Research Article Open Access
Sulfur Amino Acid Metabolic Process Pathway may Modulate Bipolar Disorder with Alcohol Dependence Comorbidity
Enrico Cocchi1, Antonio Drago2* and Alessandro Serretti11Department of Biomedical and NeuroMotor Sciences, University of Bologna, Italy
2IRCCS Centro S. Giovanni di Dio, Fatebenefratelli, Brescia, Italy
- Corresponding Author:
- Antonio Drago
Institute of Psychiatry, University of Bologna
Viale Carlo Pepoli 5, 40123 Bologna, Italy
Tel: +39 051 6584233, +39 320 4269332, +39 347 3024020
Fax: +39 051 521030
E-mail: antonio.drago@unibo.it
Received date: January 16, 2014; Accepted date: March 29, 2014; Published date: March 31, 2014
Citation: Cocchi E, Drago A, Serretti A (2014) Sulfur Amino Acid Metabolic Process Pathway may Modulate Bipolar Disorder with Alcohol Dependence Comorbidity. J Addict Res Ther 5:177. doi:10.4172/2155-6105.1000177Copyright: © 2014 Cocchi E, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Journal of Addiction Research & Therapy
Abstract
Background: A relationship between alcohol use disorder and Bipolar Disorder (BD) has far been detected. A record of alcohol dependence may worsen the course of BD. Nevertheless, the genetic underpinnings of this comorbidity have not been completely elucidated. Authors investigated the impact of a set of genetic variations as possible risk factors for the pathological mood swings in bipolar patients with a record of alcohol dependence.
Methods: A list of candidate genes identified as risk loci by GWAS studies in last 10 years were tested in a sample of 802 bipolar patients from the public available STEP-BD study. Variations harbored by these genes were checked for quality, imputed and pruned. A set of 260 genes embedded in 160 different pathways were analyzed as predictors of the frequency of severe (YMRS>11) manic events and depressive phases (MADRS>19) during the period of observation (1139 days for manic records and 1856 for depressive records). Their effect was tested in combination with alcohol comorbidity. Clinical and sociodemographic variables entered the study as covariates when significantly associated with the phenotypes.
Results: We found an impact of alcohol dependence positive record with a higher frequency of severe manic (p=0.02) and depressive (p=0.0006) records. A positive association between a pathway related to the Sulfur amino acid metabolic process (GO:0000096) and an increased frequency of severe depressive records was detected for BD subjects with a record of alcohol dependence.
Discussion: We found an association between GO: 0000096 (Sulfur amino acid metabolic process pathway) and severe depressive episodes in BD patients with a record of alcohol dependence in their clinical story.
Keywords
Alcohol; Dependence; Bipolar disorder; Genetics; Pathway; Severity
Introduction
Alcohol comorbidity in Bipolar Disorder (BD) attracted much attention during the last years [1] because of its relevant impact on prognosis. Co-occurrence of alcohol misuse during BD is common [2,3] and patients with both types of disorder are typically more difficult to treat than patients who have either problem alone [4]. BD shows a 1% prevalence worldwide, which rises up to 6.4% when subthreshold cases are, included [5]. Its costs were estimated to be $45.2 billion in 1991 (in the U.S.) [6]. Alcohol use disorders concern 25% of adults in the U.S. [7] overall economic cost was estimated to be $148 billion for 1992, $166.6 billion in 1995 and $184.6 billion in 1998 [8]. Moreover, alcohol use is the third leading preventable cause of death [9]. Thus, the costs of the combination of BD and alcohol use disorders are extremely high [10] and projected to increase. In this prospective alcohol dependence is one of the biggest problems related to alcohol [11] and its impact is particularly high in BD patients. Studies showed that 27.6% of any BD and especially 31.5% of BD type I patients suffered of alcohol dependence [12]. This comorbidity resulted in an increased risk for suicide attempt, greater severity of symptoms and impaired functioning. Compared with the lower rate of patients that suffer comorbidity of alcohol dependence with unipolar depression (11.6%) or with any mood disorder (4.9%), BD and alcohol dependence reveals a even stronger relationship [12]. As a consequence, the proper treatment of BD patients and the identification of subjects at risk is a priority. Consistently, a brain degeneration during BD is reported to be a late-phase characteristic that depends on the number of episodes of illness [13] and alcohol dependence could play a role in worsening this relapses number. Moreover, a proper treatment following the identification of BD patients at risk for alcohol dependence could successfully decrease the number of bipolar relapses through life and their severity. As genetics reveals, both BD and alcohol dependence are heritable [14,15]. Interestingly, in BD, there are lines of evidence that 71% of the genetic variance for mania was not shared with depression [15]. Several attempts tried to elucidate the genetic background leading alcohol dependence but results are not conclusive so far. Aldehyde dehydrogenase 2 (ALDH2), alcohol dehydrogenase 1B (ADH1B), gamma-aminobutyric acid (GABA) A receptor, and the GABA alpha 2 (GABRA2) receptors pathway were recently reported to be central to alcohol misbehavior [16]. Nevertheless, there is sufficient evidence to say that the genetic liability to BD and alcohol dependence is derived from the orchestrated activity of a set of genes and not by a single one. Starting from this perspective, we 1) collected all the genes identified as possible risk loci for alcohol dependence in the last 10 years of GWAS studies in the field and 2) described the biological pathways they are relate to 3) analyzed the influence of the variations on the genes that code for the proteins that are embedded in these pathways on the frequency of both severe (Young Mania Rating Scale (YMRS) overall score>11) manic events and depressive phases (Montgomery-Asberg Depression Scale (MADRS) overall score >19) in BD patients. Considering the enormous difference in GWAS results, we also enriched our initial subset of genes with Cytoscape [17] in order to considering the major number of associated genes and pathways in the analysis. As a clear clinical conclusion, the definition of the genetic impact on alcohol dependence and BD comorbidity could be instrumental to the early diagnosis and even to primary prevention of BD in patients who also present with alcohol dependence.
Methods
Sample under investigation
The sample under investigation was retrieved from the public available STEP-BD protocol [18]. During the study, bipolar patients of every subtype with age ≥ 15 years are accessioned into a study registry. All patients receive a systematic assessment battery at entry and are treated by a psychiatrist (trained to deliver care and measure outcomes in patients with BD) using a series of model practice procedures consistent with expert recommendations. At every follow-up visit, the treating psychiatrist completes a standardized assessment and assigns an operationalized clinical status based on DSM-IV criteria. Patients have independent evaluations at regular intervals throughout the study and remain under the care of the same treating psychiatrist while making transitions between randomized care studies and the standard care treatment pathways. We were able to identify 802 patients assessed with the MADRS and YMRS scales. Characteristics of the samples under analysis is reported in Table 1.
| Variable | mean±SD all | mean±SD alcohol-abuse | mean±SD all non-alcohol-abuse | statistics |
|---|---|---|---|---|
| Age | 41.18±12.37 | 41.00±11.02 | 41.26±12.96 | T = 0.3007
df = 582.468 p-value = 0.7637 |
| Sample size | 802(100%) | 257(32%) | 545(68%) | / |
| Gender | Female=446(55.6%) Male=354(44.2%) Transgender=2(0.2%) |
Female=120(46.7%) Male=137(53.3%) Transgender=0(0.0%) |
Female=326(59.8%) Male=217(39.8%) Transgender=2(0.2%) |
X-squared = 13.5534 df = 2 p-value = 0.00114 |
| YMRS > 11 manic episode frequency / times observed | 0.16±0.24 | 0.19±0.26 | 0.14±0.23 | T = -2.3327
df = 445.442 p-value = 0.02011 |
| MADRS > 19 episode frequency / times observed | 0.31±0.33 | 0.37±0.34 | 0.28±0.32 | T = -3.4181
df = 474.914 p-value = 0.0006848 |
| Ethnicity | White or Caucasian=802(100.0%) | White or Caucasian=257(100.0%) | White or Caucasian=545(100.0%) | / |
The table is a resumen of the clinical characteristics of the patients taken into account for the study.
