal [9,10]. Blast epidemics result in a complete loss of seedlings in the seedbed [6,11-16]
Varying tools have been used as a blast management toolkit such as knowledge tools, communication, physical and policy tools. Each tool is rationalized in terms of having an effect either on initial inoculum or disease (x0), the epidemic infection rate(R) or duration of epidemic (D). Certain tools like biological control agents and confirmatory serological tools are still unknown to blast control. Nepalese farmers do not have efficient knowledge and understanding about the use of fungicide and the effect of nutrient (nitrogen, phosphorus, potassium), and nonnutrient (silicon) amendments on blast epidemic development. Water management to reduce stress on plants at blast susceptible stages [17] are still in the dark to the Nepalese common farmers. Cultural practices seem ineffective due to no clear cut fallow period between any two rice seasons making blast pathosystem a continuous pathosystem and given the dispersal pattern of conidia initial inoculum will always be available for matching alloinfection. Hence, the most effective physical tool seems to be provision of resistant genotype. Seed possessing resistant genes to blast have been the basis for plant protection for centuries [13].
Materials and Methods
Experimental setup
Field experiment was set up in Agronomy farm of IAAS, Rampur, Chitwan. The experiment was conducted in single factorial RCBD design with 3 replication. Each plot was 5 mX1 m, in each replication 50 rows was made to sow the 50 different genotypes. Seed was sown randomly in such a way that, genotypes was not repeated in line in the replications. Seeding was done in 2nd week of July.
Observation
Disease assessment
Disease incidence: Appearance of first symptoms of disease among all the plants germinated will be recorded. Here, total no. of plants in a row and Number of plants showing the symptoms will be recorded (Figures 1 and 2).
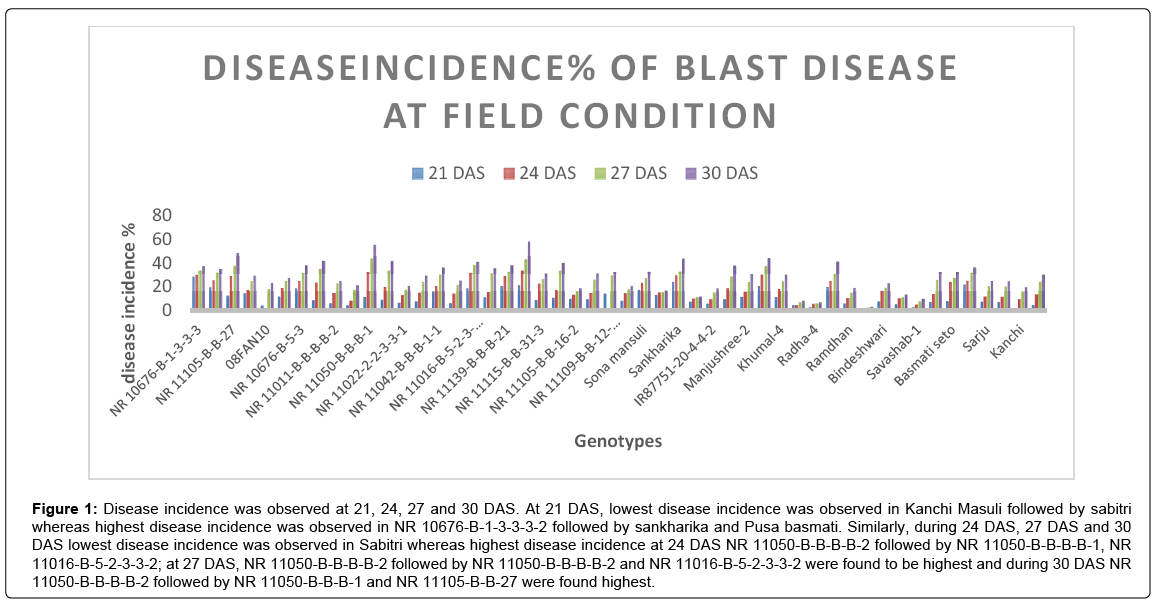
Figure 1: Disease incidence was observed at 21, 24, 27 and 30 DAS. At 21 DAS, lowest disease incidence was observed in Kanchi Masuli followed by sabitri whereas highest disease incidence was observed in NR 10676-B-1-3-3-3-2 followed by sankharika and Pusa basmati. Similarly, during 24 DAS, 27 DAS and 30 DAS lowest disease incidence was observed in Sabitri whereas highest disease incidence at 24 DAS NR 11050-B-B-B-B-2 followed by NR 11050-B-B-B-B-1, NR 11016-B-5-2-3-3-2; at 27 DAS, NR 11050-B-B-B-B-2 followed by NR 11050-B-B-B-B-2 and NR 11016-B-5-2-3-3-2 were found to be highest and during 30 DAS NR 11050-B-B-B-B-2 followed by NR 11050-B-B-B-1 and NR 11105-B-B-27 were found highest.
