Editorial Open Access
APE1: A Molecule of Focus with Neuroprotective and Anti-Cancer Properties
Anil K Mantha1,2*1Center for Biosciences, School of Basic and Applied Sciences, Central University of Punjab, Bathinda, Punjab, India, Pin Code: 151 001
2Department of Biochemistry and Molecular Biology, University of Texas Medical Branch, Galveston, TX-77555, USA (Adjunct Assistant Professor)
- Corresponding Author:
- Dr. Anil K. Mantha
Assistant Professor
Center for Biosciences
School of Basic and Applied Sciences
Central University of Punjab
Bathinda, 151 001, Punjab, India
E-mail: anilmantha@gmail.com; Anil.kumar@cup.ac.in
Received date: June 25, 2013; Accepted date: June 26, 2013; Published date: June 28, 2013
Citation: Mantha AK (2013) APE1: A Molecule of Focus with Neuroprotective and Anti-Cancer Properties. J Biotechnol Biomater 3:e120. doi:10.4172/2155-952X.1000e120
Copyright: © 2013 Mantha AK. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Journal of Biotechnology & Biomaterials
Abstract
Apurinic/Apyrimidinic endonuclease (APE1) is a multi-functional, central enzyme of base excision repair (BER) pathway that takes care of oxidized base damage (AP sites and strand breaks) caused by both endogenous and exogenous oxidative DNA damaging agents. In repair function, APE1 exhibits majorly abasic (AP) endonuclease activity and stable interaction(s) with BER-pathway participant proteins. Second function of APE1 is redox activation of various transcription factors (TFs e.g., c-jun, NF-kB, p53 and HIF1α) and also named as redox effector factor 1(Ref-1). In redox function, APE1 reductively activates TFs involved in regulation of gene expression for cell survival mechanisms through stable pair-wise interaction(s). Recent studies have indicated that APE1 also possesses other distinct functions such as RNA metabolism, riboendonuclease activity and protein-protein interaction for maintaining cellular homeostasis. Altered APE1 expression has been reported in various cancers and neurodegenerative diseases. Taken together such findings advocates the necessity to delineate the underlying molecular mechanism(s) for understanding its role in various biological functions, that could be translated to its application in therapeutics against human diseases like cancer, neurodegenerative diseases and other pathologies such as cardiovascular diseases.
Abbreviations
APE1, apurinic/apyrimidinic endonuclease; BER, base excision repair; Ref-1, redox effector factor 1; ROS, reactive oxygen species; TF, transcription factor; HIF-1α, hypoxia inducible factor 1 alpha; NLS, nuclear localization signal; AD, Alzheimer’s disease; PD, Parkinson’s disease; ALS, amyotrophic lateral sclerosis; AP sites, Apurinic/Apyrimidinic sites; AP-1: Activator Protein 1; Aβ, amyloid beta; PARP-1, poly [ADP-ribose] polymerase 1; STAT3, signal transducer activator of transcription 3; WRN, Werner syndrome helicase; YB-1, Y-box-binding protein 1; XRCC-1, X-ray repair crosscomplementing protein 1; and SSBs, single strand breaks
Introduction
DNA damage/lesions caused by oxidizing/alkylating exogenous agents (ionizing radiation, pesticides, and chemotherapeutic agents) and endogenous agents (metabolites, oxidative phosphorylation, and free radicals) are implicated in several human pathologies including cancer and neurodegenerative diseases [1]. BER-pathway is the primary repair system against these DNA lesions [2]. Two types of BER sub-pathways exists; short-patch and long-patch BER takes place in mammalian cells depending upon the type of damage, concentration of the participant BER protein(s) and thus, finally allowing the replacement of the damaged DNA gap by single nucleotide and multinucleotides respectively [3]. APE1 is a central enzyme of BER-pathway with primary two major independent cellular functions; repair and redox [4-6]. Both functions are displayed by two different regions of the protein. Primarily, repair function through C-terminus, which involves removal of AP sites in DNA. Second function is reductive activation of several TFs displayed via disordered N-terminus which also possesses the Nuclear Localization Signal (NLS), controlling gene expression for significant cellular pathways [6,7]. Because of redox activity, APE1 is also known as redox effector factor-1 (Ref-1). It was found to be involved in reductive activation of TFs like c-Jun via a thiol exchange reaction involving Cys65 residue [8]. Later on, it was also demonstrated to reductively activate p53 [9], PAX-8 [10], AP-1 (fos/jun) [11], PAX- 5 [12], HIF-1α and NF-κB by same redox-based process involving reduction of key Cys residues in TF’s DNA–binding domains, thus increasing their DNA-binding affinity [4,6,13]. All these TF’s regulates the transcription of various genes responsible for cell survival pathways. Thus, APE1 regulates the pathways that are involved in normal as well as cancer cell survival.
