Review Article Open Access
Argonaute-1 Machinery Silent Cancer Noises
Tanmoy Mondal1,3, Dhananjaya Pal1, Manika Pal Bhadra1* and Utpal Bhadra2*
1Centre for Chemical Biology, Indian Institute of Chemical Technology, Hyderabad, India
2Functional Genomics and Gene Silencing Group, Centre For Cellular and Molecular Biology, Hyderabad, India
3Academy of Scientific and Innovative Research, Aruna Asaf Ali Marg, New Delhi, India
- *Corresponding Author:
- Manika Pal Bhadra
Centre for Chemical Biology, CSIRIndian Institute of Chemical Technology
Hyderabad, India
Tel: 91 40 27193236
Email: manikapb@gmail.com
- Utpal Bhadra
CSIR - Centre for Cellular and MolecularBiology
Hyderabad, India
Tel: 91 40 27192513
E-mail: utpal@ccmb.res.in
Received Date: February 01, 2016 Accepted Date: February 25, 2016 Published Date: March 03, 2016
Citation: Mondal T, Pal D, Bhadra MP, Bhadra U (2016) Argonaute-1 Machinery Silent Cancer Noises. Can surg 1:102.
Copyright: © 2016 Mondal T, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Cancer Surgery
Abstract
Small non-coding RNA mediated that is micromanaging of global gene regulation was well established as fundamental principle in cell or tissue functions. To carry out, biological role small RNA needs a unique batch of protein family named Argonaute (AGO). The Argonaute proteins carry sophisticated and highly delicate interacting modules common for three specialized small RNAs, microRNA (miRNA), short interfering RNA (siRNA) and Piwi interacting RNA (piRNA). These RNA-proteins complexes are unique to coordinate silencing events in the genome. Recent works have made a novel role of AGO1 protein by extending its tentacle towards cancer monitoring pathways. It makes a constructive bridge between a direct and constructive link between cancer and RNAi (RNA interference) machinery. Apart from multifarious classical functions such as disruption of mRNA translation, decay, transcriptional regulation and splicing, we demonstrate a new concept to narrate the role of AGO1 proteins in different cancer regulatory pathways via microRNA- Argonaute circuit specifically.
Keywords
Argonaute-1; microRNA; piwi associated RNA; RNA interference; Cancer
Introduction
Small regulatory RNA research is a cutting edge ingredient for finetuning of gene regulation. RNA mediated post-transcriptional inhibition was first noticed by antisense RNA expressed in transgenic plants [1,2]. Gene regulation through small regulatory RNAs is operated by both transcriptional and post-transcriptional gene regulatory mechanisms. Small non-coding RNA mediated transcriptional silencing is involved with new family members of proteins coincides the name of Greek warriors from mythology, the Argonautes. The Argonaute (AGO) proteins are very important in all small RNA guided gene-silencing mechanisms identified so far. AGO family proteins are highly found in all organism, except Saccharomyce scerevisiae that has no small RNA machinery [3]. AGO family protein contains a specific interacting module that interacting with three small RNAs (microRNA, siRNA and piRNA) to induce silencing process. The AGO family members are classified into two broad area a. AGO proteins (also known as AGO clade), alike Arabidopsis thaliana AGO1 and b. PIWI (P-element induced wimpy testis) proteins, close to Drosophila PIWI with significant homology (also known as PIWI clade). AGO proteins mainly related to posttranscriptional gene silencing (PTGS) through their interaction with microRNAs (miRNAs) or short interfering RNAs (siRNAs) [4,5]. Germ line cells are the main site of expression for PIWI protein. Transposable genetic element silencing occurs through the binding of PIWI proteins to PIWI-interacting RNAs (piRNAs) in germ line cells [6]. A third clade (referred as WAGO) has evolved in Caenorhabditis elegans where 26 different Argonaute genes exist. This is recognizably different in nature from the AGO and PIWI clades. Role of these proteins are mainly as secondary Argonaute proteins. They obtain their small RNA load through ‘primary’ Argonaute proteins or through small RNA amplification system [7]. Some of these Argonaute proteins cluster to the AGO clade [8], are exclusively expressed in the germ line and also have plant-specific functions [9].
In the last decades, the numerous functional insights or mechanisms of Argonaute protein have been evaluated. A new vistas including additional function of Argonaute protein that demonstrate their strong link to tumor regulatory pathways [10] was also improved our current knowledge. This review describes a novel cellular aspect of Argonaute tentacle, which is emerged towards cancer controlling pathways. A direct relationship between Argonaute and tumor formation was supported from human. Argonaute homologue in human elF2C [11] proteins might have a direct function in cancer stem cell renewal via RNA-dependent silencing mechanism as a positive ingredient of RISC (RNA-induced silencing complex). This source might function as a direct consequence for unique function of different diverse aspects of Argonaute protein with an emphasis for tumor formation.
Argonaute and its Unique Features
The Argonaute family protein plays a critical role in RNA mediated silencing processes, as crucial catalytic components of RNA-induced silencing complex (RISC). RNA interference (RNAi) is a kind of gene silencing phenomenon mediated through small RNAs forming RISC complex. Argonaute proteins have an ability to bind different classes of small non-coding RNAs, including small interfering RNAs (siRNAs), microRNAs (miRNAs) and Piwi-interacting RNAs (piRNAs). Role of small RNAs is to guide AGO proteins to their targets through sequence specific (base pairing) manner. This process leads to the inhibition of translation or the degradation of target mRNA [12].
Though RNA mediated gene silencing was first observed and reported on 1986-1990 time still the understanding of actual mechanism of RNA silencing began only with the experiments of double-stranded RNA triggered gene silencing (RNAi) by Andrew Fire and colleagues in the year of 1998 [13]. In RNA silencing pathway, long RNAs are cleaved into small RNA fragments that direct the repression of transcription of a gene or translation of a mRNA target in sequence specific way. Single-stranded RNAs, known as guide strands which are incorporated into RNA-induced silencing complex (RISC) where Argonaute family proteins take part a critical role to induce efficient silencing.
Structural domains of argonaute
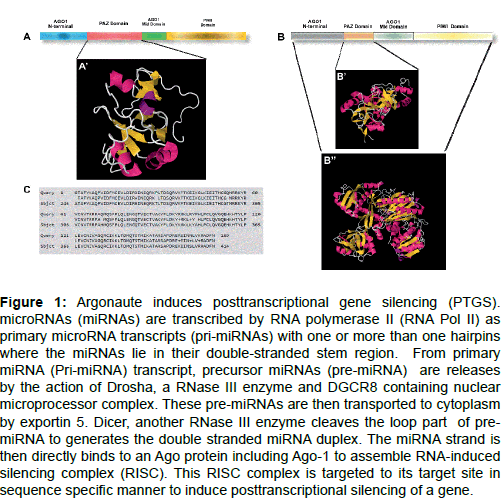
Argonaute proteins contain four domains in their structure- a) The N-terminal domain, b) PAZ domain, c) Mid domain and d) PIWI domain (Figure 1A-C).