YMRS=Young Mania Rating Scale
MADRS=Montgomery Asberg Depression Rating Scale
SD=Standard Deviation
Table 1: Sample under analysis.
Definition of phenotype
The phenotype under analysis was the number of severe manic phases (manic relapses with an overall YMRS [19] point>11) and severe depressive episodes (depressive episodes with MADRS [20] total score >19) corrected for the number of observations during the STEP-BD period of observation. We took these thresholds (YMRS>11 and MADRS>19) accordingly to the standard YMRS and MADRS interpretation to defining manic phases [19] and depressive ones [21] severity in order to avoid stratification factors due to different interpretation of these assessment scales. This particular phenotype was chosen to limit the impact of missing values in order to increase the power of the study. Care was taken to control for a possible clinical bias: more severe patients might have been seen more often compared to less severe ones. In order to do so, the correlations between the phenotype of choice and a set of other phenotypes calculated at standard time points (number of manic/depressive phases from 30-90- 120 and so forth days from the beginning of the study) - a more classical approach to this kind of studies - were calculated. We had confirmation that the phenotype under analyses significantly correlated with almost all the phenotypes at different timepoints, with the advantage of having 0% of missing values. Results are reported in Table 2 (mania) and Table 3 (depression). The only timepoint at which the correlation was not significant was after 30 days from the beginning of the study for the manic relapses. Nevertheless, the number of missing information for this timepoint (43.89%) may be held accountable for the lack of association. Subjects were labeled as alcohol dependent when there was a record of alcohol dependence in the past. This variable was used instead of alcohol dependence during the manic/depressive phase to distinguish the hedonistic use of alcohol during manic/depressive phases vs. a record of alcohol dependence which was deemed to be more related to a chronic habit. The alcohol dependence record in STEP-BD was assessed using the Mini Mental State Evaluation as the presence of clearly symptoms of alcohol dependence (not only alcohol abuse) during the last 12 months [19].
| days from baseline | manic relapses per time | missing (%)* | mean manic relapses per visit unit | p | t | conf 1 | conf 2 |
|---|---|---|---|---|---|---|---|
| 0 | 0.18 | 43.89 | 0.16±0.24 | 0.18 | -1.35 | -0.09 | 0.02 |
| 30 | 0.18 | 8.98 | 0.16±0.24 | 0.02 | -2.27 | -0.09 | -0.01 |
| 60 | 0.18 | 2.87 | 0.16±0.24 | 0.02 | -2.33 | -0.09 | -0.01 |
| 90 | 0.18 | 2.49 | 0.16±0.24 | 0.02 | -2.33 | -0.09 | -0.01 |
| 120 | 0.18 | 1.62 | 0.16±0.24 | 0.02 | -2.41 | -0.09 | -0.01 |
| 150 | 0.17 | 1.37 | 0.16±0.24 | 0.02 | -2.39 | -0.09 | -0.01 |
| 180 | 0.17 | 1.37 | 0.16±0.24 | 0.02 | -2.39 | -0.09 | -0.01 |
| 210 | 0.17 | 0.75 | 0.16±0.24 | 0.02 | -2.34 | -0.08 | -0.01 |
| 240 | 0.17 | 0.5 | 0.16±0.24 | 0.02 | -2.27 | -0.08 | -0.01 |
| 270 | 0.17 | 0.37 | 0.16±0.24 | 0.02 | -2.29 | -0.08 | -0.01 |
| 300 | 0.17 | 0.12 | 0.16±0.24 | 0.02 | -2.32 | -0.08 | -0.01 |
| 330 | 0.17 | 0.12 | 0.16±0.24 | 0.02 | -2.32 | -0.08 | -0.01 |
| 360 | 0.18 | 0.12 | 0.16±0.24 | 0.02 | -2.32 | -0.08 | -0.01 |
| 390 | 0.18 | 0.12 | 0.16±0.24 | 0.02 | -2.32 | -0.08 | -0.01 |
| 420 | 0.18 | 0.12 | 0.16±0.24 | 0.02 | -2.32 | -0.08 | -0.01 |
| 450 | 0.18 | 0.12 | 0.16±0.24 | 0.02 | -2.32 | -0.08 | -0.01 |
| 480 | 0.18 | 0.12 | 0.16±0.24 | 0.02 | -2.32 | -0.08 | -0.01 |
| 510 | 0.18 | 0.12 | 0.16±0.24 | 0.02 | -2.32 | -0.08 | -0.01 |
| 540 | 0.18 | 0.12 | 0.16±0.24 | 0.02 | -2.32 | -0.08 | -0.01 |
| 570 | 0.18 | 0.12 | 0.16±0.24 | 0.02 | -2.32 | -0.08 | -0.01 |
| 600 | 0.18 | 0.12 | 0.16±0.24 | 0.02 | -2.32 | -0.08 | -0.01 |
| 630 | 0.18 | 0.12 | 0.16±0.24 | 0.02 | -2.32 | -0.08 | -0.01 |
| 660 | 0.18 | 0.12 | 0.16±0.24 | 0.02 | -2.32 | -0.08 | -0.01 |
| 690 | 0.18 | 0.12 | 0.16±0.24 | 0.02 | -2.32 | -0.08 | -0.01 |
| 720 | 0.18 | 0.12 | 0.16±0.24 | 0.02 | -2.32 | -0.08 | -0.01 |
| 750 | 0.17 | 0.12 | 0.16±0.24 | 0.02 | -2.32 | -0.08 | -0.01 |
| 780 | 0.17 | 0.12 | 0.16±0.24 | 0.02 | -2.32 | -0.08 | -0.01 |
| 810 | 0.17 | 0.12 | 0.16±0.24 | 0.02 | -2.32 | -0.08 | -0.01 |
| 840 | 0.17 | 0.12 | 0.16±0.24 | 0.02 | -2.32 | -0.08 | -0.01 |
| 870 | 0.17 | 0.12 | 0.16±0.24 | 0.02 | -2.32 | -0.08 | -0.01 |
| 900 | 0.17 | 0.12 | 0.16±0.24 | 0.02 | -2.32 | -0.08 | -0.01 |
| 930 | 0.17 | 0 | 0.16±0.24 | 0.02 | -2.33 | -0.08 | -0.01 |
| 960 | 0.17 | 0 | 0.16±0.24 | 0.02 | -2.33 | -0.08 | -0.01 |
| 990 | 0.17 | 0 | 0.16±0.24 | 0.02 | -2.33 | -0.08 | -0.01 |
| 1020 | 0.17 | 0 | 0.16±0.24 | 0.02 | -2.33 | -0.08 | -0.01 |
| 1050 | 0.17 | 0 | 0.16±0.24 | 0.02 | -2.33 | -0.08 | -0.01 |
| 1080 | 0.17 | 0 | 0.16±0.24 | 0.02 | -2.33 | -0.08 | -0.01 |
| 1110 | 0.17 | 0 | 0.16±0.24 | 0.02 | -2.33 | -0.08 | -0.01 |
The table shows the statistics of association between the analyzed phenotype of the patients (total number of severe manic relapses corrected for the total number of observation for each patient) and the number of manic relapses at specific timepoints (every 30 days from the beginning of the study) that is a more classical approach for these kind of studies but is related to a bigger number of missings (totally avoided with our phenotype) and clinical bias (e.g. more severe patients might have been seen more often compared to less severe ones.).