Now it is well-known that APE1 is essential for cell survival [14]. Studies have indicated that altered APE1 expression is associated with various cancers and neurodegenerative diseases such as Alzheimer’s disease (AD), Parkinson’s disease (PD) and Amyotrophic Lateral Sclerosis (ALS) [15-18]. APE1 overexpression has been linked with chemoresistance in cancer cells [19,20]. In neuronal cells, it provides neuroprotection against oxidative assault [21,22]. It is known that not only oxidatively damaged DNA, but also oxidatively damaged RNA (involving mRNA, rRNA and tRNA) has been suggested to be associated with neurodegenerative diseases [23]. Recently, APE1’s role in RNA metabolism including its function as riboendonuclease has been identified, pointing to have role in post-transcriptional regulation of gene expression [24,25]. Therefore, it can be stated that APE1 and its distinct functions in repair, redox as well as in RNA metabolism (Figure 1) can be employed for possible candidate for anti-cancer and as a neuroprotection molecule for better therapies.
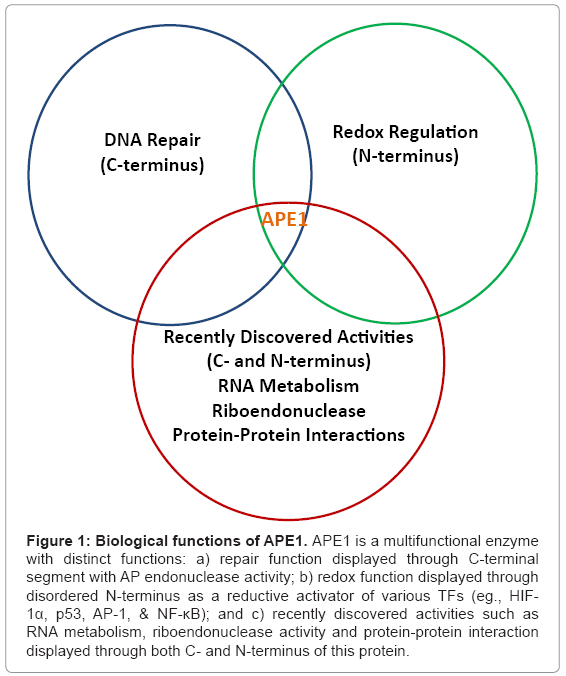
Figure 1: Biological functions of APE1. APE1 is a multifunctional enzyme with distinct functions: a) repair function displayed through C-terminal segment with AP endonuclease activity; b) redox function displayed through disordered N-terminus as a reductive activator of various TFs (eg., HIF- 1α, p53, AP-1, & NF-κB); and c) recently discovered activities such as RNA metabolism, riboendonuclease activity and protein-protein interaction displayed through both C- and N-terminus of this protein.
APE1 as a Therapeutic Target
An Anti-Cancer Agent
Main treatment options for cancer includes chemotherapy and radiotherapy which exhibits their cytotoxic effects via inducing DNA damage in cancerous cells. Likewise normal healthy cells, cancerous cells also tend to repair such DNA damage via BER-pathway exploiting DNA repair protein, APE1. Therefore, obvious approach is to inhibit APE1 thereby reducing DNA-damage repair in cancerous cells. This will help to combat chemoresistance and to potentiate the cytotoxic effects of chemotherapeutic agents [26]. Inhibition of both DNA repair function, as well as redox function can act as a rational cancer target [27]. Data from various studies have shown that APE1 overexpression is responsible for chemoresistance in cancer cells and blocking APE1 activities have helped to sensitize cancerous cell against chemotherapeutic agents such as Bleomycin, Temozolomide (TMZ), and Methyl Methane Sulfonate (MMS) [28-30]. Some studies involving use of specific small-molecule inhibitors have tried to understand the APE1’s individual causative function responsible for cancer cell survival and proliferation [31,32]. APE1’s endonuclease activity was also found to be significantly higher in advanced tumors as compared to lower grade tumors [29]. Currently, it is clear that both activities of APE1 play an important role in cancer cell survival and proliferation, thus research involving identification of inhibitor molecules of APE1 repair and redox function are required for development of new cancer therapeutics.
