Research Article Open Access
Development of Methods for Immobilization and Usages of Immobilized Recombinant Bioluminescent Strain, Pseudomonas Putida Mt-2 KG1206, Stored by Deep-Freezing
Kim M1 and Kong IC2*1Infra Technology Team, Samsung Display Co., Ltd, Chungnam 336-741, Korea
2Department of Environmental Engineering, Yeungnam University, Kyungbuk 38541, Korea
- Corresponding Author:
- In Chul Kong
Department of Environmental Engineering
Yeungnam University
Kyungbuk 38541, Korea
Tel: +82-53-810-2546
Fax: +82- 53-810-4624
E-mail: ickong@ynu.ac.kr
Received date: September 05, 2015; Accepted date: November 20, 2015; Published date: November 27, 2015
Citation: Kim M, Kong IC (2015) Development of Methods for Immobilization and Usages of Immobilized Recombinant Bioluminescent Strain, Pseudomonas putida Mt-2 KG1206, Stored by Deep-Freezing. J Biotechnol Biomater 5:208. doi:10.4172/2155-952X.1000208
Copyright: © 2015 Kim M, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Journal of Biotechnology & Biomaterials
Abstract
Deep-freezing process can be used practically for environmental monitoring. However, for the organism to be useful in environmental applications, the strain of bacteria exhibits a high intensity of activity following reconstitution after deep-freezing. An overall protocol for immobilization and reconstitution of recombinant bioluminescent bacteria, preserved by deep-freezing was investigated for future biomonitoring. Tested strain KG1206 has ability to produce bioluminescence in the presence of toluene analogs and those intermediates. Immobilization using glass beads with subcultured cells showed better results than other conditions, such as alginate beads or centrifuged cell pellets. Beads number, thawing time, and amount of reconstitution solution influence on bioluminescence activity. Among tested conditions, followings were appeared as optimum conditions: working volume 15 mL of serum vials, approximately 20-30 glass beads per vial, 3 h thawing time, and 2-5 mL reconstitution solution. Potassium nitrate (KNO3) greatly stimulates the low bioluminescence activity of immobilized strain, preserved by deep freezing, in the range of 2.5 to 21 times of untreated one.
Keywords
Bioluminescence; Biomonitoring; Deep-freezing; Immobilization; Recombinant
Significance and Impact of the Study
This study indicated that type of immobilization matrix, cell conditions (subcultured or centrifuged pellets), matrix (beads) numbers, volume of reconstitution solution, and thawing time are important factors on bioluminescence activity of reconstituted culture, preserved by deep freezing. Overall, this investigation demonstrated the ability of immobilized bacteria on glass beads, preserved by deep freezing, for the monitoring of a specific group of environmental pollutants using a stimulant KNO3, suggesting the potential method for preliminary application in a field-ready portable bioassay.
Introduction
Petroleum hydrocarbons (PHCs) are one of the most widely used compound with huge amounts of them used for powering vehicles and heating. Leaking and spill accidents related to those compounds have become a rising problem that cannot be neglected. Petroleum consists of hazardous chemicals including benzene, toluene, ethylbenzene, xylenes and naphthalene which are very toxic not only to plants and animals, but also to humans [1-3]. As the number of reports on the pollution of soil and groundwater caused by PHCs has increased, the United States Environmental Protection Agency (USEPA), the Quebec Ministry of Environment (MENV), and the Organizations of Environment and Public Health around the world have treated PHCs as a significant contaminant [4].
Genetically engineered bioluminescent bacteria, such as Pseudomonas putida mt-2 KG1206, have shown a high feasibility for use as a tool for biomonitoring PHCs in environments [5]. The tested recombinant strain KG1206 has been shown to contain an intact TOL plasmid, Pm-lux fusion plasmid, and promoter Pm as the lower (meta) operon (13 genes) of the TOL catabolic plasmid pWW0, and it is responsible for producing bioluminescence in the presence of toluene analogs as well as their metabolites [6]. Intermediates of toluene analogs, such as benzoate and m-toluate, can activate XylS regulatory protein, whereas toluene, m-methyl benzyl alcohol (m-MBA), and isomers of xylene can activate XylR regulatory protein. Both XylS and XylR proteins positively regulate promoters. XylR controls the upper pathway promoter Pu as well as induces expression of the xylS gene, which is responsible for the production of XylS protein and regulating the lower pathway promoter Pm. For KG1206 strain, inducer chemicals bind to inactive XylS protein and induce a conformational change in the protein that results in enhanced interaction with the promoter site. This active protein then binds to the promoter Pm and produces bioluminescence from the Pm-lux gene in the recombinant strain [7].