Figure 1: Argonaute induces posttranscriptional gene silencing (PTGS). microRNAs (miRNAs) are transcribed by RNA polymerase II (RNA Pol II) as primary microRNA transcripts (pri-miRNAs) with one or more than one hairpins where the miRNAs lie in their double-stranded stem region. From primary miRNA (Pri-miRNA) transcript, precursor miRNAs (pre-miRNA) are releases by the action of Drosha, a RNase III enzyme and DGCR8 containing nuclear microprocessor complex. These pre-miRNAs are then transported to cytoplasm by exportin 5. Dicer, another RNase III enzyme cleaves the loop part of premiRNA to generates the double stranded miRNA duplex. The miRNA strand is then directly binds to an Ago protein including Ago-1 to assemble RNA-induced silencing complex (RISC). This RISC complex is targeted to its target site in sequence specific manner to induce posttranscriptional silencing of a gene.
PAZ domain: Two protein families, Dicer and Argonaute which take part a key role in RNAi mechanisms. Both of them contain the PAZ domain in their protein structures. The PAZ domain has two subdomains. One of the subdomain of PAZ domain displays OB-like folding (oligonucleotide/oligosaccharide binding) which indicated that the PAZ motif can bind the single-stranded nucleic acids [14-16]. Biochemical approaches combined with crystallographic studies showed that the PAZ domain has an ability to bind with ssRNAs (single strand RNA) at a low affinity and independent of sequence specific manner [17,18]. A notable feature of PAZ domain is that, it has an ability to recognize the 3′-ends of an ssRNA (single stranded RNA). Both miRNAs and distinct types of siRNAs are trimmed by the action of RNase III enzymes, acting as a sequential manner (Drosha and Dicer in animals, Dicer alone in plants and yeast). As a result of this, formation of two characteristically 3′-overhangs occur on the processed product. In this way, the PAZ domain could initially recognize these small regulatory RNAs from other RNAs (degraded RNAs) that are derived from other non-related pathways.
The PIWI domain: An absence of the N-terminal and the PAZ domain in the Piwi-like protein of the structure of full length archaeal and eubacterial Argonautes and also the archaeal archaeoglobus fulgidus, revealed that the PIWI domain can show an RNase-H like activity [19-22]. RNase-H mediated catalysis needs a conserved Asp- Asp-Asp/Glu motif in the catalytic site and binding of two divalent cations by the ribonuclease and the cleavage of RNA depends on a DNA template. The cleavage site of Argonaute proteins also has an Asp-Asp- Asp/Glu/His/Lys motif which requires the binding of a divalent metal ion for their activity [8]. Cleavage products of this catalysis contain 3′-OH and 5′-phosphate, a characteristic of RNase-H like activity [23,24]. The 5’ phosphate group of a siRNA or a miRNA is crucial for their activity [25]. A divalent metal ion is the responsible factor for the anchoring of this 5′ phosphate at the interface between mid and the PIWI domains [20,21]. Metazoan AGO protein mid domain also has a specific motif, the MC domain which has a homology to the translation initiation factors [eIF4E (eukaryotic translation initiation factor 4E)] cap binding motif. For this reason, MC domain is required for efficient translational regulation through the cap binding [26].
Endonuclease activity of the Piwi-like and Argonaute-like proteins is crucial for the proper function of RISC [27] in fission yeast, fungi, plants, flies and mammals. Argonaute-like proteins take an important role in the process of siRNA maturation by removing non-active siRNA strand [28-30] and initiating sequence specific chopping of target RNAs [31]. The maturation of repeat-associated small interfering RNAs (rasiRNA) and Piwi interacting RNAs (piRNAs) may be also dependent on the cleavage activity of Piwi-like proteins in flies and mammals [32]. The presence of a full PIWI-domain in Argonaute proteins serve as a catalytic centre which only partially explains the chopping activity of Argonaute proteins.
Diverse argonaute activities
Human AGO1 and AGO4, one of the Piwi-like protein in human HIWI2 (alternatively PIWIL4), and most of the C. elegans group 3 Argonaute proteins have diverged their catalytic motif, probably impairs their endonuclease activity [8]. Argonautes are important for small RNA maturation and small RNA induced gene silencing which may require interactions between protein complexes. It is reported that human AGO1 and AGO2 interact with a number of proteins in three complexes of distinct size. These proteins are mainly RNA binding proteins, which are involved in RNA processing, maturation, transport of RNA and the regulation of translation and the stability of RNA. Some of these interactions are RNA mediated, some may bind directly to Argonautes or associations are through the help of other interacting partners [33]. In the selection of active siRNA and miRNA strand both Argonaute and DICER proteins take part an active role. PIWI box of human Argonaute is responsible for the binding of Dicer’s RNase III domain [34]. The PIWI domain of Drosophila Argonaute1 protein (AGO1) directly interacts with a characteristic protein, GW182, which is responsible for the formation of cytoplasmic processing bodies (P bodies). GW182 is also important for miRNA mediated gene regulation. In fly, this might function downstream of AGO1 [35]. Around 22 amino acid long part of PIWI domain which is known as AGO hook, is the responsible factor for the accommodation of guide stand’s 5’ phosphate group of a siRNA [36]. The GW/WG repeats and the Trp residues in the domain are crucial for the interaction. A number of unrelated Argonaute interactor proteins; large subunit of plant polymerase IV [NRPD1b (nuclear RNA polymerase D1b)], metazoan GW182 protein family orthologues and yeast Tas3 also, all of them contains this type of amino acid repeat [36,37]. Numerous copies of GW/WG motifs of NRPD1b and GW182 proteins allow them to bind with multiple Argonaute proteins to assemble the regulatory complex. Probably this motif takes part in the process of small RNA mediated gene regulation also [36].
Argonate protein in transcription machinery and epigenetic regulation
Early models hypothesize that direct base pairing of siRNA with DNA might necessary to recruit DNA methyltransferase (DNMT), but recent study has shown DNA-RNA hybrid structures cannot methylated by DNMTs [38]. But, AGO-siRNA complex can binds to the newly transcribed RNA in the nucleus [39,40]. A number of reports suggest that TGS (transcriptional gene silencing) needs active transcription and sense-strand mRNA expression through the promoter of the gene [41,42]. Nevertheless, other data shows that AGO1 and AGO2 bind to the antisense transcripts during the process of TGS [43,44]. Transfection of only anti-sense siRNA is sufficient to induce the silencing of the EEF1A1 (eukaryotic translation elongation factor 1 alpha 1) gene [39,41]. Microarray data of MCF7 cell reveals that miRNA with AGO2 association are mainly derived from the sense strands of the corresponding pre-miRNA, whereas the association of AGO1 are predominantly from the anti-sense strand [45].