A Neuroprotective Agent
Given the fact that brain is more vulnerable to oxidative DNA damage and neurons do not regenerate readily, it becomes important to find ways for neuroprotection [33,34]. Manipulating BER-pathway by altering APE1 expression and functions in neuronal cells can help in preventing neurotoxicity caused by environmental agents (radiation) and abnormal accumulation of metabolites-mediated oxidative DNA damage [35]. Overexpression of APE1 and increased APE1’s activities might help in neuroprotection by switching-on the cell survival mechanisms. In a study performed on nuclear extracts of brain of AD patients, increased APE1 expression was observed [16]. Another immuno histochemical study reported elevated nuclear APE1 expression in cerebral cortex of AD patients [35]. Elevated APE1 expression was also observed in brain and spinal cord samples of ALS patients [36]. These finding advocates for role of elevated APE1 expression in maintaining neuronal cell survival in response to oxidative stress. It was also found that APE1 is able to protect Dorsal Root Ganglion (DRG) cells from IR-induced neurotoxicity, through its repair activity [22]. To address the issue of responsible APE1 individual activity behind neuroprotection, various studies have been conducted. In another in-vitro study conducted on SH-SY5Y cell line, it was demonstrated that APE1’s repair function provides neuroprotection and promotes cell survival after oxidative DNA damage [37]. Recently Mantha et al. through 2-D proteomics using PC12 and SH-SY5Y cells have identified some APE1 interacting proteins associated with neuronal cell survival pathways suggesting role of APE1 in neuroprotection against oxidative damage caused by Aβ-toxicity that is observed in case of AD [38].
Recent Discoveries in APE1 Field
APE1 as a Riboendonuclease
Riboendonucleases hold significance by the fact that they control mRNA degradation and hence, controls gene expression [39,40]. To date, many riboendonucleases have been identified and still, many are remaining [41,42]. Recently APE1 has been identified as a riboendonuclease that cleaves and regulates c-myc mRNA [25]. It was demonstrated that APE1 cleaves c-myc Coding Region Determinant (CRD) RNA at UA, UG and CA sites, and they also showed that APE1 controls c-myc mRNA levels as well as half life in cells [24,25,42]. Altogether these findings suggest the possible role of APE1 in regulating other mRNAs involved in various diseases.
In RNA Metabolism
From DNA-RNA-Protein, each step is prone to oxidative DNA damage and ultimately, can affect cell viability. Oxidatively damaged RNA can impair protein synthesis or can lead to inaccurate translation. Therefore, cell needs to cope with damaged RNA also; likewise it copes with damaged DNA. Studies have demonstrated the role of APE1 in RNA metabolism [25]. In an experiment, APE1-depleted cells showed accumulation of oxidized rRNA after oxidative stress, which suggests APE1’s role in cleansing oxidatively damaged RNA [25]. Recent study has indicated APE1’s ability to cleave AP site in single stranded RNA [43]. In an another study it has been demonstrated that APE1 interacts through N-terminal domain with nucleophosmin (NPM1) protein, a ribosome processing protein, that regulates rRNA metabolism in ribosome biogenesis [44]. APE1 also reported to interact with YB-1 and hnRNP-L, which are involved in RNA metabolism [45,46]. All together, these findings strongly suggests for possibility of APE1 to be an evolutionary protein with previously unknown functions that needs to be understood/revealed, and possibility to have role in miRNA metabolism [25].
Protein-Protein Interaction
APE1 displays binary interactions with participant protein(s) of BER-pathway as well as with proteins of other biologically relevant pathways involving TFs, so as to stabilize or stimulate each other’s functions [5,47]. These interactions play important role in maintenance of cell survival. In DNA repair, weakened interactions among the key downstream participant proteins of BER-pathway can lead to reduced repair efficiency [5,47]. Important enzymes of BER-pathway and TFs to which APE1 interacts are summarized in Table 1.
| Protein Pathway | Participant Proteins | Reference |
|---|---|---|
| BER-Pathway Proteins | • XRCC-1 • PARP-1 • OGG1 • pol β • DNA ligase I |
[48] [49] [50] [51] [52] |
| Redox Regulation of Key TFs | • NF-κB • HIF-1α • p53 • AP-1 • Pax-8 |
[53] [54] [9] [11] [10] |
Table 1: Important APE1 Protein-Protein Interaction.