Table 2: Study of the correlation between the phenotype under analysis and the number of manic relapses at specific timepoints.
| days from baseline | depressive phases per time | missing (%)* | mean depressive phases per visit unit | p | t | conf 1 | conf 2 |
|---|---|---|---|---|---|---|---|
| 0 | 0.54 | 43.64 | 0.31±0.33 | 0.02 | -2.35 | -0.15 | -0.01 |
| 30 | 0.58 | 8.48 | 0.31±0.33 | 0.002 | -3.03 | -0.13 | -0.03 |
| 60 | 0.58 | 2.74 | 0.31±0.33 | 0.001 | -3.13 | -0.13 | -0.03 |
| 90 | 0.58 | 2.37 | 0.31±0.33 | 0.001 | -3.22 | -0.13 | -0.03 |
| 120 | 0.59 | 1.5 | 0.31±0.33 | 0.001 | -3.17 | -0.13 | -0.03 |
| 150 | 0.59 | 1.25 | 0.31±0.33 | 0.001 | -3.28 | -0.14 | -0.03 |
| 180 | 0.59 | 1.12 | 0.31±0.33 | 0.001 | -3.26 | -0.14 | -0.03 |
| 210 | 0.58 | 0.75 | 0.31±0.33 | <0.001 | -3.36 | -0.14 | -0.04 |
| 240 | 0.58 | 0.62 | 0.31±0.33 | <0.001 | -3.34 | -0.14 | -0.04 |
| 270 | 0.58 | 0.5 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 300 | 0.58 | 0.25 | 0.31±0.33 | <0.001 | -3.43 | -0.14 | -0.04 |
| 330 | 0.58 | 0.25 | 0.31±0.33 | <0.001 | -3.43 | -0.14 | -0.04 |
| 360 | 0.58 | 0.25 | 0.31±0.33 | <0.001 | -3.43 | -0.14 | -0.04 |
| 390 | 0.57 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 420 | 0.57 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 450 | 0.57 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 480 | 0.57 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 510 | 0.57 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 540 | 0.58 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 570 | 0.57 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 600 | 0.57 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 630 | 0.57 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 660 | 0.57 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 690 | 0.57 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 720 | 0.57 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 750 | 0.56 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 780 | 0.56 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 810 | 0.56 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 840 | 0.56 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 870 | 0.56 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 900 | 0.56 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 930 | 0.56 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 960 | 0.56 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 990 | 0.56 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1020 | 0.56 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1050 | 0.56 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1080 | 0.56 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1110 | 0.56 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1140 | 0.55 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1170 | 0.55 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1200 | 0.55 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1230 | 0.55 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1260 | 0.55 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1290 | 0.55 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1320 | 0.55 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1350 | 0.55 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1380 | 0.55 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1410 | 0.55 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1440 | 0.55 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1470 | 0.55 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1500 | 0.55 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1530 | 0.55 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1560 | 0.55 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1590 | 0.55 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1620 | 0.55 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1650 | 0.55 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1680 | 0.55 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1710 | 0.55 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1740 | 0.55 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1770 | 0.55 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1800 | 0.55 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
| 1830 | 0.55 | 0.12 | 0.31±0.33 | <0.001 | -3.42 | -0.14 | -0.04 |
The table shows the statistics of association between the analyzed phenotype of the patients (total number of severe depressive phases corrected for the total number of observation for each patient) and the number of depressive phases at specific timepoints (every 30 days from the beginning of the study) that is a more classical approach for these kind of studies but is related to a bigger number of missings (totally avoided with our phenotype) and clinical bias (e.g. more severe patients might have been seen more often compared to less severe ones.).
Table 3: Study of the correlation between the phenotype under analysis and the number of depressive phases at specific time points.
Study of stratification factors
The sociodemographic and clinical variables were investigated as possible stratification factors. Age and gender were included in the analysis as covariates. Gender of patients resulted to be significantly associated with the phenotype (p=0.001). Ethnicity was not included as covariate because all patients resulted to be Caucasian. Table 1 reports the characteristics of the sample and the strength of association between the sociodemographic variables and the record of alcohol dependence in the past in bipolar subjects.
Power of the study and correction for multiple testing
We had sufficient power (0.80) to detect a medium effect size of 0.09 with a significance level of 0.05 between two alleles represented in a sample of 401 subjects each. Both a Bonferroni correction (p<0.05/160=0.0003) and a False Discovery Rate correction were applied to avoid false positive findings after checking 160 different molecular pathways.