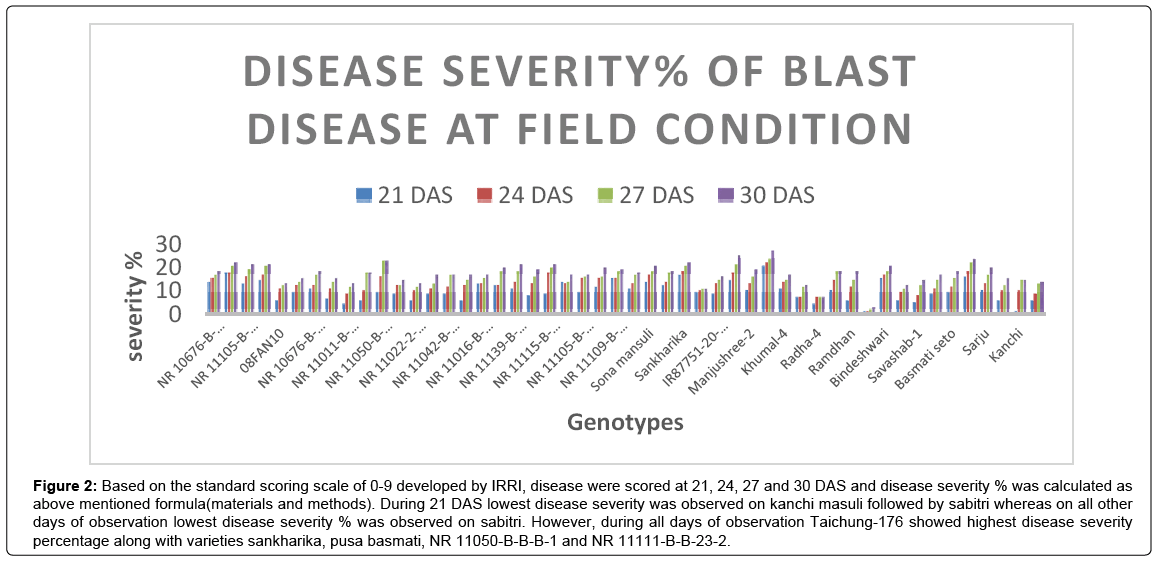
Figure 2: Based on the standard scoring scale of 0-9 developed by IRRI, disease were scored at 21, 24, 27 and 30 DAS and disease severity % was calculated as above mentioned formula(materials and methods). During 21 DAS lowest disease severity was observed on kanchi masuli followed by sabitri whereas on all other days of observation lowest disease severity % was observed on sabitri. However, during all days of observation Taichung-176 showed highest disease severity percentage along with varieties sankharika, pusa basmati, NR 11050-B-B-B-1 and NR 11111-B-B-23-2.
Percent disease incidence will be calculated by using the formula:

Disease scoring: Disease scoring will be done according to standard scoring scale developed by International Rice Research Institute (IRRI) using a scale of 0-9.
• Small brown specks of pin point size.
• Small roundish to slightly elongated, necrotic gravy spots, about 1-2 mm in diameter, with a distinct brown margin, lesions are mostly found on the lower leaves.
• Lesion type is the same as in 2, but significant number of lesion are on the upper leaves.
• Typical susceptible blast lesions, 3 mm or longer, infecting less than 4% of the leaf area.
• Typical susceptible blast lesions, 3 mm or longer, infecting less than 4-10% of the leaf area.
• Typical susceptible blast lesions, 3 mm or longer, infecting less than 11-25% of the leaf area.
• Typical susceptible blast lesions, 3 mm or longer, infecting less than 26-50% of the leaf area.