In addition APE1 has been shown to be interacting or cross-talking with proteins such as heat shock protein 70 (hsp70) [55], WRN protein [56], STAT3 and p300 [57], YB-1 for activation of multidrug resistance (MDR) gene [58]. A recent study by Mantha et al. has addressed the role of APE1 and protein-protein interaction(s) in neuronal cell survival [38]. This study identified various APE1 interacting proteins upon Aβ(25- 35) stress in PC12 and SH-SY5Y cells, associated with stress-dependent events and neuronal cell survival pathways. Key proteins identified are: Pyruvate kinase 3 isoform 2 (PKM2), Tropomodulin 3 (Tmod3), and hetero genous nuclear ribonucleoprotein-H1 (hn-RNP-H) [38]. Future studies are required to unveil other APE1 and protein-protein interactions which may have key biologically significant functions.
Conclusions and Future Perspectives
APE1 is a key regulating enzyme of BER-pathway taking care of oxidative base damage resulting due to oxidative stress. Recent findings from various molecular studies have indicated association of differential APE1 expression pattern with various cancers and neurodegenerative disorders. It can be speculated that manipulating APE1 expression and function(s) can affect the underlying process of pathogenesis. In addition, recently discovered function of APE1 in RNA metabolism also broadens the perspectives of APE1’s exploitation as a therapeutic target for various diseases. In future, new approaches are needed to delineate the molecular mechanisms underlying APE1’s role in chemo/ radioresistance in cancer cells, and neuromodulatory role in neuronal cell survival. More proteomic studies are required to elucidate the specific APE1 protein-protein interaction in other biological pathways, in response to oxidative damage caused by exogenous and endogenous agents. Investigations elucidating the nature of molecular modifications (post-transcriptional and post-translational) and molecular signaling mechanisms regulating the functions of APE1 are need to be characterized in order to translate the APE1-mediated therapeutic interventions.
Acknowledgements
A.K.M is supported by Alzheimer’s Association, USA, NIRG-11-203527 grant. Thanks are to Ms. Shweta Thakur for assistance in drafting the article and to Dr. Monisha Dhiman, Centre for Genetic Diseases and Molecular Medicine, School of Emerging Life Science Technologies,Central University of Punjab, Bathinda for her critical comments and suggestions. Because of the short editorial/limited focus of the article, many relevant and appropriate references could not be included, for which the author sincerely apologize. The CUPB institutional number allocated for this manuscript is P-50.
References
- Hoeijmakers JH (2009) DNA damage, aging, and cancer. N Engl J Med 361: 1475-1485.
- Slupphaug G, Kavli B, Krokan HE (2003) The interacting pathways for prevention and repair of oxidative DNA damage. Mutat Res 531: 231-251.
- Fortini P, Dogliotti E (2007) Base damage and single-strand break repair: mechanisms and functional significance of short- and long-patch repair subpathways. DNA Repair (Amst) 6: 398-409.
- Bhakat KK, Mantha AK, Mitra S (2009) Transcriptional regulatory functions of mammalian AP-endonuclease (APE1/Ref-1), an essential multifunctional protein. Antioxid Redox Signal 11: 621-638.
- Hegde ML, Mantha AK, Hazra TK, Bhakat KK, Mitra S, et al. (2012) Oxidative genome damage and its repair: implications in aging and neurodegenerative diseases. Mech Ageing Dev 133: 157-168.
- Tell G, Quadrifoglio F, Tiribelli C, Kelley MR (2009) The many functions of APE1/Ref-1: not only a DNA repair enzyme. Antioxid Redox Signal 11: 601-620.
- Mol CD, Izumi T, Mitra S, Tainer JA (2000) DNA-bound structures and mutants reveal abasic DNA binding by APE1 and DNA repair coordination [corrected]. Nature 403: 451-456.
- Abate C, Patel L, Rauscher FJ 3rd, Curran T (1990) Redox regulation of fos and jun DNA-binding activity in vitro. Science 249: 1157-1161.