Selection of genes
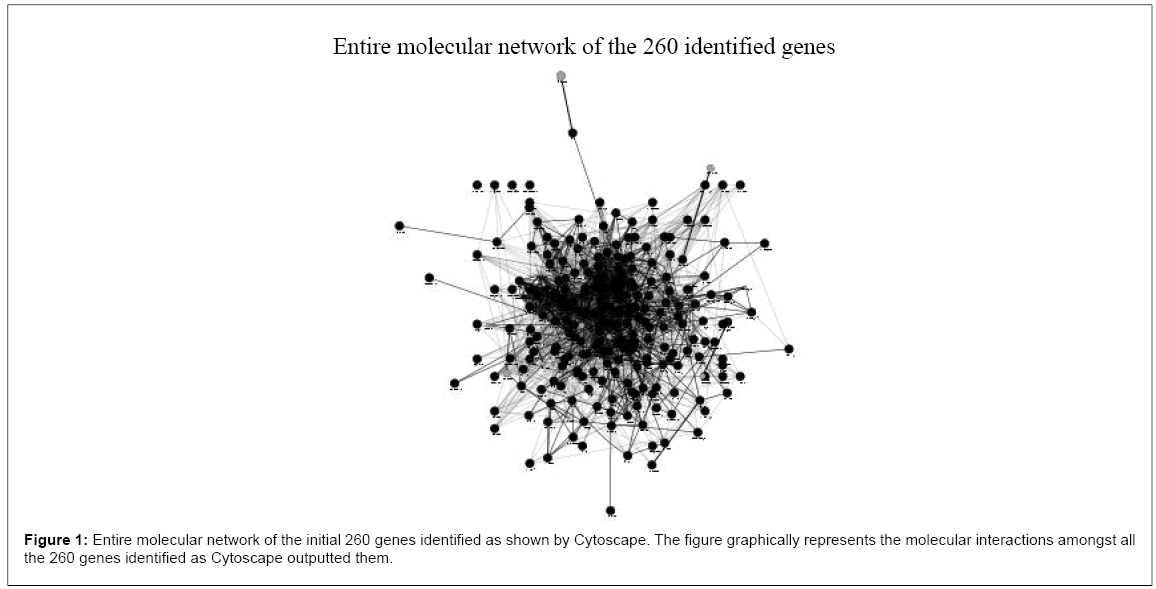
We selected 260 genes that showed a possible association as risk loci for alcohol dependence in literature available GWAS studies of the last 10 years (from 1993 to January 2014). Table 4 reports the studies analyzed. Genes and their characteristics are reported in Table 1, supplementary materials. We then enriched the gene subset and analyzed the pathway in which these genes play role using Cytoskape, Table 2, supplementary materials, reports the pathways characteristics and Figure 1 shows the entire molecular network of our genes.
| TITLE | Kind of study | Main result | REFERENCE | |
|---|---|---|---|---|
| 1 | The genetics of alcohol dependence. | Review | Genes involved in GABAergic, endogenous opioid, dopaminergic, cholinergic, and serotonergic transmission are selected as best candidates for AD related genes | [35] |
| 2 | Genome-wide association study of alcohol dependence:significant findings in African- and European-Americans including novel risk loci. | GWAS | 16,087 AD subjects; ADH1B, ADH1C, ADH gene cluster, PDLIM5, METAP, rs1437396, between MTIF2 and CCDC88A may be associated with AD | [36] |
| 3 | Family-Based Association Analysis of Alcohol Dependence Criteria and Severity. | GWAS | 118 extended European American families (n = 2,322 individuals); NALCN, OR51L1 may be associated with AD | [37] |
| 4 | Introduction to Deep Sequencing and Its Application to Drug Addiction Research with a Focus on Rare Variants. | Review | ALDH2 may be associated with AD | [38] |
| 5 | A genome-wide association study of behavioral disinhibition. | GWAS | 7,188 Caucasian individuals clustered in 2,300 nuclear families; rs1868152 (intronic) associated with the use of illicit drugs | [39] |
| 6 | Association, interaction, and replication analysis of genes encoding serotonin transporter and 5-HT3 receptor subunits A and B in alcohol dependence. | Candidate gene | 500 AD and 280 healthy control individuals of European descent; SLC6A4-HTR3A-HTR3B interact in affecting AD | [40] |
| 7 | Dosage transmission disequilibrium test (dTDT) for linkage and association detection. | TDT | Rs4903712 may affect AD behavior | [41] |
| 8 | A meta-analysis of two genome-wide association studies to identify novel loci for maximum number of alcoholic drinks. | Meta-analysis of GWAS | two GWAS using maxdrinks as an excessive alcohol consumption phenotype: one in 118 extended families (N = 2,322) selected from COGA,, and the other in a case-control sample (N = 2,593) derived from SAGE. Rs9523562 and rs67666182 were associated with AD. LMO1, AUTS2, INADL, HIP1 and PLCL1 were associated with alcohol related phenotypes. | [42] |
| 9 | Common biological networks underlie genetic risk for alcoholism in African- and European-American populations. | GWAS re-analysis | Chloride transporters and glycine metabolism genes are associated with AD | [43] |
| 10 | Replication of genome wide association studies of alcohol dependence: support for association with variation in ADH1C. | GWAS | 808 alcohol-dependent cases and 1,248 controls; ADH1C may be associated with AD | [44] |
| 11 | Extended genetic effects of ADH cluster genes on the risk of alcohol dependence: from GWAS to replication. | GWAS | ADH1B may be associated with AD | [45] |
| 12 | Multi-species data integration and gene ranking enrich significant results in an alcoholism genome-wide association study. | Mixed animal and GWAS in humans approach | COGA and SAGE samples; brain responses to ethanol and neural adaptations genes are involved in AD | [46] |
| 13 | Using genetic information from candidate gene and genome-wide association studies in risk prediction for alcohol dependence. | GWAS | COGA and SAGE samples | [47] |
| 14 | Genome-wide association study identifies a potent locus associated with human opioid sensitivity. | GWAS | Family history was a better predictor of AD than genes | [48] |
| 15 | A genome-wide association study of alcohol-dependence symptom counts in extended pedigrees identifies C15orf53. | GWAS | Collaborative Study on the Genetics of Alcoholism sample; C15orf53 may be associated with AD | [49] |
| 16 | TACR1 genotypes predict fMRI response to alcohol cues and level of alcohol dependence. | Imaging study | 326 individuals with alcohol use disorders; rs3771863, rs3755459, and rs1106855 (TACR1) predicted BOLD activation in response to alcohol cues | [50] |
| 17 | ANKRD7 and CYTL1 are novel risk genes for alcohol drinking behavior. | GWAS | 1972 Caucasians in 593 nuclear families, 761 unrelated Caucasian subjects, and 2955 unrelated Chinese Hans; ANKRD7, CYTL1 may be associated with AD | [51] |
| 18 | Genetic influences on craving for alcohol. | Candidate gene analysis and GWAS | a sample of unrelated adults ascertained for alcohol dependence (N=3976); DRD3, ITGADmay be associated with AD (craving) | [52] |
| 19 | A novel, functional and replicable risk gene region for alcohol dependence identified by genome-wide association study. | GWAS | a discovery sample of 681 African-American (AA) cases with alcohol dependence and 508 AA controls were retested in a primary replication sample of 1,409 European-American (EA) cases and 1,518 EA controls; PHF3-PTP4A1 locus may be associated with AD | [53] |
| 20 | A haplotype analysis is consistent with the role of functional HTR1B variants in alcohol dependence. | Candidate study | 136 Brazilian alcoholics of European descendent and 237 controls; rs11568817 (HTR1B) may be associated with AD | [54] |
| 21 | Genome-wide significant association between alcohol dependence and a variant in the ADH gene cluster. | GWAS | 1333 male in-patients with severe AD according to the Diagnostic and Statistical Manual of Mental Disorders, 4th edition, and 2168 controls; ADH1B, ADH1C may be associated with AD | [55] |
| 22 | Genome-wide association study of alcohol dependence implicates KIAA0040 on chromosome 1q. | GWAS | 4116 subjects (1409 European-American (EA) cases with AD, 1518 EA controls, 681 African-American (AA) cases, and 508 AA controls); TNN-KIAA0040 may be associated with AD | [56] |
| 23 | A quantitative-trait genome-wide association study of alcoholism risk in the community: findings and implications. | GWAS | 3393 Australians with genome-wide SNP data; ANKS1A and TMEM108 may be associated with AD | [57] |
| 24 | Genome-wide association study of theta band event-related oscillations identifies serotonin receptor gene HTR7 influencing risk of alcohol dependence. | GWAS | 1,064 unrelated individuals, ARID5A, HTR7 may be associated with event-related brain oscillations | [59] |
| 25 | Genome-wide association study of alcohol dependence implicates a region on chromosome 11. | GWAS | Collaborative Study on the Genetics of Alcoholismsample; SLC22A18, PHLDA2, NAP1L4, SNORA54, CARS, and OSBPL5 may be associated with AD | [60] |
| 26 | A genomewide association study of nicotine and alcohol dependence in Australian and Dutch populations. | GWAS | 1224 cases and 1162 controls plus comorbid AD and ND, 599 cases and 488 controls; ARHGAP10, MARK1, DDX6, KIAA1409 may be associated with AD or ND | [61] |
| 27 | Genome-wide association study of alcohol dependence | GWAS | 487 male inpatients with alcohol dependence, 1358 population-based control individuals; follow-up study included 1024 male inpatients and 996 age-matched male controls. CDH13 and ADH1C may be involved in AD | [62] |
The table is a resumen of the studies analyzed to extract the initial subset of genes that showed a possible association with alcohol dependence phenotypes in last 10 years. AD: Alcohol-dependence; ND: Nicotine-dependence COGA: Collaborative Study on the Genetics of Alcoholism (http://www.niaaa.nih.gov/research/major-initiatives/collaborative-studies-genetics-alcoholism-coga-study) SAGE: Study of Addiction: Genes and Environment (http://www.ncbi.nlm.nih.gov/projects/gap/cgi-bin/study.cgi?study_id=phs000092.v1.p1)
Table 4: Studies analyzed. The GWAS studies were used to obtain the list of genes for prioritizing the molecular cascades to be analyzed. The other articles provided material for the background and discussion.