• Typical susceptible blast lesions, 3 mm or longer, infecting less than 51-75% of the leaf area, many leaves dead.
• Typical susceptible blast lesions, 3 mm or longer, infecting more than 75% of the leaf area
Disease intensity/index: Disease severity will be scored on the basis of standard scoring scale developed by International Rice Research institute (IRRI). 5 plants from each row showing the symptoms will be selected at random for observation and scored at a scale of 0-9 and average will be taken.
Disease severity will be calculated as:
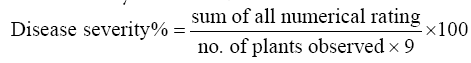
AUDPC:
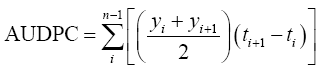
Where, yi: initial infection percentage (disease score)
Yi+1: progressive infection percentage
Ti+1-ti: time interval between the readings
Area under disease progressive curve
Total AUDPC value lied in the range of 17.78-210%. AUDPC 1&2 were calculated based on the disease severity percentage and calculated using formula as presented in the materials and methods above. Lowest total AUDPC was observed on Sabitri whereas highest was observed on Taichung-176 followed by pusa basmati, NR 11111-B-B-23-2, Sankharika and NR 10490-89-3-2-1. Based on the Total AUDPC value rice genotypes were listed on the five categories from resistant to highly susceptible which are shown in the Tables 1 and 2.
| Genotypes | AUDPC1 | AUDPC2 | AUDPC 3 | TOTAL AUDPC |
|---|---|---|---|---|
| NR 10676-B-1-3-3-3 NR 10490-89-3-2-1 NR 11105-B-B-27 NR 11052-B-B-B-B_6 08FAN10 NR10769-4-2-2 NR 10676-B-5-3 NR 11011-B-B-B-B-3 NR 11011-B-B-B-B-2 Sugandha-2 NR 11050-B-B-B-1 NR 11037-B-B-B-B-5 NR 11022-2-2-3-3-1 NR 11092-B-B-B-12 NR 11042-B-B-B-1-1 NR 11082-B-B-B-5-3 NR 11016-B-5-2-3-3-2 NR 11011-B-B-B-B-6 NR 11139-B-B-B-21 NR 11050-B-B-B-B-2 NR 11115-B-B-31-3 NR 11130-B-B-B-19 NR 11105-B-B-16-2 NR 11111-B-B-23-2 NR 11109-B-B-12-3-2 Madhya dhan -845 Sonamansuli Radha-22 Sankharika Radha-11 IR87751-20-4-4-2 NR 11111-B-B-23 Manjushree-2 Taichung-176 Khumal-4 Kalomasino Radha-4 Sawamansuli Ramdhan Sabitri Bindeshwari Sukkah-3 Savashab-1 Jethimansuli Basmati seto Pusa basmati Sarju Hardinath Kanchi Makwanpure |
44.44bcdefg 53.33ab 44.44bcdefg 47.78abcde 25.56hijkl 33.33defghijkl 35.56cdefghijk 26.67hijkl 20.00klm 24.44hijkl 38.89bcdefghij 32.22defghijkl 24.44hijkl 30.00fghijkl 31.11efghijkl 27.78ghijkl 40.00bcdefghi 37.78bcdefghij 37.77bcdefghij 32.22defghijkl 40.00bcdefghi 41.11bcdefgh 37.78bcdefghij 41.11bcdefgh 46.67bcdef 36.67bcdefghijk 46.67bcdef 40.00bcdefghi 53.33ab 30.00fghijkl 33.33defghijkl 48.89abcde 35.56cdefghijk 64.44a 37.78bcdefghij 22.22jkl 17.78lm 37.78bcdefghij 26.67hijkl 4.44m 48.89abcd 23.33ijkl 20.00klm 30.00fghijkl 32.22defghijkl 52.22abc 35.56cdefghijk 24.44hijkl 17.78lm 22.22jkl |