A number of documentation showed that, less RNAPII (RNA polymerase II) is available at promoter region after TGS [40,42,44,46-51] where as there is an enrichment of RNAPII in case of RNAa [50,52-54]. Studies have reported modified histones enriched promoters having well known silencing marks after TGS and active marks just after RNAa. TGS specifically is associated with high profusion of repressive H3K9me2, H3K9me3 (histone3, lysine9 di- and tri-methylation) and H3K27me3 (histone3, lysine27 tri-methylation) marks, with reduction of active H3K9Ac (histone 3 lysine 9 acetylation) and H3K14Ac (histone3, lysine 14 acetylation) marks and sometimes loss of H3K4me3 mark. For RNAa, regulation of histone is just opposite, where loss of H3K9me2, H3K27me3 and gain of H3K4me2, H3K4me3 marks are well documented. In other side, RNAa is also associated with the depletion of H3K9ac and H3K14ac marks alike TGS. Association of these marks with active and bivalent promoters was found in mouse cells [55]. An increased level in DNA methylation at different targeted promoter region has been reported in some TGS study. The de novo methyltransferase Dnmt3a is essential for the establishment of DNA methylation at the promoter region. Dnmt1 is necessary for the maintenance of that methylation [43,56]. Requirement of TGS to induce chromatin condensation is mainly associated with histonedeacetylase HDAC, most probably the H3K9 methyltransferase G9a, but not with EZH2 (Enhancer of zeste homolog 2), a polycombgroup H3K27 methyltransferase [43,56]. Enrichment of EZH2 at target promoters was confirmed by ChIP (Chromatin immunoprecipitation) experiments [57,58]. The requirement of TARBP2 (Trans-Activation- Responsive RNA-Binding Protein 2) in TGS is also important for the assembly of RISC-loading complex subunit [59]. For RNAa, it has been described that the PTGS2 activationis associated with WDR5 (WD repeat-containing protein 5) and the GW182, an Argonaute interacting protein. Interestingly, the lncRNA (long intergenic non-coading RNA), HOTTIP (HOXA transcript at the distal tip), transcribed from the HOXA 5′-end, binds to the WDR5 and in turn, MLL (mixed-lineage leukaemia) histone methyltransferase which induces gene activation along with H3K4 tri-methylation (H3K4me3). Same histone mark has demonstrated to be enriched in RNAa [59-61]. This suggests short RNA and lncRNA (long non coding RNA) mediated transcriptional gene activation may share a number of common features in their process.
Alternative splicing: Redirection of exon splicing in aberrant splice sites of the disease-associated SMN2 (survival of motor neuron 2) and dystrophin genes has been reported as a result of exogenous duplex RNAs introduction into the nuclear region of the cell [62]. Duplex RNA mediates the recruitment of AGO2 to the pre-mRNA transcripts and results the alteration in splicing without nuclear pre-mRNA cleavage. Regulation of pre-mRNAs by MALAT (Metastasis Associated Lung Adenocarcinoma Transcript 1) was the first demonstration of the regulation of alternative splicing by endogenous sRNA pathways [63]. Subsequently, physical association of MALAT with AGO1 and AGO2 has been identified as a splicing factors and chromatin modifiers [64]. Dicer dependent AGO1 and AGO2 recruitment facilitates spliceosome function and modulates RNAPII elongation rate and affects alternative splicing thereby. This AGO1 and AGO2 recruitment to the transcribed regions of CD44 (CD44 molecule) gene required the Dicer and histone modifying enzymes. This histone modifying enzymes results an increased H3K9 methylation on variant heterochromatin and TGS associated exons [64].
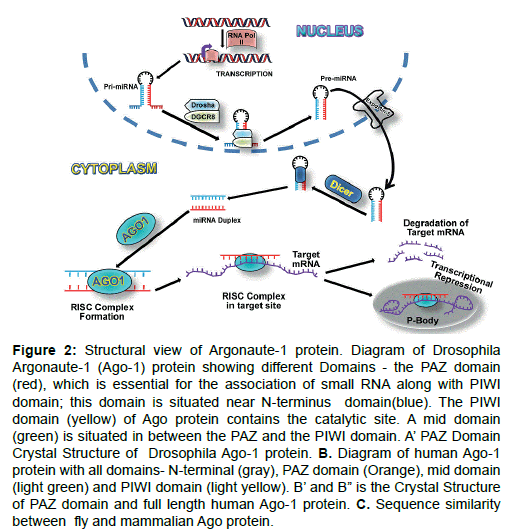
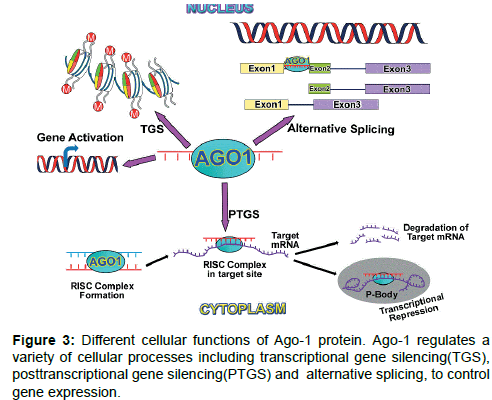
Functional location: AGO family proteins generally induces posttranscriptional gene silencing (PTGS) in cytoplasmic part of a cell (Figures 2 and 3). Nuclear function of AGO proteins also reported as a regulator of transcriptional gene silencing (TGS) in fission yeast and plants. Nuclear role of mammalian AGO proteins include gene activation [52,60,65], TGS [57,66-68] and alternative splicing [64] (Figure 3). AGO1 and AGO2 both play an important role in neuronal differentiation by miRNA-mediated process [69] through binding with the same pool of miRNAs and mRNA targets [70].
Figure 2: Structural view of Argonaute-1 protein. Diagram of Drosophila Argonaute-1 (Ago-1) protein showing different Domains - the PAZ domain (red), which is essential for the association of small RNA along with PIWI domain; this domain is situated near N-terminus domain(blue). The PIWI domain (yellow) of Ago protein contains the catalytic site. A mid domain (green) is situated in between the PAZ and the PIWI domain. A’ PAZ Domain Crystal Structure of Drosophila Ago-1 protein. B. Diagram of human Ago-1 protein with all domains- N-terminal (gray), PAZ domain (Orange), mid domain (light green) and PIWI domain (light yellow). B’ and B” is the Crystal Structure of PAZ domain and full length human Ago-1 protein. C. Sequence similarity between fly and mammalian Ago protein.
AGO1 overexpression causes slow down the cell cycle process may be due to G1/S transition delay. And it induces apoptosis after getting UV exposure. Thus, AGO1 overexpression acts as a tumor suppressor element for the cell [69,71]. Significantly lower AGO1 levels were found in several tumor cell lines [73]. Deletion of AGO1 locus which is present at 1p34–35 of chromosome 1 is often associated with neuroectodermal tumours, found in Wilms’ tumors [11,73]. AGO1’s role in the cell cycle modulation is also confirmed by its microarray expression profiles of differentiating neuronal cells during brain development. Very low level of expression was reported at undifferentiated neural progenitor cells and significantly increment expression profile in the time of differentiation into neurons [74].
AGO1 and the regulation of cell cycle
In fly system, the regulatory role of AGO-1 in cell cycle control was established. In this process, AGO-1 is associated with cyclinB, which acts as a G2/M cyclin in cell cycle progression. In mitotic division, AGO-1 is very crucial for proper segregation of chromosome and spindle fiber assembly in the time of early embryonic development. Increased activity of cyclinB-Cdk1 (cyclinB- cyclin-dependent kinase 1) and decreased activity of cell cycle check point proteins such as, p53, grp (grapes), mei-41(meiotic 41) and Wee1(Wee1 kinase) was reported as a result of AGO-1 mutation in the division process of fly embryogenesis. Involvement of 2 embryonic miRNAs; miR-317 and miR-981 for the spatiotemporal regulation of cyclin B was also reported in the same study [75].
AGO1 machinery genes and cancer-related pathways
Clustering AGO1-bound genes (AbGs) by their chromosomal location (-5 kb, -1 kb or -0.5 kb genes) uncovers several cyto-bands implicated in various overrepresented human cancers. Further, Gene pathway enrichment analysis also revealed the same [10]. Top enriched cyto-bands 19p13.3 and 16p13.3 has been reported by a number of studies for several cancer types including lymphoma, breast, prostate and thyroid cancer [76-79].