Imputation
Imputation was run for the genes under analysis. The CEU HapMap 1000 genomes served for the analysis. From the original (not pruned) set of 3473 SNPs from the pathways under analysis we obtained 2208 SNPs that passed the imputation quality control (info>0.9) and the pruning (r2>0.2). Pruning was undertaken after imputation.
Statistical methods
Covariated linear regression was the statistical model for the analysis. Plink served for the analysis [22]. We analyzed the results organized as pathways comparing the distribution of the p values<0.05 (of association with the phenotype under analysis) between each pathway subset of SNPs and an equal number of SNPs association p values randomly chosen from the genome. Fisher exact test was the statistical method for the analysis. We then permuted these p values randomly re-assigning the SNPs in the two groups 100000 times for each pathway. The resulting permuted p for each pathway is a strong association p calculated on the frequency that one of these random groups (with the same number of SNPs of the pathway) could reach a significance level equal or stronger than the pathway. Table 2 supplementary materials reports the statistics of all the SNPs of all the pathways.
Results
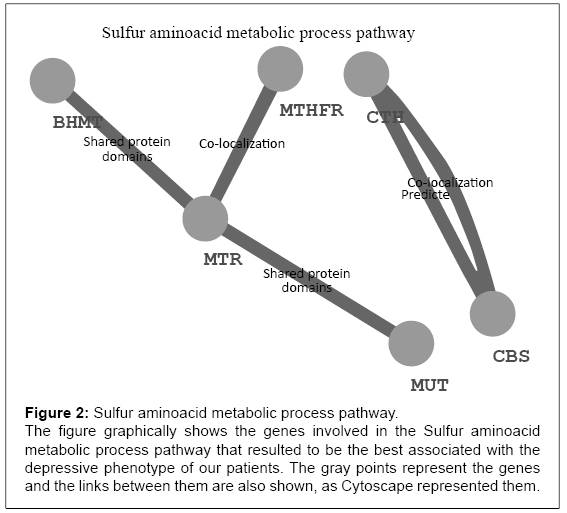
Table 1 reports the characteristics of the sample under analysis. Briefly, as sociodemographic variables, gender but not age resulted to be correlated with alcohol dependence record (p = 0.001). We also found an impact of alcohol dependence on an higher frequency of severe manic (T=-2.3, p=0.02) and depressive (T=-3.4, p=0.0006) phases (Table 1). The molecular pathways associated with the number of manic and depressive records corrected for the times patients were assessed during the period of observation are reported in Table 5 and 6. None of these pathways survived neither the Bonferroni nor the False Discovery Rate correction for multitesting. The sulfur amino acid metabolic process (GO:0000096) almost reached the threshold for significance (p<0.0003). Figure 2 reports the pathway representation.
| Molecular pathway | Permutated p value | False discovery rate q value | Activity |
|---|---|---|---|
| GO:0022838 | 0.001 | 0.18 | Substrate-specific channel activity |
| GO:0005216 | 0.018 | 1 | Ion channel activity |
| GO:0015837 | 0.029 | 1 | Amine transport |
| GO:0007210 | 0.034 | 1 | Serotonin receptor signaling pathway |
| GO:0015267 | 0.040 | 1 | Channel activity |
The table shows the results of the association between the pathways under analysis and the manic phenotype of our patients.
Table 5: Pathways associated with manic phases.
| Molecular pathway | Permutated p value | False discovery rate q value | Activity |
|---|---|---|---|
| GO:0000096 | 0.0007 | 0.10 | Sulfur aminoacid metabolic process |
| GO:0019932 | 0.0053 | 0.40 | Second-messenger-mediated signaling activity |
| GO:0016054 | 0.0085 | 0.43 | Organic acid catabolic process |
| GO:0009069 | 0.0120 | 0.45 | Serine family aminoacid process |
| GO:0045121 | 0.0153 | 0.46 | glycolipid-enriched membrane domain |
| GO:0022834 | 0.0196 | 0.49 | Ligand-gated channel activity |
The table shows the results of the association between the pathways under analysis and the depressive phenotype of our patients.
Table 6: Pathways associated with depressive phases.
Figure 2: Sulfur aminoacid metabolic process pathway.
The figure graphically shows the genes involved in the Sulfur aminoacid
metabolic process pathway that resulted to be the best associated with the
depressive phenotype of our patients. The gray points represent the genes
and the links between them are also shown, as Cytoscape represented them.