48.49bcdefghi 57.78abcd 53.33abcdef 56.67abcde 35.56ghijklmn 40.00fghijkl 44.44cdefghijkl 37.78fghijklm 31.11klm 42.22defghijkl 58.89abc 37.78fghijklmn 33.33ijklm 36.67ghijklm 43.33cdefghijkl 41.11efghijkl 43.33cdefghijkl 46.67bcdefghijk 48.89bcdefghi 44.44cdefghijkl 56.67abcde 41.11efghijkl 47.78bcdefghij 47.78bcdefghij 51.11bcdefg 45.56bcdefghijk 53.33abcdef 47.78bcdefghij 58.89abc 32.22jklm 42.22defghijkl 58.89abc 44.44cdefghijkl 68.89a 43.33cdefghijkl 28.89lm 22.22m 50.00bcdefgh 40.00fghijkl 5.56n 53.33abcdef 31.11klm 31.11klm 38.89fghijkl 41.11efghijkl 61.11ab 45.56bcdefghijk 34.44hijklm 37.78fghijklm 33.33ijklm |
53.33bcdefghij 64.44abc 61.11abcde 63.33abc 38.89fghijk 44.44cdefghij 53.33bcdefghij 44.44cdefghij 37.78ghijk 53.33bcdefghij 68.89ab 41.11efghijk 37.78ghijk 45.56cdefghij 51.11bcdefghij 47.78cdefghij 47.78cdefghij 57.78abcdefghij 60.00abcde 53.33bcdefhij 62.22abcd 45.56cdefghij 50.00bcdefghij 54.44bcdefghi 56.67abcdefgh 52.22bcdefghij 58.89abcdef 54.44bcdefghij 64.44abc 33.33jk 46.67cdefghij 70.00ab 53.33bcdefghij 76.67a 47.78cdefghij 36.67hijk 22.22kl 55.56bcdefghi 50.00bcdefghij 7.77l 58.88abcdef 35.55ijk 41.11efghijk 47.78cdefghij 51.11bcdefghij 68.89ab 55.56bcdefghi 42.22defghijk 44.44cdefghij 41.11efghijk |
146.67bcdefghij 175.56abcd 158.89bcdefgh 167.78abcde 100.00jklm 117.78fghijkl 133.33bcdefghijkl 108.89ijklm 88.89lm 120.00efghijkl 166.66abcdef 111.11hijklm 95.56klm 112.22ghijkl 125.56efghijkl 116.67ghijkl 131.11cdefghijkl 142.22bcdefghijk 146.67bcdefghij 130.00cdefghijkl 158.89bcdefgh 127.78defghijkl 135.56bcdefghijkl 143.33bcdefghijk 154.44bcdefghi 134.44bcdefghijkl 158.89bcdefgh 142.22bcdefghijk 176.67abcd 95.56klm 122.22efghijkl 177.78abc 133.33bcdefghijkl 210.00a 128.89cdefghijkl 87.78lm 62.22mn 143.33bcdefghijk 116.67ghijkl 17.78n 161.11abcdefg 90.00lm 92.22lm 116.67ghijkl 124.44efghijkl 182.22ab 136.67bcdefghijkl 101.11jklm 100.00jklm 96.67klm |
Table 1: AUDPC values of rice genotypes.
| Category | Range | Genotypes |
|---|---|---|
| Resistant | 0-70 | Sabitri Radha-4 |
| Moderately resistant | 71-120 | Kalomasino Sukkah-3 NR 11011-B-B-B-B-2 Savashab-1 NR 11022-2-2-3-3-1 KanchiMansulimasuli Hardinath Radha-11 08FAN10 Makwanpure NR 11092-B-B-B-12 NR 11037-B-B-B-B-5 NR 11011-B-B-B-B-3 NR 11082-B-B-B-5-3 Ramdhan Jethimansuli NR 10769-4-2-2 Sugandha-2 |
| Moderately susceptible | 121-140 | IR87751-20-4-4-2 Basmati seto NR 11042-B-B-B-1-1 NR 11130-B-B-B-19 Khumal-4 NR 11050-B-B-B-B-2 NR 11016-B-5-2-3-3-2 NR 10676-B-5-3 Manjushree-2 Madhya dhan-845 NR 11105-B-B-16-2 Sarju |
| Susceptible Highly susceptible | 141-170 171-225 | Radha-22 NR 11111-B-B-23-2 NR 11011-B-B-B-6 NR 10676-B-1-3-3-3 NR 11139-B-B-B-21 SawaMansuli NR 11109-B-B-12-3-2 Bindeshwari NR 11105-B-B-27 NR 11115-B-B-31-3 NR 11050-B-B-B-1 SonaMansuli NR 11052-B-B-B-B-6 NR 11111-B-B-23-2 Sankharika Pusa Basmati Taichung-176 NR 10490-89-3-2-1 |
Table 2: Based on the AUDPC value rice genotypes are listed on the five categories from resistant to highly susceptible.