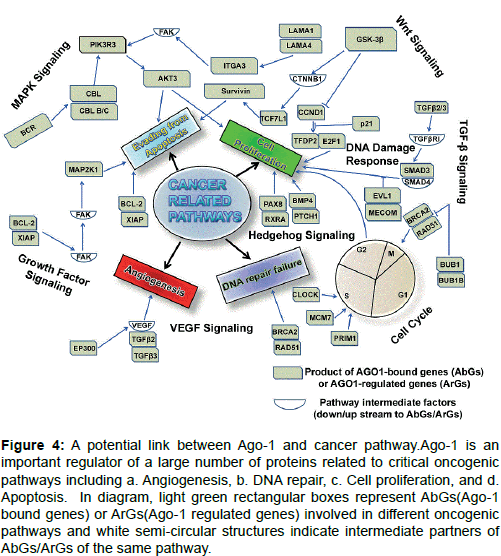
High enrichment of AbGs-5 kb genes includes MAPK (Mitogenactivated protein kinases) signalling, Wnt signing, endocytosis and focal adhesion (Figure 4). Many proliferation promoting and protooncogenes are illustrated in these pathways including serine/threonine/ tyrosine kinases, G-protein coupled receptors (GPCs), membrane associated G-proteins, growth factors; DNA-binding protein factors and transcription factors as well. A large number of genes grouped under AbG-0.5 kb gene group such as CDC20 (Cell Division Cycle 20), SMC1A (Structural Maintenance of Chromosomes 1A), BUB1 (budding uninhibited by benzimidazoles 1) and SMAD3 (Mothers against Decapentaplegic Homolog 3) are known to promote cell cycle progression and proliferation in different cancer types [80-83]. Gene Ontology (GO) classification and KEGG (Kyoto Encyclopedia of Genes and Genomes) pathway enrichment analysis also reported that AGO1- regulated genes (ArGs), regulate various cancer related pathways and they shared this with AbGs-5 kb genes including cell cycle regulation, MAPK signalling pathway, p53 signaling pathway in various cancers (Figure 4) including with prostate and colorectal cancer [10].
Figure 4: A potential link between Ago-1 and cancer pathway.Ago-1 is an important regulator of a large number of proteins related to critical oncogenic pathways including a. Angiogenesis, b. DNA repair, c. Cell proliferation, and d. Apoptosis. In diagram, light green rectangular boxes represent AbGs(Ago-1 bound genes) or ArGs(Ago-1 regulated genes) involved in different oncogenic pathways and white semi-circular structures indicate intermediate partners of AbGs/ArGs of the same pathway.
Function of AGO-1 during tumor formation
Argonaute proteins are well known for their cytoplasmic microRNA (miRNA) mediated processes in which they regulate post-transcriptional gene silencing (PTGS) to regulate the expressions of transcripts [84,85]. Apart from this, they also take part a role in the process of heterochromatin formation and the establishment of repressive epigenetic marks, known as transcriptional gene silencing (TGS), in the nuclear part of fission yeast and plants. In fission yeast, AGO proteins interact with the antisense strand transcripts to assemble the RNAinduced transcriptional silencing (RITS) complex at the centromeric regions of the chromosome to induce heterochromatinization [86]. Similarly, to induce histone and DNA methylation Argonaute proteins interact with ribonucleoprotein complexes in plant [87]. In mammals, AGO members are associated with a spectrum of nuclear processes including gene activation [10,52,60,65], TGS [57,66-68] and alternative splicing [64]. Further, the direct interaction of nuclear AGO1 with RNA Polymerase II (RNAPII) and subsequent wide association with chromosomal loci of the transcriptionally active gene’s promoter region has been elicited by ChIP-seq analysis. Further analysis show the regulation of AGO1 to the expression of nuclear AGO1-bound genes which are implicated in carcinogenic pathways including cellular growth, cell cycle progression, and survival [10].
MicroRNAs (miRNAs) are 22 nucleotide long single-stranded noncoding RNAs which repress gene expression through the interaction with mRNA (messenger RNA) by suppressing the translation of that particular mRNA or by inducing the cleavage of that, depending on the degree of homology to the target sequence [88-90]. More than 500 miRNAs have been identified in human and almost 30% protein coding genes are estimated to be miRNA regulated in Homo sapiens [89-92]. A number of vital biological processes, such as cell proliferation, apoptosis, cellular differentiation and organ development are under the regulation of miRNA activity [93,94]. Furthermore, several studies indicate the role of miRNAs in the development and progression of cancers, including lung cancer, by changing the normal expression of tumor suppressor genes or proto-oncogenes or both the tumor suppressor and proto-onco genes [95,96]. Mature miRNAs are processed by two step processing mechanism. First miRNA genes are transcribed by RNA polymerase II and form relatively larger primary miRNAs which are then capped and polyadenylated [97]. After transcription of primary miRNAs, they are clipped in the nucleus by the nuclear microprocessor machinery which is consist of the RNAse III DROSHA and the DGCR8/PASHA (partner of drosha), one kind of double strand RNA binding protein [98]. 70 nucleotide precursor miRNAs are produced after this process [99]. These precursor miRNAs are then exported to the cytoplasm by the help of RanGTP-dependent double stranded RNA (dsRNA) transporter exportin 5 (XPO5) [100]. In next step, precursor miRNAs are processed by DICER, one more RNAse III enzyme, to produce 22 nucleotide double stranded miRNA molecules [101] (Figure 2). One strand of the miRNA duplex is take part a role in the formation of ribonucleoprotein effector complex, known as RNA induced silencing complex (RISC) which consists of Argonaute proteins (AGO1, AGO2 and HIWI), human immunodeficiency virus trans-activating response RNA binding protein (TRBP), GEMIN3 (Gem (Nuclear Organelle) Associated Protein 3) and GEMIN4 (Gem (Nuclear Organelle) Associated Protein 4) [102,103]. Furthermore, the deregulation of miRNAs along with the altered regulation of miRNA processing genes has also been implicated in tumorigenesis process [104-107]. It has been described that an abnormal expression of DICER adapts the development of lung cancers [104]. Down regulation of the miRNA processing enzymes DROSHA and DICER result in reduced global miRNA expression and subsequent enhanced cellular transformation and tumorigenesis in mammalian cell lines [105]. As Argonaute proteins have an important role in miRNA function, so they also been associated with a number of cancer types [106]. TRBP also has been shown to have an oncogenic potential, and is able to induce tumors in nude mice [108]. Considering these emerging lines of evidence it is suggest that AGO1 may play an indispensable role to development and the progression of cancer like other miRNA processing proteins. As AGO1 is an crucial player of RNAi mechanism so obviously questions comes –“The effects of AGO1 on cellular differentiation, cell cycle regulation and apoptosis is the result of miRNA misregulation only, or any miRNA independent process also involved there?”