Discussion
Bipolar Disorder is a severe psychiatric disease that further worsens when it is associated with alcohol comorbidity [4]. Costs for society, patients and their relatives are impressive and chronic [10]. Pharmacological treatments are only partially effective also because the biological causes of both disorders and their connections are only partially known. In the present paper we investigated the genetic liability to depressive and manic records of BD patients with alcohol dependence in a sample of 802 bipolar patients. The STEP-BD was an open labeled study and patients were assessed according to the clinical choices. We then corrected the number of manic and depressive records for the times patients were assessed during the period of observation in order to correct the possible bias more severe patients being assessed more frequently compared to less severe patients. Sociodemographic variables entered the study as usual in this kind of investigation. Other Authors investigated this issue before [23] but with a smaller sample (n=278) and with a strict candidate gene approach. The limit of such approach is that it adds or detracts evidence from the theory it starts from, but it lacks in providing clues to new theories. This approach was reversed some years ago when the first platforms for the interrogation of the whole genome became available [24,25]. In the present paper we took advantage of the evidence gathered from this latter approach. Genes were selected starting from an atheoretical point of view (previous GWAS studies in literature) in order to maximize the chances to identify all the variations possibly associated with BD and alcohol dependence yet associated in literature. Moreover, the possible candidate genes associated with these disorders were further enriched taking advantage from the known metabolic cascades in which the first identified genes are embedded. This strategy is not without limits:
1. Pathway analyses are based on the current knowledge on the genes, their products and their interaction. This knowledge is incomplete at its best. Known genes could code for unknown proteins. Unknown genes could be yet discovered. Thus, the findings reported in the present paper are to be considered preliminary.
2. Longer genes are more likely to harbour more variations associated with the phenotype under analysis compared with shorter genes. We tried to balance this possible caveat by selecting random pathways of the same exact length of the index pathway under analysis.
3. Genes contain regulatory, exonic and intronic variations. Variations may be in LD with each other. Moreover, epistatic control is also possible. This genetic scenario is difficult to be taken into account in a statistical algorithm. This is a true limit of our analysis.
4. A high number of statistical tests are needed to compare the high number of genes that are embedded in a molecular cascade. As a consequence, false positive findings are a true concern. We tried to overcome this possible caveat by permutation. Nevertheless, independent research is needed to confirm our results.
Another limit of this study is the absence of comparison between BD patients and healthy controls because authors investigated the STEPBD database that only includes bipolar patients. Moreover in STEP-BD database a record of alcohol dependence was available only referred to last 12 months and not life trough. Finally, our study does not take into account the epigentic mechanisms of control of genome including for example the methylation rates and the activity of iRNA. These molecular mechanisms may deeply impact on the genetic makeup, covering the effects of the genetic variations or stressing their role towards the final phenotype and metabolomic studies could help to clarify changes in the metabolic pathways related to the SNPs variations effect. Despite these are true concerns of our analysis, we had an interesting finding for a pathway related to the Sulfur aminoacid metabolic process (GO:0000096) that includes methionine, cysteine, homocysteine, and taurine. Variations in this pathway genes resulted to be associated with a higher number of severe depressive records during the period of observation for patients that showed a record of alcohol dependence in past. Methionine is among the most hydrophobic of the amino acids and it has antioxidant properties, a proposed mechanism for depression [26]. Moreover, its related compound the S-Adenosylmethionine is the methyl donor for the methylation process that also includes the DNA and RNA silencing process. Thus its role in controlling the cell life is pivotal. Vitamins are required for methionine metabolism, and methionine metabolism plays a crucial role in the cellular assimilation of folate. Intriguingly, a vitamine and folate deficiency is a typical finding during alcohol dependence, suggesting that the combination of alcohol misbehavior and the oxidative stress or the DNA deregulation that is caused by the alteration of the sulfur aminoacid balance may precipitate depression in bipolar patients. Cysteine is semi-essential amino acid has known has antioxidant properties, a proposed mechanism for depression [26], and it has been proposed as a preventative or antidote for some of the negative effects of alcohol, including liver damage and hangover [27]. This is of particular relevance and it is of particular consistency when investigating depressive phenotypes in BD patients with alcohol dependence. Consistently, it was shown in animal models that blocking the cystein system results in a liability to exogenous stress [28], and epidemiological studies in humans would suggest that alcohol dependence is more frequent in subjects exposed to higher levels of stress, especially during childhood [29,30]. Thus, our result would suggest that a genetic liability in the Sulfur aminoacidic metabolic process would expose bipolar subjects to a higher sensitivity to stress and to a higher likelihood to treat it with alcohol dependence during the early stages. Of note this would result in a worse response to treatment in an open label environment. Taurine may play a role as well. It acts as an antioxidant and as an intracellular osmolyte, a membrane stabilizer, and a neurotransmitter [31]. In particular, it has been proposed that taurine may be involved in the mechanisms of action of valproic acid, a stronghold in the treatment of bipolar disorder [32]. Moreover, serine which is found in the same pathway as cysteine, is a potent agonist at the glycine site of the NMDA-type glutamate receptor, where glycine plays an inhibitory role. Glutamate is the main excitatory mediator of the central nervous system. It is then tempting to assume that an number of genetic variations located in this pathway able to disrupt its balance may interfere with the activation of some relevant neuronal networks. This may play a role in the pathological mood swings that characterize BD and may be triggered or worsened by the alcohol dependence. Finally, as shown in Table 5, we had some suggestive evidence that the control of mood in BD patients with alcohol dependence was driven by molecular pathways related to the permeability of membranes (GO:0022838 (substrate-specific channel activity), GO:0005216 (ion channel activity) and GO:0015267 (channel activity )) and to the aminergic tone (GO:0015837 (amine transport) and GO:0007210 (serotonin receptor signaling pathway)). Intriguingly, this is also related to the pharmacodynamics of mood stabilizers and to the second-generation antipsychotics, which are commonly prescribed [33] to treat manic phases and associated with several adverse effects related with pathways in which these activities can play role [34]. In conclusion, despite the limitations of our study, we had a marginal finding that point at a molecular cascade worthy of further investigation by independent groups. The main finding of our study is that the inconsistency that is found in literature when a single phenotype is investigated by different groups of research that look for single genes or single variations can be better understood if complete molecular pathways are taken into consideration [63].
Acknowledgements
We thank the NIMH for having had the possibility of analyzing their data on the STEP-BD sample. We also thank the Authors of previous publications in this dataset and foremost we thank the patients and their families who accepted to be enrolled in the study. Data and biomaterials were collected for the Systematic Treatment Enhancement Program for Bipolar Disorder (STEP-BD), a multi-center, longitudinal (5-8 year) project selected from responses to RFP #NIMH-98-DS-0001, “Treatment for Bipolar Disorder.” The project was led by Gary Sachs, M.D., and coordinated by Massachusetts General Hospital in Boston, MA. The NIMH grant number was 2N01MH080001-001. Given the major public health implications of identifying genes responsible for severe neuropsychiatric disorders, the National Institute of Mental Health (NIMH) has funded a Human Genetics Initiative. The goal of this Initiative is to establish a national resource of clinical data and biomaterials that are collected from individuals with Alzheimer disease, schizophrenia, or bipolar I disorder (BP), in order to aid researchers in understanding the genetic bases of these disorders. The NIMH Bipolar Disorder Genetics Initiative is supported by the Office of Human Genetics & Genomic Resources in NIMH’s Division of Neuroscience and Basic Behavioral Science (DNBBS). Since 1996, data and biomaterials (cell lines and DNA samples) have been available to qualified investigators who study the genetics of BP, and may be accessed by following a set of instructions.