Discussion
Disease incidence was observed at 21, 24, 27 and 30 DAS. At 21 DAS, lowest disease incidence was observed in Kanchi Masuli followed by sabitri whereas highest disease incidence was observed in NR 10676- B-1-3-3-3-2 followed by sankharika and Pusa basmati. Similarly, during 24 DAS, 27 DAS and 30 DAS lowest disease incidence was observed in Sabitri whereas highest disease incidence at 24 DAS NR 11050-B-BB- B-2 followed by NR 11050-B-B-B-B-1, NR 11016-B-5-2-3-3-2; at 27 DAS, NR 11050-B-B-B-B-2 followed by NR 11050-B-B-B-B-2 and NR 11016-B-5-2-3-3-2 were found to be highest and during 30 DAS NR 11050-B-B-B-B-2 followed by NR 11050-B-B-B-1 and NR 11105-B-B-27 were found highest [18-20]. Sabitri was reported to be most resistant by Chaudary et al. [5]. Genotypes starting with NR initial were breeding lines developed by Nepal Agriculture Research Council (NARC) as they were developed for high hills and this experiment being conducted in the terai region might had induced blast incidence on these lines due to unsuitable temperature to the genotypes.
During 21 DAS lowest disease severity was observed on kanchi masuli followed by sabitri whereas on all other days of observation lowest disease severity % was observed on sabitri. However, during all days of observation Taichung-176 showed highest disease severity percentage along with varieties sankharika, pusa basmati, NR 11050-B-B-B-1 and NR 11111-B-B-23-2. Experiment by Manandher et al. [11] presented sankharika to be most susceptible variety and established that it is adversely affected by blast pathogen whereas Taichung-176 were found to be highly susceptible variety by Manandhar et al. [9]. Kumar et al. [21] reported pusa basmati as most susceptible to blast disease.
Similarly sabitri showed lowest level of AUDPC value and was categorized as the resistant genotype along with Radha-4 which is supported by Chaudary et al. [5] suggesting that Sabitri and Radha varieties to be resistant to blast pathogen; whereas Taichung-176 pusa basmati and Sankharika were categorized as the most susceptible varieties which coincides with the result presented by (Manandhar et al. and Manandhar et al.) and Kumar et al. [9,11,21]. Similary, NR 11111-B-B-23-2 and NR 10490-89-3-2-1 were also categorized as the most susceptible and more conclusive result are yet to be drawn of these genotypes.
Conclusion
50 rice genotypes were sown in 6th July in Randomized complete block design at chitwan. The experiment was only limited to seedling stage and its purpose was to identify the resistant and susceptible variety among the different rice genotypes collected all over the country along with some of the breedling lines provided by the NARC khumaltar. Sankharika, Taichung-176, pusa basmati, NR 11111-B-B-23 and NR 10490-89-3-2-1 were found most susceptible and sabitri and Radha-4 were found to be resistant.
As Taichung-176, sankharika was found to be most susceptible to blast on both field and lab condition as NARC has described Taichung-176 as susceptible variety to mid-hills and sankharika to Terai region. Sabitri was found to be most resistant among all genotypes. Further research is recommended on the varieties mentioned above for further certainty; in addition, further research work such as comparison of plant yield with disease can be done and also molecular study of the plant varieties is further recommended.
Acknowledgements
I would like to express my sincere gratitude to prof. Dr. Sundarman Shrestha and Mr. Shankhar Gaire, Faculty of Plant pathology at Agriculture and Forestry University for their guidance and support during all the activity of this research. I would also like to acknowledge Mr. Bikash adhikari for his help during all the field activities; Mr. Ramesh Acharya for his support; Mrs. Shawantana Ghimire Khanal for her encouragement to write this paper.
References
- Kato H (1974) Epidemiology of rice blast disease. REV plant prot Res 7:1-20.
- Rossman AY (1990) Pyriculariagrisea the correct name for the Rice Blast disease fungus. Mycologia 82:509-512.