In the miRNA dependent storyline, AGO1 was identified as a part of large multiprotein complex execute to miRNA maturation from pre-miRNA. May be these effects also depend on accessory factors associated with AGO1. TRIM32 (Tripartite Motif Containing 32) is an example of this phenomenon; TRIM32 protein induces neuronal differentiation by direct binding with AGO1 and increasing the specific miRNA activity [109]. In the other scenario, the tumor suppressor like activities was reported only upon AGO1 overexpression, not in AGO2 [69]. In S. Pombe, it has shown that AGO1 is important for a normal G1/S transition of cell cycle; cytokinesis and the DNA damage response although miRNAs are not expressed in S. Pombe [110]. In this case, AGO1 protein directly associates with factors involved in regulating the cell cycle [111]. AGO1 mutants isolated in Arabidopsis also support miRNA-independent functions of AGO1. These mutants either defective in RNAi still functional in development or vice versa which indicates the control of these two processes by AGO1 can be uncoupled [112]. So, it is noted that the effects of AGO1 on the cell cycle at G1/S transition are in compliance with both miRNA dependent and independent framework. Actually, most miRNAs, targeting cell cycle modulate this specific checkpoint [113]. By the same token, in S. Pombe where miRNAs are not present, AGO1 mutants are defective in G1 arrest [110]. The level of p53 is strongly upregulated in AGO1 overexpressing cells; this high p53 expression level may give molecular evidence underlying the phenotypic alterations of AGO1 over expressing cells. Recent studies suggest that there is a relationship between p53 and miRNA pathway, whereas p53 plays as a miRNA target, a miRNA transcriptional regulator and also has an involvement in miRNA biogenesis [114,115], a pathway where p53 is a component of AGO1 activated tumor suppressor cascade is like-minded with both miRNA-dependent and independent framework. Further work needed to explore the detailed molecular cascade of AGO1 overexpression effects in context of tumor genesis and tumor suppression.
Now it is known that AGO proteins play a role to regulate gene expression at multiple levels. It is also reported that AGO1 directly interacts with RNA polymerase II (RNAP II) in nucleus. Additionally, AGO1 prevalently bound with multiple genomic region including euchromatic sites and repetitive elements of transposons, defined by the histone mark H3K4me3. Further, binding of AGO1 with the promoter of a gene regulates the expression of that particular gene. Loss of function of AGO1 results the reduction of that particular gene expression due to decreased level of AGO1 and corresponding RNAP II occupancy at transcription start sites and gain of function results opposite effect of this. The mechanism of AGO1 target finding to a particular chromosomal loci remain unclear. It is may be involvement of miRNA which mediates the interaction between nuclear AGO1, chromosomal loci and/or RNAP II. AGO1 bound sequences carry specific miRNA target sites and any disturbance in Dicer function results suppression of AGO1 binding activity to RNAP II [10]. Deletion of the PAZ domain which functions as RNA-binding domain in AGO1, interfered with gene activation; further suggesting a role of RNAs (miRNA) in this process. Exogenous miRNA transfection can promote AGO1 enrichment at highly-complementary sites of gene promoters to control transcription of those genes [51,52,57]. In context of AGO1- RNAP II association, formation of a duplex with complementary target sequence may be plays a role to load AGO1 with miRNA which protects bound RNA from RNase (RNase A/T1) digestion, similar to canonical target recognition manner [116,117]. Other classes of small RNAs may also mediate AGO1 interactions with genomic loci. Association of AGO1 with small RNA from non-miRNA sources also have shown by deep sequencing studies [118,119]. AGO1 fine-tunes gene expression in a miRNA dependent manner at both transcriptional and posttranscriptional levels [120].
Ago-1 counts for disease therapy
The specificity and longterm efficacy of potential exogenous sRNA (Small RNA) in vitro brings the hope for a new class of drugs, based on RNA. Development of new RNA based cancer therapies or suppression strategy of HIV-1 (human immunodeficiency virus-1) replication in the T cells of HIV patients may be possible using this new class of drugs. In vivo efficacy of sRNA mediated transcriptional silencing or activation has been tested in mouse xenograft model system. Stable in vivo CCNB1 (Cyclin B1) RNAa (RNA activation) by miRNA constructs resulted in reduction of tumor size compared to control [52]. In the same way, established tumors treated with lipid transfections of siRNA which targets CDH1 (Cadherin 1) [121] or CDKN1A (Cyclin-Dependent Kinase Inhibitor 1A) [122] in every three days interval showed decreased growth rate and reduced tumor size, relative to control. Further, chemically modified dsRNA, specially designed for lipidoid encapsulated nanoparticle delivery system is efficiently promotes RNAa of CDKN1A in mouse xenograft model [123,124]. Successful lentiviral shRNA (short hairpin RNA) delivery has also been revealed in vivo. Efficient lentiviral delivery of shRNA which targets VEGF (Vascular endothelial growth factor) promoter has also been observed to increase the flow of blood in the hind limb area of ischemic mice [59]. These results demonstrate the promise of translational regulation and the exogenous sRNA therapy as an efficient medicinal system to the clinic. So, it is anticipate the RNA-based therapeutic approach may eventually be used to manipulate the epigenetic state of a particular locus with much targeted manner.
Diseases like pancreatic cancer which are involved with elevated expression of particular genes, at that case RNAi may function as an effective tool to combat the situation. The sequence specificity activity of RNA interference may serve an appropriate treatment particularly for mutated endogenous gene sequence associated cancers. A large number of non-coding RNAs (microRNAs, long non-coding RNAs) are related with different types of human cancers. It is also reported that miR-15a and miR-16a are generally deleted and/or down-regulated in some cancer patients. Although the miRNA functions are not completely understood, the role of miRNAs in cell proliferation coordination and the regulation of cell death during development and other conditions have been uncovered. It is believed that the miRNAs have ability to direct negative or positive regulation at different levels. This is a specific miRNA and target base pair interaction and the recognizing cofactor dependent process [125].
It is well known that instead of DNA, many viruses use RNA as their genetic material and undergo at least one stage of their life cycle when they make their double-stranded genetic element. RNA interference is an evolutionarily ancient mechanism to defend against viruses and other unwanted genetic materials to protect organisms from these kind of threats. The small interfering RNAs cause sequence specific posttranscriptional gene silencing (PTGS) by guiding an endonuclease to the RNA-induced silencing complex (RISC) which cleaves the target mRNA. In Neurospora fungus this process is named quelling and in plants this phenomenon is known as post-transcriptionl gene silencing (PTGS); in mammalian cells this is familiar as RNA interference (RNAi). In response to a complete or very close sequence complementarity between the target and the small RNA, the AGO-protein component of RISC complex mediates the cleavage of target transcript to induce the repression of translation predominantly.
However, more work is necessary to understand the detailed molecular mechanism of AGO1 effects in different cellular pathways and to test potential therapeutic applications to treat a disease state.
Conclusion
Short term (48 hours) AGO1 siRNA transfection may also be able to induce a stable change in gene expression. AGO1 proteins elicit pleiotropic effects on gene expression through the utilization of miRNAs to silence multiple transcripts and regulate various cellular processes in the cytoplasmic part of a cell [84]. In the same way, nuclear AGO1 also shows a pleiotrophy by regulating the transcription process of multiple genes. In PC-3 prostate cancer cells, AGO1 selectively drives those gene expressions, which are involved in oncogenic pathways. Report suggests that it may help to induce cancer phenotype [10], whereas other reports depict just opposite view of this portrait [11,69,71-73]. In this scenario we can take a stand that the effect of AGO1 on cancer may be depends on different context and on cell types also; based on its activities both at nucleus and cytoplasm as well as gene profiles, regulated by it. Further study is necessary to explore the network of AGO1 mediated gene regulatory hub in the context of oncogenic signaling pathways.
Diverse research over last decades has documented the main principle of Argonaute protein function is diverse human diseases via micromanaging small RNA regulatory pathways. However there is potential gaps in our understating is that how Argonaute function in diverse cancer related pathways, Many diversified role of Argonaute protein including uniqueness of structural motif, facilitation of AGO loading to their target small RNA and microRNA, even relieving of silencing causes detachment of AGO proteins from their immediate target was illustrated. However experiments of interlinking AGO-1 proteins and microRNA target has just been started.