- Jayaraman L, Murthy KG, Zhu C, Curran T, Xanthoudakis S, et al. (1997) Identification of redox/repair protein Ref-1 as a potent activator of p53. Genes Dev 11: 558-570.
- Tell G, Pellizzari L, Cimarosti D, Pucillo C, Damante G (1998) Ref-1 controls pax-8 DNA-binding activity. Biochem Biophys Res Commun 252: 178-183.
- Xanthoudakis S, Miao G, Wang F, Pan YC, Curran T (1992) Redox activation of Fos-Jun DNA binding activity is mediated by a DNA repair enzyme. EMBO J 11: 3323-3335.
- Tell G, Zecca A, Pellizzari L, Spessotto P, Colombatti A, et al. (2000) An 'environment to nucleus' signaling system operates in B lymphocytes: redox status modulates BSAP/Pax-5 activation through Ref-1 nuclear translocation. Nucleic Acids Res 28: 1099-1105.
- Evans AR, Limp-Foster M, Kelley MR (2000) Going APE over ref-1. Mutat Res 461: 83-108.
- Izumi T, Brown DB, Naidu CV, Bhakat KK, Macinnes MA, et al. (2005) Two essential but distinct functions of the mammalian abasic endonuclease. Proc Natl Acad Sci U S A 102: 5739-5743.
- Al-Attar A, Gossage L, Fareed KR, Shehata M, Mohammed M, et al. (2010) Human apurinic/apyrimidinic endonuclease (APE1) is a prognostic factor in ovarian, gastro-oesophageal and pancreatico-biliary cancers. Br J Cancer 102: 704-709.
- Davydov V, Hansen LA, Shackelford DA (2003) Is DNA repair compromised in Alzheimer's disease? Neurobiol Aging 24: 953-968.
- Huang E, Qu D, Zhang Y, Venderova K, Haque ME, et al. (2010) The role of Cdk5-mediated apurinic/apyrimidinic endonuclease 1 phosphorylation in neuronal death. Nat Cell Biol 12: 563-571.
- Yoo DG, Song YJ, Cho EJ, Lee SK, Park JB, et al. (2008) Alteration of APE1/ref-1 expression in non-small cell lung cancer: the implications of impaired extracellular superoxide dismutase and catalase antioxidant systems. Lung Cancer 60: 277-284.
- Jeon BH, Irani K (2009) APE1/Ref-1: versatility in progress. Antioxid Redox Signal 11: 571-574.
- Wang D, Xiang DB, Yang XQ, Chen LS, Li MX, et al. (2009) APE1 overexpression is associated with cisplatin resistance in non-small cell lung cancer and targeted inhibition of APE1 enhances the activity of cisplatin in A549 cells. Lung Cancer 66: 298-304.
- Li P, Hu X, Gan Y, Gao Y, Liang W, et al. (2011) Mechanistic insight into DNA damage and repair in ischemic stroke: exploiting the base excision repair pathway as a model of neuroprotection. Antioxid Redox Signal 14: 1905-1918.
- Vasko MR, Guo C, Thompson EL, Kelley MR (2011) The repair function of the multifunctional DNA repair/redox protein APE1 is neuroprotective after ionizing radiation. DNA Repair (Amst) 10: 942-952.
- Nunomura A, Moreira PI, Takeda A, Smith MA, Perry G (2007) Oxidative RNA damage and neurodegeneration. Curr Med Chem 14: 2968-2975.
- Barnes T, Kim WC, Mantha AK, Kim SE, Izumi T, et al. (2009) Identification of Apurinic/apyrimidinic endonuclease 1 (APE1) as the endoribonuclease that cleaves c-myc mRNA. Nucleic Acids Res 37: 3946-3958.
- Tell G, Wilson DM 3rd, Lee CH (2010) Intrusion of a DNA repair protein in the RNome world: is this the beginning of a new era? Mol Cell Biol 30: 366-371.
- Zhu Y, Hu J, Hu Y, Liu W (2009) Targeting DNA repair pathways: a novel approach to reduce cancer therapeutic resistance. Cancer Treat Rev 35: 590-596.
- Bapat A, Fishel ML, Kelley MR (2009) Going ape as an approach to cancer therapeutics. Antioxid Redox Signal 11: 651-668.