- Swaminathan MS (1982) Biotechnology factors in the epidemiology of rice blast. Annual review of phytopathology 13: 139-256.
- Barman RS, Chattoo BB (2005) Rice blast fungus sequenced. Current Science 89: 930-931
- Chaudary B, Shrestha SM, Sharma RC (2001) Resistance in Rice breeding lines to blast fungus in Nepal. Journal of institute of agriculture and animal sciences (Nepal)2001. Nepal agriculture research journal 6: 49-56.
- Adhikari TB, Shrestha SM(1986) Blast epidemic in chitwan valley, Nepal. International Rice Research Newsletter 11:22.
- Zeigler RS, Leong SA,Teng PS (1994) Rice Blast Disease. Commonwealth Agricultural Bureau International, Wallingford, England.
- Swelgard JA, Carrol AM, Kang S, Farral L, Chumley FG, et al.(1995) Identification,cloning and characterization of PWL2 of gene for host species specificity in Rice Blast fungus. The plant cell 7:1221-1233.
- Manandhar HK, Shrestha K, Amatya P (1992) Seed-borne diseases. In: Plant diseases, seed production and seed health testing in Nepal. Danish Government, Institute of Seed Pathology for Developing Countries, Copenhagen, Denmark. pp: 59-74
- Chaudary B (1999) Effect of blast disease on rice yield. Nepal Agricultural Research Journal 3:8-13.
- Manandhar HK, Thapa BJ, Amatya P (1985) Efficacy of various fungicides on the control of rice blast disease. Journal of Institute of agriculture and animal sciences 6:21-29.
- Pradhanang PM (1988) Outbreak of blast disease at Lumle Agricultural Centre (LAC) and its extension command area (ECA). In: Proceedings of the first national rice blast workshop. National Agricultural Research and Services Center, National Rice Research Program, pp:61-69.
- Reissig W, Heinrichs E, Litsinger J, Moody K, Fiedler L (1986) Illustrated guide to integrated pest management in rice in tropical Asia. IRRI, Los Banos, Laguna, Philippines.
- Chaudhary B, Karki PB, Lal KK (1994) Neck blast resistant lines of Radha-17 isolated. International Rice Research Notes 19:11
- Chaudhary B, Sah DN (1997) Effect of promising rice genotypes on leaf blast disease progression. Nepal Agriculture Research Journal 1:27-31.
- Chaudhary B, Sah DN (1998) Efficacy of Beam 75 WP in controlling leaf blast disease at the seedling stage of rice. Nepal Agriculture Research Journal 2:42-47.
- Teng PS, Klein-Gebbink HW, Pinnschmidt H (1991) An analysis of the blast pathosystem to guide modelling and forecasting. In: Rice blast modelling and forecasting. International Rice Research institute, Manila, Philippines. pp:1-30.
- Kingslover CH, Mackenzie DR, Rush MC (1988) Field testing a computerized forecasting systems for rice blast disease. Phytopathology 78: 931- 934.
- Sah DN (1989) Effects of flooding and leaf wetness duration on resistance of rice lines to Pyricularia oryzae. Journal of Institute of Agriculture and Animal Sciences 10:41-48.
- Suryanarayan S (1966) Environment and the blast disease. Indian phytopathological society 3: 100-114.
- Kumar N, Singh D, Gupta S, Sirohi A, Ramesh B, et al. (2013) Determination and expression of genes for resistant to blast(M. oryzae) in Basmati and Non-basmati Indica rices (Oryzae sativa L.). African journal of Biotechnology12: 4098-4104.
Relevant Topics
- Agricultural science
- Agronomy
- Climate impact on crops
- Crop Productivity
- Crop Sciences
- Crop Technology
- Field Crops Research
- Hybrid Seed Technology
- Irrigation Technology
- Organic Cover Crops
- Organic Crops
- Pest Management
- Plant Genetics
- Plant Breeding
- Plant Nutrition
- Seed Production
- Seed Science and Technology
- Soil Fertility
- Weed Control
Recommended Journals
Article Tools
Article Usage
- Total views: 13332
- [From(publication date):
August-2016 - Jul 12, 2025] - Breakdown by view type
- HTML page views : 12196
- PDF downloads : 1136