A new emerging area that has not yet been described in depth that is a cross talk between Ago signaling pathways and microRNA mediated gene silencing mechanism. The Argonaute proteins are involved in small RNA target function that is participated in phosphorylation cascade of different cancer regulatory pathways. The protein is involved in large regulatory tumor networks. The dynamicity of these networks during apoptosis or signal transduction is feebly illustrated. However their multifarious changes might lead a penetrating but mild chronic effect on cancer pathways. Therefore a microRNA based understanding in subtle changes in gene expression of Argonaute protein might facilitate to unravel the regulatory device of different cancer. We imagined that further studies will be required to elucidate the exact biological function of Argonaute proteins in cancer.
References
- Ecker JR, Davis RW (1986) Inhibition of gene expression in plant cells by expression of antisense RNA. ProcNatlAcadSci USA 83:5372-5376.
- Napoli C, Lemieux C, Jorgensen R(1990) Introduction of a Chimeric Chalcone Synthase Gene into Petunia Results in Reversible Co-Suppression of Homologous Genes in trans. Plant Cell 2: 279-289.
- Drinnenberg IA (2009)RNAi in budding yeast. Science 326: 544-550.
- Peters L, Meister G(2007) Argonaute proteins: mediators of RNA silencing. Mol Cell 26: 611-623.
- Hutvagner G, Simard MJ(2008) Argonaute proteins: key players in RNA silencing. Nat Rev Mol Cell Biol 9: 22-32.
- Siomi MC, Sato K, Pezic D,AravinAA(2011) PIWI-interacting small RNAs: the vanguard of genome defence. Nat Rev Mol Cell Biol 12: 246-258.
- Yigit E(2006) Analysis of the C. elegansArgonaute family reveals that distinct Argonautes act sequentially during RNAi. Cell 127: 747-757.
- Toli NH, Joshua-TorL(2007) Slicer and the argonautes. Nat ChemBiol 3: 36-43.
- Van Ex F, Jacob Y, Martienssen RA (2011)multiple roles for small RNAs during plant reproduction. CurrOpin Plant Biol 14: 588-593.
- Huang V (2013) Ago1 Interacts with RNA polymerase II and binds to the promoters of actively transcribed genes in human cancer cells. PLoS Genet 9: 1003821.
- Koesters R (1999) Human eukaryotic initiation factor EIF2C1 gene: cDNA sequence, genomic organization, localization to chromosomal bands 1p34-p35, and expression. Genomics 61: 210-218.
- Czech B, Hannon GJ(2011) Small RNA sorting: matchmaking for Argonautes. Nat Rev Genet 12:19-31.
- FireA(1998) Potent and specific genetic interference by double-stranded RNA in Caenorhabditiselegans. Nature 391: 806-811.
- Song JJ (2003) The crystal structure of the Argonaute2 PAZ domain reveals an RNA binding motif in RNAi effector complexes. Nat StructBiol 10: 1026-1032.
- Yan KS (2003) Structure and conserved RNA binding of the PAZ domain. Nature 426: 468-474.
- Lingel A, Simon B, Izaurralde E, Sattler M (2003) Structure and nucleic-acid binding of the Drosophila Argonaute 2 PAZ domain. Nature 426: 465-469.
- Ma JB, Ye K, Patel DJ(2004) Structural basis for overhang-specific small interfering RNA recognition by the PAZ domain. Nature 429: 318-322.
- Lingel A, Simon B, Izaurralde E, Sattler M(2004) Nucleic acid 3'-end recognition by the Argonaute2 PAZ domain. Nat StructMolBiol 11: 576-577.
- Yuan YR(2005) Crystal structure of A. aeolicusargonaute, a site-specific DNA-guided endoribonuclease, provides insights into RISC-mediated mRNA cleavage. Mol Cell 19: 405-419.
- Ma JB(2005) Structural basis for 5'-end-specific recognition of guide RNA by the A. fulgidusPiwi protein. Nature 434: 666-670.
- Parker JS, Roe SM,Barford D(2004) Crystal structure of a PIWI protein suggests mechanisms for siRNA recognition and slicer activity. EMBO J 23: 4727-4737.
- Song JJ, Smith SK, Hannon GJ, JoshuaTor L (2004) Crystal structure of Argonaute and its implications for RISC slicer activity. Science 305: 1434-1437.
- Martinez J,Tuschl T (2004) RISC is a 5' phosphomonoester-producing RNA endonuclease. Genes Dev 18: 975-980.
- Schwarz DS, Tomari Y,ZamorePD (2004) The RNA-induced silencing complex is a Mg2+-dependent endonuclease. CurrBiol 14: 787-791.
- Nykanen A, Haley B, Zamore PD (2001) ATP requirements and small interfering RNA structure in the RNA interference pathway. Cell 107: 309-321.
- Kiriakidou M(2007) An mRNA m7G cap binding-like motif within human Ago2 represses translation. Cell 129: 1141-1151.
- Irvine DV (2006)Argonaute slicing is required for heterochromatic silencing and spreading. Science 313: 1134-1137.
- Miyoshi K, Tsukumo H, Nagami T, Siomi H,Siomi MC(2005) Slicer function of Drosophila Argonautes and its involvement in RISC formation. Genes Dev 19: 2837-2848.
- Matranga C, Tomari Y, Shin C, Bartel DP, Zamore, PD (2005) Passenger-strand cleavage facilitates assembly of siRNA into Ago2-containing RNAi enzyme complexes. Cell 123: 607-620.
- Leuschner PJ, Ameres SL, Kueng S, Martinez J (2006) Cleavage of the siRNA passenger strand during RISC assembly in human cells. EMBO Rep 7: 314-320.
- Hutvagne G, Zamore PD (2002) A microRNA in a multiple-turnover RNAi enzyme complex. Science 297: 2056-2060.
- Gunawardane LS(2007) A slicer-mediated mechanism for repeat-associated siRNA 5' end formation in Drosophila. Science 315: 1587-1590.
- Hock J (2007) Proteomic and functional analysis of Argonaute-containing mRNA-protein complexes in human cells. EMBO Rep 8: 1052-1060.
- TahbazN (2004) Characterization of the interactions between mammalian PAZ PIWI domain proteins and Dicer. EMBO Rep 5: 189-194.
- BehmAnsmant I (2006) mRNA degradation by miRNAs and GW182 requires both CCR4:NOTdeadenylase and DCP1:DCP2 decapping complexes. Genes Dev 20: 1885-1898.
- Till S (2007) A conserved motif in Argonaute-interacting proteins mediates functional interactions through the Argonaute PIWI domain. Nat StructMolBiol 14: 897-903.
- El Shami M (2007) Reiterated WG/GW motifs form functionally and evolutionarily conserved ARGONAUTE-binding platforms in RNAi-related components. Genes Dev 21: 2539-2544.
- Ross JP, Rand KN, Molloy PL (2010)Hypomethylation of repeated DNA sequences in cancer. Epigenomics 2: 245-269.
- Han J, Kim D, Morris KV(2007) Promoter-associated RNA is required for RNA-directed transcriptional gene silencing in human cells. ProcNatlAcadSci USA 104: 12422-12427.
- Schwartz JC (2008) Antisense transcripts are targets for activating small RNAs. Nat StructMolBiol 15: 842-848.