References
- Farren CK, Hill KP, Weiss RD (2012) Bipolar disorder and alcohol use disorder: a review. Curr Psychiatry Rep 14: 659-666.
- Kessler RC, Chiu WT, Demler O, Merikangas KR, Walters EE (2005) Prevalence, severity, and comorbidity of 12-month DSM-IV disorders in the National Comorbidity Survey Replication. Arch Gen Psychiatry 62: 617-627.
- Hasin DS, Goodwin RD, Stinson FS, Grant BF (2005) Epidemiology of major depressive disorder: results from the National Epidemiologic Survey on Alcoholism and Related Conditions. Arch Gen Psychiatry 62: 1097-1106.
- Frye MA, Salloum IM (2006) Bipolar disorder and comorbid alcoholism: prevalence rate and treatment considerations. Bipolar Disord 8: 677-685.
- Judd LL, Akiskal HS (2003) The prevalence and disability of bipolar spectrum disorders in the US population: re-analysis of the ECA database taking into account subthreshold cases. J Affect Disord 73: 123-131.
- Kleinman L, Lowin A, Flood E, Gandhi G, Edgell E, et al. (2003) Costs of bipolar disorder. Pharmacoeconomics 21: 601-622.
- Friedmann PD, Saitz R, Gogineni A, Zhang JX, Stein MD (2001) Validation of the screening strategy in the NIAAA "Physicians' Guide to Helping Patients with Alcohol Problems". J Stud Alcohol 62: 234-238.
- Harwood HJ, Fountain D, Fountain G (1999) Economic cost of alcohol and drug abuse in the United States, 1992: a report. Addiction 94: 631-635.
- Mokdad AH, Marks JS, Stroup DF, Gerberding JL (2004) Actual causes of death in the United States, 2000. JAMA 291: 1238-1245.
- Hirschfeld RM, Vornik LA (2005) Bipolar disorder--costs and comorbidity. Am J Manag Care 11: S85-90.
- Grant BF, Dawson DA, Stinson FS, Chou SP, Dufour MC, et al. (2004) The 12-month prevalence and trends in DSM-IV alcohol abuse and dependence: United States, 1991-1992 and 2001-2002. Drug Alcohol Depend 74: 223-234.
- Regier DA, Farmer ME, Rae DS, Locke BZ, Keith SJ, et al. (1990) Comorbidity of mental disorders with alcohol and other drug abuse. Results from the Epidemiologic Catchment Area (ECA) Study. JAMA 264: 2511-2518.
- Berk M, Hallam K, Malhi GS, Henry L, Hasty M, et al. (2010) Evidence and implications for early intervention in bipolar disorder. J Ment Health 19: 113-126.
- Walters GD (2002) The heritability of alcohol abuse and dependence: a meta-analysis of behavior genetic research. Am J Drug Alcohol Abuse 28: 557-584.
- McGuffin P, Rijsdijk F, Andrew M, Sham P, Katz R, et al. (2003) The heritability of bipolar affective disorder and the genetic relationship to unipolar depression. Arch Gen Psychiatry 60: 497-502.
- Kimura M, Higuchi S (2011) Genetics of alcohol dependence. Psychiatry ClinNeurosci 65: 213-225.
- Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, et al. (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13: 2498-2504.
- Sachs GS, Thase ME, Otto MW, Bauer M, Miklowitz D, et al. (2003) Rationale, design, and methods of the systematic treatment enhancement program for bipolar disorder (STEP-BD). Biol Psychiatry 53: 1028-1042.
- Young RC, Biggs JT, Ziegler VE, Meyer DA (1978) A rating scale for mania: reliability, validity and sensitivity. Br J Psychiatry 133: 429-435.
- Williams JB, Kobak KA (2008) Development and reliability of a structured interview guide for the Montgomery Asberg Depression Rating Scale (SIGMA). Br J Psychiatry 192: 52-58.
- Montgomery SA, Asberg M (1979) A new depression scale designed to be sensitive to change. Br J Psychiatry 134: 382-389.
- Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, et al. (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81: 559-575.
- Yasseen B, Kennedy JL, Zawertailo LA, Busto UE (2010) Comorbidity between bipolar disorder and alcohol use disorder: association of dopamine and serotonin gene polymorphisms. Psychiatry Res 176: 30-33.
- Visscher PM, Brown MA, McCarthy MI, Yang J (2012) Five years of GWAS discovery. Am J Hum Genet 90: 7-24.
- Sullivan PF (2010) The psychiatric GWAS consortium: big science comes to psychiatry. Neuron 68: 182-186.
- Behr GA, Moreira JC, Frey BN (2012) Preclinical and clinical evidence of antioxidant effects of antidepressant agents: implications for the pathophysiology of major depressive disorder. Oxid Med Cell Longev 2012: 609421.
- Bosch-Morell F, MartÃnez-Soriano F, Colell A, Fernández-Checa JC, Romero FJ (1998) Chronic ethanol feeding induces cellular antioxidants decrease and oxidative stress in rat peripheral nerves. Effect of S-adenosyl-L-methionine and N-acetyl-L-cysteine. Free RadicBiol Med 25: 365–368.
- Sienko M, Natorff R, Skoneczny M, Kruszewska J, Paszewski A, et al. (2014) Regulatory mutations affecting sulfur metabolism induce environmental stress response in Aspergillusnidulans. Fungal Genet Biol 65: 37-47.
- Blanco C, Xu Y, Brady K, Pérez-Fuentes G, Okuda M, et al. (2013) Comorbidity of posttraumatic stress disorder with alcohol dependence among US adults: results from National Epidemiological Survey on Alcohol and Related Conditions. Drug Alcohol Depend 132: 630–638.
- Brady KT, Back SE (2012) Childhood trauma, posttraumatic stress disorder, and alcohol dependence. Alcohol Res 34: 408-413.
- Kumari N, Prentice H, Wu JY (2013) Taurine and its neuroprotective role. AdvExp Med Biol 775: 19-27.
- Anyanwu E, Harding GF (1993) The involvement of taurine in the action mechanism of sodium valproate (VPA) in the treatment of epilepsy. Actaphysiologica, pharmacologica et therapeuticalatinoamericana: órgano de la AsociaciónLatinoamericana de CienciasFisiológicas y [de] la AsociaciónLatinoamericana de FarmacologÃa 43: 20–27.
- Scherk H, Pajonk FG, Leucht S (2007) Second-generation antipsychotic agents in the treatment of acute mania: a systematic review and meta-analysis of randomized controlled trials. Arch Gen Psychiatry 64: 442–455.
- Cocchi E, Drago A, de Ronchi D, Serretti A (2014) The genetics of vascular incidents associated with second-generation antipsychotic administration. Expert Rev ClinPharmacol 7: 75-90.
- Dick DM, Bierut LJ (2006) The genetics of alcohol dependence. Curr Psychiatry Rep 8: 151-157.
- Gelernter J, Kranzler HR2, Sherva R3, Almasy L4, Koesterer R3, et al. (2014) Genome-wide association study of alcohol dependence:significant findings in African- and European-Americans including novel risk loci. Mol Psychiatry 19: 41-49.