- Bapat A, Glass LS, Luo M, Fishel ML, Long EC, et al. (2010) Novel small-molecule inhibitor of apurinic/apyrimidinic endonuclease 1 blocks proliferation and reduces viability of glioblastoma cells. J Pharmacol and Exp Ther 334: 988-998.
- Bobola MS, Blank A, Berger MS, Stevens BA, Silber JR (2001) Apurinic/apyrimidinic endonuclease activity is elevated in human adult gliomas. Clin Cancer Res 7: 3510-3518.
- Silber JR, Bobola MS, Blank A, Schoeler KD, Haroldson PD, et al. (2002) The apurinic/apyrimidinic endonuclease activity of Ape1/Ref-1 contributes to human glioma cell resistance to alkylating agents and is elevated by oxidative stress. Clin Cancer Res 8: 3008-3018.
- Jiang Y, Zhou S, Sandusky GE, Kelley MR, Fishel ML (2010) Reduced expression of DNA repair and redox signaling protein APE1/Ref-1 impairs human pancreatic cancer cell survival, proliferation, and cell cycle progression. Cancer Invest 28: 885-895.
- Luo M, Caldwell D, Xu Y, He Y, Reed A, et al. (2004) Inhibition of the human apurinic/apyrimidinic endonuclease DNA base excision repair enzyme/redox factor (APE1/Ref-1) using small molecule redox and repair inhibitors: Therapeutic implications. Experimental and Molecular Therapeutics 29: Novel Therapeutic Agents II.
- Floyd RA (1999) Antioxidants oxidative stress and degenerative neurological disorders. Experimental Biology and Medicine 222: 236-245.
- Bosshard M, Markkanen E, van Loon B (2012) Base excision repair in physiology and pathology of the central nervous system. Int J Mol Sci 13: 16172-16222.
- Marcon G, Tell G, Perrone L, Garbelli R, Quadrifoglio F, et al. (2009) APE1/Ref-1 in Alzheimer's disease: an immunohistochemical study. Neurosci Lett 466: 124-127.
- Coppedè F (2011) An overview of DNA repair in amyotrophic lateral sclerosis. Scientific World Journal 11: 1679-1691.
- Jiang Y, Guo C, Fishel ML, Wang ZY, Vasko MR, et al. (2009) Role of APE1 in differentiated neuroblastoma SH-SY5Y cells in response to oxidative stress: use of APE1 small molecule inhibitors to delineate APE1 functions. DNA Repair (Amst) 8: 1273-1282.
- Mantha AK, Dhiman M, Taglialatela G, Perez-Polo RJ, Mitra S (2012) Proteomic study of amyloid beta (25-35) peptide exposure to neuronal cells: Impact on APE1/Ref-1's protein-protein interaction. J Neurosci Res 90: 1230-1239.
- Li WM, Barnes T, Lee CH (2010) Endoribonucleases--enzymes gaining spotlight in mRNA metabolism. FEBS J 277: 627-641.
- Rodgers ND, Wang Z, Kiledjian M (2002) Characterization and purification of a mammalian endoribonuclease specific for the alpha -globin mRNA. J Biol Chem 277: 2597-2604.
- Tomecki R, Dziembowski A (2010) Novel endoribonucleases as central players in various pathways of eukaryotic RNA metabolism. RNA 16: 1692-1724.
- Kim SE, Gorrell A, Rader SD, Lee CH (2010) Endoribonuclease activity of human apurinic/apyrimidinic endonuclease 1 revealed by a real-time fluorometric assay. Anal Biochem 398: 69-75.
- Berquist BR, McNeill DR, Wilson DM 3rd (2008) Characterization of abasic endonuclease activity of human Ape1 on alternative substrates, as well as effects of ATP and sequence context on AP site incision. J Mol Biol 379: 17-27.
- Vascotto C, Fantini D, Romanello M, Cesaratto L, Deganuto M, et al. (2009) APE1/Ref-1 interacts with NPM1 within nucleoli and plays a role in the rRNA quality control process. Mol Cell Biol 29: 1834-1854.
- Chattopadhyay R, Das S, Maiti AK, Boldogh I, Xie J, et al. (2008) Regulatory role of human AP-endonuclease (APE1/Ref-1) in YB-1-mediated activation of the multidrug resistance gene MDR1. Mol Cell Biol 28: 7066-7080.