- Weinberg MS (2006) The antisense strand of small interfering RNAs directs histone methylation and transcriptional gene silencing in human cells. RNA 12: 256-262.
- Napoli S, Pastori C, Magistri M, Carbone GM,Catapano CV(2009) Promoter-specific transcriptional interference and c-myc gene silencing by siRNAs in human cells. EMBO J 28: 1708-1719.
- Hawkins PG, Santoso S, Adams C, Anest V, Morris KV(2009) Promoter targeted small RNAs induce long-term transcriptional gene silencing in human cells. Nucleic Acids Res 37: 2984-2995.
- Chu Y, Yue X, Younger ST, Janowski BA, Corey DR (2010) Involvement of argonaute proteins in gene silencing and activation by RNAs complementary to a non-coding transcript at the progesterone receptor promoter. Nucleic Acids Res 38: 7736-7748.
- Turchinovich A, BurwinkelB(2012) Distinct AGO1 and AGO2 associated miRNA profiles in human cells and blood plasma. RNA Biol 9: 1066-1075.
- Janowski, BA (2006) Involvement of AGO1 and AGO2 in mammalian transcriptional silencing. Nat StructMolBiol 13: 787-792.
- Zhang MX (2005) Regulation of endothelial nitric oxide synthase by small RNA. ProcNatlAcadSci USA 102:16967-16972.
- Jiang G (2012) Small RNAs targeting transcription start site induce heparanase silencing through interference with transcription initiation in human cancer cells. PLoS One 7: 31379.
- Roberts TC, Andaloussi SE, Morris KV, McClorey G, Wood MJ(2012) Small RNA-Mediated Epigenetic Myostatin Silencing. MolTher Nucleic Acids 1: 23.
- Yue X (2010) Transcriptional regulation by small RNAs at sequences downstream from 3' gene termini. Nat ChemBiol 6: 621-629.
- Younger ST, Corey DR(2011) Transcriptional gene silencing in mammalian cells by miRNA mimics that target gene promoters. Nucleic Acids Res 39: 5682-5691.
- Huang V(2012) Upregulation of Cyclin B1 by miRNA and its implications in cancer. Nucleic Acids Res 40: 1695-1707.
- Place RF, Li LC, Pookot D, Noonan EJ, Dahiya R (2008) MicroRNA-373 induces expression of genes with complementary promoter sequences. ProcNatlAcadSci USA 105: 1608-1613.
- Matsui M (2010) Activation of LDL receptor expression by small RNAs complementary to a noncoding transcript that overlaps the LDLR promoter. ChemBiol 17: 1344-1355.
- Karmodiya K, Krebs AR, OuladAbdelghani M, Kimura H, Tora L (2012) H3K9 and H3K14 acetylation co-occur at many gene regulatory elements, while H3K14ac marks a subset of inactive inducible promoters in mouse embryonic stem cells. BMC Genomics 13: 424.
- Turner AM, De La Cruz J, Morris KV(2009) Mobilization-competent Lentiviral Vector-mediated Sustained Transcriptional Modulation of HIV-1 Expression. MolTher 17: 360-368.
- Kim DH, Villeneuve LM, Morris KV, Rossi JJ (2006) Argonaute-1 directs siRNA-mediated transcriptional gene silencing in human cells. Nat StructMolBiol 13: 793-797.
- Kim DH, Saetrom P, Snove O, Jr, Rossi JJ(2008) MicroRNA-directed transcriptional gene silencing in mammalian cells. ProcNatlAcadSci USA 105: 16230-16235.
- Turunen MP(2009)Efficient regulation of VEGF expression by promoter-targeted lentiviralshRNAs based on epigenetic mechanism: a novel example of epigenetherapy. Circ Res 105: 604-609.
- Janowski BA (2007) Activating gene expression in mammalian cells with promoter-targeted duplex RNAs. Nat ChemBiol 3: 166-173.
- Matsui M(2013) Promoter RNA links transcriptional regulation of inflammatory pathway genes. Nucleic Acids Res 41: 10086-10109.
- Liu J, Hu J, Corey DR(2012) Expanding the action of duplex RNAs into the nucleus: redirecting alternative splicing. Nucleic Acids Res 40: 1240-1250.
- Tripathi V (2010)The nuclear-retained noncoding RNA MALAT1 regulates alternative splicing by modulating SR splicing factor phosphorylation. Mol Cell 39: 925-938.
- AmeyarZazoua M (2012) Argonaute proteins couple chromatin silencing to alternative splicing. Nat StructMolBiol 19: 998-1004.
- Li LC (2006) Small dsRNAs induce transcriptional activation in human cells. ProcNatlAcadSci USA 103: 17337-17342.
- Morris KV, Chan SW, Jacobsen SE, Looney DJ (2004) Small interfering RNA-induced transcriptional gene silencing in human cells. Science 305: 1289-1292.
- Ting AH, Schuebel KE, Herman JG, Baylin SB(2005) Short double-stranded RNA induces transcriptional gene silencing in human cancer cells in the absence of DNA methylation. Nat Genet 37: 906-910.
- Benhamed M, Herbig U, Ye T, DejeanA,BischofO (2012) Senescence is an endogenous trigger for microRNA-directed transcriptional gene silencing in human cells. Nat Cell Biol 14: 266-275.
- Parisi C (2011) Ago1 and Ago2 differentially affect cell proliferation, motility and apoptosis when overexpressed in SH-SY5Y neuroblastoma cells. FEBS Lett 585: 2965-2971.
- Hafner M (2010) Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell 141: 129-141.
- Barrett T N (2011) CBI GEO: archive for functional genomics data sets--10 years on. Nucleic Acids Res 39: 1005-1010.
- Talantov D (2005) Novel genes associated with malignant melanoma but not benign melanocytic lesions. Clin Cancer Res 11: 7234-7242.
- Dome JS, Coppes MJ(2002) Recent advances in Wilmstumor genetics. CurrOpinPediatr 14: 5-11.
- Hartl D (2008) Transcriptome and proteome analysis of early embryonic mouse brain development. Proteomics 8: 1257-1265.
- Pushpavalli SN(2014) Argonaute-1 functions as a mitotic regulator by controlling Cyclin B during Drosophila early embryogenesis. FASEB J 28: 655-666.
- Bertucci F (2012) 8q24 Cancer risk allele associated with major metastatic risk in inflammatory breast cancer. PLoS One 7: e37943.
- Neta G (2012) Common genetic variants in the 8q24 region and risk of papillary thyroid cancer. Laryngoscope 122: 1040-1042.
- Brisbin AG (2011) Meta-analysis of 8q24 for seven cancers reveals a locus between NOV and ENPP2 associated with cancer development. BMC Med Genet 12: 156.
- Witte JS(2007) Multiple prostate cancer risk variants on 8q24. Nat Genet 39: 579-580.
- Hernando E(2001) Molecular analyses of the mitotic checkpoint components hsMAD2, hBUB1 and hBUB3 in human cancer. Int J Cancer 95: 223-227.
- Kidokoro T (2008) CDC20, a potential cancer therapeutic target, is negatively regulated by p53. Oncogene 27: 1562-1571.
- Lu S, Lee J, Revelo M, Wang X, Dong Z (2007) Smad3 is overexpressed in advanced human prostate cancer and necessary for progressive growth of prostate cancer cells in nude mice. Clin Cancer Res 13: 5692-5702.
- Barber TD (2008) Chromatid cohesion defects may underlie chromosome instability in human colorectal cancers. ProcNatlAcadSci USA 105: 3443-3448.