- Wetherill L, Kapoor M, Agrawal A, Bucholz K, Koller D, et al. (2014) Family-based association analysis of alcohol dependence criteria and severity. Alcohol ClinExp Res 38: 354-366.
- Wang S, Yang Z, Ma JZ, Payne TJ, Li MD (2014) Introduction to deep sequencing and its application to drug addiction research with a focus on rare variants. MolNeurobiol 49: 601-614.
- McGue M, Zhang Y, Miller MB, Basu S, Vrieze S, et al. (2013) A genome-wide association study of behavioral disinhibition. Behav Genet 43: 363-373.
- Seneviratne C1, Franklin J, Beckett K, Ma JZ, Ait-Daoud N, et al. (2013) Association, interaction, and replication analysis of genes encoding serotonin transporter and 5-HT3 receptor subunits A and B in alcohol dependence. Hum Genet 132: 1165-1176.
- Zhang Z, Wang JC, Howells W, Lin P, Agrawal A, et al. (2013) Dosage transmission disequilibrium test (dTDT) for linkage and association detection. PLoS One 8: e63526.
- Kapoor M1, Wang JC, Wetherill L, Le N, Bertelsen S, et al. (2013) A meta-analysis of two genome-wide association studies to identify novel loci for maximum number of alcoholic drinks. Hum Genet 132: 1141-1151.
- Kos MZ, Yan J, Dick DM, Agrawal A, Bucholz KK, et al. (2013) Common biological networks underlie genetic risk for alcoholism in African- and European-American populations. Genes Brain Behav 12: 532-542.
- Biernacka JM, Geske JR, Schneekloth TD, Frye MA, Cunningham JM, et al. (2013) Replication of genome wide association studies of alcohol dependence: support for association with variation in ADH1C. PLoS One 8: e58798.
- Park BL, Kim JW, Cheong HS, Kim LH, Lee BC, et al. (2013) Extended genetic effects of ADH cluster genes on the risk of alcohol dependence: from GWAS to replication. Hum Genet 132: 657-668.
- Zhao Z, Guo AY, van den Oord EJ, Aliev F, Jia P, et al. (2012) Multi-species data integration and gene ranking enrich significant results in an alcoholism genome-wide association study. BMC Genomics 13 Suppl 8: S16.
- Yan J, Aliev F, Webb BT, Kendler KS, Williamson VS, et al. (2013) Using genetic information from candidate gene and genome-wide association studies in risk prediction for alcohol dependence. Addict Biol .
- Nishizawa D, Fukuda K2, Kasai S1, Hasegawa J1, Aoki Y3, et al. (2014) Genome-wide association study identifies a potent locus associated with human opioid sensitivity. Mol Psychiatry 19: 55-62.
- Wang JC, Foroud T, Hinrichs AL, Le NX, Bertelsen S, et al. (2013) A genome-wide association study of alcohol-dependence symptom counts in extended pedigrees identifies C15orf53. Mol Psychiatry 18: 1218-1224.
- Blaine S, Claus E, Harlaar N, Hutchison K (2013) TACR1 genotypes predict fMRI response to alcohol cues and level of alcohol dependence. Alcohol ClinExp Res 37 Suppl 1: E125-130.
- Chen XD, Xiong DH, Yang TL, Pei YF, Guo YF, et al. (2012) ANKRD7 and CYTL1 are novel risk genes for alcohol drinking behavior. Chin Med J (Engl) 125: 1127-1134.
- Agrawal A, Wetherill L, Bucholz KK, Kramer J, Kuperman S, et al. (2013) Genetic influences on craving for alcohol. Addict Behav 38: 1501-1508.
- Zuo L , Zhang CK, Wang F, Li CS, Zhao H, et al. (2011) A novel, functional and replicable risk gene region for alcohol dependence identified by genome-wide association study. PLoS One 6: e26726.
- Contini V, Bertuzzi GP, Polina ER, Hunemeier T, Hendler EM, et al. (2012) A haplotype analysis is consistent with the role of functional HTR1B variants in alcohol dependence. Drug Alcohol Depend 122: 100-104.
- Frank J, Cichon S, Treutlein J, Ridinger M, Mattheisen M, et al. (2012) Genome-wide significant association between alcohol dependence and a variant in the ADH gene cluster. Addict Biol 17: 171-180.
- Zuo L, Gelernter J, Zhang CK, Zhao H, Lu L, et al. (2012) Genome-wide association study of alcohol dependence implicates KIAA0040 on chromosome 1q. Neuropsychopharmacology 37: 557-566.
- Heath AC, Whitfield JB, Martin NG, Pergadia ML, Goate AM, et al. (2011) A quantitative-trait genome-wide association study of alcoholism risk in the community: findings and implications. Biol Psychiatry 70: 513-518.
- Treutlein J, Rietschel M (2011) Genome-wide association studies of alcohol dependence and substance use disorders. Curr Psychiatry Rep 13: 147-155.
- Zlojutro M, Manz N, Rangaswamy M, Xuei X, Flury-Wetherill L, et al. (2011) Genome-wide association study of theta band event-related oscillations identifies serotonin receptor gene HTR7 influencing risk of alcohol dependence. Am J Med Genet B Neuropsychiatr Genet 156B: 44–58.
- Edenberg HJ, Koller DL, Xuei X, Wetherill L, McClintick JN, et al. (2010) Genome-wide association study of alcohol dependence implicates a region on chromosome 11. Alcohol ClinExp Res 34: 840-852.
- Lind PA, Macgregor S, Vink JM, Pergadia ML, Hansell NK, et al. (2010) Agenomewide association study of nicotine and alcohol dependence in Australian and Dutch populations. Twin Res Hum Genet 13: 10-29.
- Treutlein J, Cichon S, Ridinger M, Wodarz N, Soyka M, et al. (2009) Genome-wide association study of alcohol dependence. Arch Gen Psychiatry 66: 773-784.
- Tian C, Stokowski RP, Kershenobich D, Ballinger DG, Hinds DA (2010) Variant in PNPLA3 is associated with alcoholic liver disease. Nat Genet 42: 21-23.
Relevant Topics
- Addiction Recovery
- Alcohol Addiction Treatment
- Alcohol Rehabilitation
- Amphetamine Addiction
- Amphetamine-Related Disorders
- Cocaine Addiction
- Cocaine-Related Disorders
- Computer Addiction Research
- Drug Addiction Treatment
- Drug Rehabilitation
- Facts About Alcoholism
- Food Addiction Research
- Heroin Addiction Treatment
- Holistic Addiction Treatment
- Hospital-Addiction Syndrome
- Morphine Addiction
- Munchausen Syndrome
- Neonatal Abstinence Syndrome
- Nutritional Suitability
- Opioid-Related Disorders
- Relapse prevention
- Substance-Related Disorders
Recommended Journals
Article Tools
Article Usage
- Total views: 15535
- [From(publication date):
April-2014 - Aug 17, 2025] - Breakdown by view type
- HTML page views : 10836
- PDF downloads : 4699