- Kuninger DT, Izumi T, Papaconstantinou J, Mitra S (2002) Human AP-endonuclease 1 and hnRNP-L interact with a nCaRE-like repressor element in the AP-endonuclease 1 promoter. Nucleic Acids Res 30: 823-829.
- Hegde ML, Hazra TK, Mitra S (2008) Early steps in the DNA base excision/single-strand interruption repair pathway in mammalian cells. Cell Res 18: 27-47.
- Vidal AE, Boiteux S, Hickson ID, Radicella JP (2001) XRCC1 coordinates the initial and late stages of DNA abasic site repair through protein-protein interactions. EMBO J 20: 6530-6539.
- Kutuzov MM, Ilina ES, Sukhanova MV, Pyshnaya IA, Pyshnyi DV, et al. (2011) Interaction of poly(ADP-ribose) polymerase 1 with apurinic/apyrimidinic sites within clustered DNA damage. Biochemistry (Mosc) 76: 147-156.
- Hill JW, Hazra TK, Izumi T, Mitra S (2001) Stimulation of human 8-oxoguanine-DNA glycosylase by AP-endonuclease: potential coordination of the initial steps in base excision repair. Nucleic Acids Res 29: 430-438.
- Sukhanova MV, Khodyreva SN, Lebedeva NA, Prasad R, Wilson SH, et al. (2005) Human base excision repair enzymes apurinic/apyrimidinic endonuclease1 (APE1), DNA polymerase ß and poly (ADP-ribose) polymerase 1: interplay between strand-displacement DNA synthesis and proofreading exonuclease activity. Nucleic Acids Research 33: 1222-1229.
- Ranalli TA, Tom S, Bambara RA (2002) AP endonuclease 1 coordinates flap endonuclease 1 and DNA ligase I activity in long patch base excision repair. J Biol Chem 277: 41715-41724.
- Ando K, Hirao S, Kabe Y, Ogura Y, Sato I, et al. (2008) A new APE1/Ref-1-dependent pathway leading to reduction of NF-kappaB and AP-1, and activation of their DNA-binding activity. Nucleic Acids Res 36: 4327-4336.
- Ema M, Hirota K, Mimura J, Abe H, Yodoi J, et al. (1999) Molecular mechanisms of transcription activation by HLF and HIF1alpha in response to hypoxia: their stabilization and redox signal-induced interaction with CBP/p300. EMBO J 18: 1905-1914.
- Kenny MK, Mendez F, Sandigursky M, Kureekattil RP, Goldman JD, et al. (2001) Heat shock protein 70 binds to human apurinic/apyrimidinic endonuclease and stimulates endonuclease activity at abasic sites. J Biol Chem 276: 9532-9536.
- Opresko PL, Cheng WH, von Kobbe C, Harrigan JA, Bohr VA (2003) Werner syndrome and the function of the Werner protein; what they can teach us about the molecular aging process. Carcinogenesis 24: 791-802.
- Gray MJ, Zhang J, Ellis LM, Semenza GL, Evans DB, et al. (2005) HIF-1alpha, STAT3, CBP/p300 and Ref-1/APE are components of a transcriptional complex that regulates Src-dependent hypoxia-induced expression of VEGF in pancreatic and prostate carcinomas. Oncogene 24: 3110-3120.
- Sengupta S, Mantha AK, Mitra S, Bhakat KK (2011) Human AP endonuclease (APE1/Ref-1) and its acetylation regulate YB-1-p300 recruitment and RNA polymerase II loading in the drug-induced activation of multidrug resistance gene MDR1. Oncogene 30: 482-493.
Relevant Topics
- Agricultural biotechnology
- Animal biotechnology
- Applied Biotechnology
- Biocatalysis
- Biofabrication
- Biomaterial implants
- Biomaterial-Based Drug Delivery Systems
- Bioprinting of Tissue Constructs
- Biotechnology applications
- Cardiovascular biomaterials
- CRISPR-Cas9 in Biotechnology
- Nano biotechnology
- Smart Biomaterials
- White/industrial biotechnology
Recommended Journals
Article Tools
Article Usage
- Total views: 16017
- [From(publication date):
August-2013 - Dec 19, 2025] - Breakdown by view type
- HTML page views : 11189
- PDF downloads : 4828