- Joshua Tor L, Hannon GJ (2011) Ancestral roles of small RNAs: an Ago-centric perspective. Cold Spring HarbPerspectBiol 3: 003772.
- Meister G (2013) Argonaute proteins: functional insights and emerging roles. Nat Rev Genet 14: 447-459.
- Volpe TA (2002) Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 297: 1833-1837.
- Zilberman D, Cao X, Jacobsen SE(2003) ARGONAUTE4 control of locus-specific siRNA accumulation and DNA and histone methylation. Science 299: 716-719.
- Ambros V(2004) The functions of animal microRNAs. Nature 43: 350-355.
- Bartel DP(2004) MicroRNAs: genomics biogenesis mechanism and function. Cell 116: 281-297.
- Lim LP(2005) Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 433: 769-773.
- Bentwich I (2005) Identification of hundreds of conserved and non-conserved human microRNAs. Nat Genet 37: 766-770.
- Berezikov E (2005) Phylogenetic shadowing and computational identification of human microRNA genes. Cell 120: 21-24.
- Xu P, Guo M, Hay BA(2004) MicroRNAs and the regulation of cell death. Trends Genet 20: 617-624.
- Karp X, Ambros V(2005) Developmental biology. Encountering microRNAs in cell fate signaling. Science 310: 1288-1289.
- EsquelaKerscher A, Slack FJ (2006) Oncomirs - microRNAs with a role in cancer. Nat Rev Cancer 6: 259-269.
- Volinia S (2006) A microRNA expression signature of human solid tumours defines cancer gene targets. ProcNatlAcadSci USA 103: 2257-2261.
- Lee Y (2004) MicroRNA genes are transcribed by RNA polymerase II. EMBO J 23: 4051-4060.
- Lee Y (2003) The nuclear RNase III Drosha initiates microRNA processing. Nature 425: 415-419.
- Denli AM, Tops BB, Plasterk RH, Ketting RF, Hannon GJ (2004) Processing of primary microRNAs by the Microprocessor complex. Nature 432: 231-235.
- Yi R, Qin Y, Macara IG, Cullen BR (2003) Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes Dev 17: 3011-3016.
- Bernstein E, CaudyAA, Hammond SM, Hannon GJ(2001) Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409: 363-366.
- Gregory RI, Chendrimada TP, Cooch N, Shiekhattar R (2005) Human RISC couples microRNA biogenesis and posttranscriptional gene silencing. Cell 123: 631-640.
- Yue J, Tigyi G(2006) MicroRNA trafficking and human cancer. Cancer BiolTher 5: 573-578.
- Karube Y (2005) Reduced expression of Dicer associated with poor prognosis in lung cancer patients. Cancer Sci 96: 111-115.
- Kumar MS, Lu J, Mercer KL, Golub TR, Jacks T(2007) Impaired microRNA processing enhances cellular transformation and tumorigenesis. Nat Genet 39: 673-677.
- Carmell MA, Xuan Z, Zhang MQ, Hannon GJ (2002) The Argonaute family: tentacles that reach into RNAi developmental control stem cell maintenance and tumorigenesis. Genes Dev 16: 2733-2742.
- Benkirane M (1997)Oncogenic potential of TAR RNA binding protein TRBP and its regulatory interaction with RNA-dependent protein kinase PKR. EMBO J 16: 611-624.
- Spitz MR, Wei Q, Dong Q, Amos CI Wu X (2003) Genetic susceptibility to lung cancer: the role of DNA damage and repair. Cancer Epidemiol Biomarkers Prev 12: 689-698.
- Schwamborn JC, Berezikov E, Knoblich JA (2009) The TRIM-NHL protein TRIM32 activates microRNAs and prevents self-renewal in mouse neural progenitors. Cell 136: 913-925.
- Carmichael JB, Provost P, Ekwall K, Hobman TC (2004) ago1 and dcr1, two core components of the RNA interference pathway, functionally diverge from rdp1 in regulating cell cycle events in Schizosaccharomycespombe. MolBiol Cell 15: 1425-1435.
- Stoica C, Carmichael JB, Parker H, Pare J, Hobman TC (2006) Interactions between the RNA interference effector protein Ago1 and 14-3-3 proteins: consequences for cell cycle progression. J BiolChem 281: 37646-37651.
- Morel JB (2002) Fertile hypomorphic ARGONAUTE (ago1) mutants impaired in post-transcriptional gene silencing and virus resistance. Plant Cell 14: 629-639.
- Bueno MJ, Malumbres M (2011) MicroRNAs and the cell cycle. BiochimBiophysActa 1812: 592-601.
- Suzuki HI, Miyazono K(2010) Dynamics of microRNA biogenesis: crosstalk between p53 network and microRNA processing pathway. J Mol Med 88: 1085-1094.
- Boominathan L(2010) Thetumour suppressors p53, p63, and p73 are regulators of microRNA processing complex. PLoS One 5: e10615.
- De N (2013)Highly complementary target RNAs promote release of guide RNAs from human Argonaute2. Mol Cell 50: 344-355.
- Gagnon KT, Corey DR (2012)Argonaute and the nuclear RNAs: new pathways for RNA-mediated control of gene expression. Nucleic Acid Ther 22: 3-16.
- Burroughs AM (2011) Deep-sequencing of human Argonaute-associated small RNAs provides insight into miRNA sorting and reveals Argonaute association with RNA fragments of diverse origin. RNA Biol 8: 158-177.
- Polikepahad S, Corry DB (2013) Profiling of T helper cell-derived small RNAs reveals unique antisense transcripts and differential association of miRNAs with argonaute proteins 1 and 2. Nucleic Acids Res 41: 1164-1177.
- Ebert MS, Sharp PA (2012) Roles for microRNAs in conferring robustness to biological processes. Cell 149: 515-524.
- Junxia W (2010) Double strand RNA-guided endogeneous E-cadherin up-regulation induces the apoptosis and inhibits proliferation of breast carcinoma cells in vitro and in vivo. Cancer Sci 101: 1790-1796.
- Wei J (2010) p21WAF1/CIP1 gene transcriptional activation exerts cell growth inhibition and enhances chemosensitivity to cisplatin in lung carcinoma cell. BMC Cancer 10: 632.
- Place RF (2012) Formulation of Small Activating RNA IntoLipidoid Nanoparticles Inhibits Xenograft Prostate Tumour Growth by Inducing p21 Expression. MolTher Nucleic Acids 1: e15.
- Kang MR (2012)Intravesical delivery of small activating RNA formulated into lipid nanoparticles inhibits orthotopic bladder tumor growth. Cancer Res 72: 5069-5079.
- Hannon GJ (2002) RNA interference. Nature 418:244-251.
Relevant Topics
- Breast Cancer Surgery
- Colon Cancer Surgery
- Dermatologic Surgery
- Kidney Cancer Surgery
- Leukemia Surgery
- Lung Cancer Surgery
- Lymphoma Surgery
- Oesophageal Cancer Surgery
- Pancreatic Cancer Surgery
- Prostate Cancer Surgery
- Radiation Therapy
- Skin Cancer Surgery
- Stomach Cancer Surgery
- Throat Cancer Surgery
- Thyroid Cancer Surgery
Recommended Journals
Article Tools
Article Usage
- Total views: 13156
- [From(publication date):
June-2016 - Aug 31, 2025] - Breakdown by view type
- HTML page views : 12145
- PDF downloads : 1011