There are many reports of using immobilization method for microorganisms and enzymes in biological technologies. Immobilization is done by the four different methods: carrier-binding, cross-linking, entrapping and combination of other three [8]. Alginate, carrageenan, polyacrylamide and glass beads are commonly used as a carrier for the cells [9,10]. Deep-freezing is a general method for preserving bacteria for extended periods [11], and can be used practically for the portable biosensor in the field. However, these processes can affect the activity of strains that have been reconstituted after deep-freezing. Therefore, for the organism to be useful in environmental applications, it is very important that the strain of bacteria used for monitoring exhibits a high intensity of bioluminescence that can be maintained following reconstitution after deep-freezing.
The primary objective of this study is to determine a useful method for the immobilization of bioluminescence producing bioreporter strain in the presence of several inducer pollutants, and then subsequently evaluate the conditions for reconstitution of immobilized strains, preserved by deep-freezing, for future usages on biomonitoring.
Materials and Methods
Strain preparation and analysis
Pseudomonas putida mt-2 KG1206 were stored at -70°C until needed, at which they were grown overnight in Luria-Bertani (LB) medium supplemented with 50 mg/L of kanamycin at 27°C with shaking (130 rpm). Next, a 1:30 dilution was made using LB medium, and the culture was then allowed to grow until it had an optical density (OD600) of approximately 0.6. The LB medium consisted of 10g tryptone, 5g yeast extract, 5g NaCl and 0.5 mL 2.0 N NaOH per L of broth.
The KG1206 strain produces bioluminescence in the presence of appropriate inducing compounds and is also capable of degrading analogs and catabolic intermediates of toluene, including m-toluate and benzoate. The bioluminescence of KG1206 was measured by luminometer 20/20 (Turner Design, USA). It shows relative light units (RLUs) which mean the sum of the bioluminescence released from a sample. The TD 20/20 had 110.3 RLU factory standard reading and 51.5% sensitivity.
Immobilization and deep-freezing protocol for the strain
Four procedures were compared to develop efficient immobilization protocol: (method 1) Glass beads (3mm diameter) were prepared in a vial and autoclaved. Equal volume of KG1206 culture (OD600=0.6) and 24% sucrose were completely mixed and added to the vial containing beads, and continuously soaked over 20 min. After decanting the solution, the glass beads were stored at -70°C until needed. (method 2) The subcultured KG1206 (OD600=0.6) was centrifuged (3,000 rpm for 15min) and the supernatant was discarded (x 2). The pellet was resuspended in LB medium and equally mixed with 24% sucrose. This mixture was added to the vial containing glass beads and soaked (>20 min). After decanting the liquid, glass beads were stored at -70°C until needed. (method 3) The mixture of KG1206 culture (OD600=0.6) and 24% sucrose was added into an equal volume of 2% alginate. The mixture (2 mL) was injected into cold 0.1% CaCl2 solution (4 mL) to form alginate beads and left in a refrigerator for 2 h to allow strong bead formation. After decanting the CaCl2 solution, the alginate beads were stored at -70°C until needed. (method 4) The equal volume of 2% alginate solution was added into mixture of resuspended pellet solution and 24% sucrose (1:1). The final mixture was injected into 0.1% cold CaCl2 solution (4 mL). After decanting the solution, the alginate beads were stored at -70°C until needed. Following reconstitution of stored deep-freezing cells, the glass or alginate beads were amended with 1mM of m-toluate for 3 h to determine bioluminescence activity of each method.
Various conditions were further investigated. Firstly reasonable beads number for immobilization was determined at conditions of 10, 20, 30, and 50 beads. The appropriate quantity of reconstitution solution was investigated at conditions of 2, 5, and 10 mL of LB medium by adding into the deep-freezed beads (-70°C). Reasonable reconstitution (thawing) time was determined at conditions of 0.5, 1, 3, and 5 h. All tests were performed triplicates using 15 mL working volume of serum vials. Bioluminescence activity of each condition was evaluated in the presence of 1 mM of m-toluate.
Effect of enhancer on bioluminescence activity of reconstituted beads
Stimulating chemicals on bioluminescence activity of reconstituted beads were investigated. Immobilized beads stored in deep-freezing were reconstituted with LB medium for 3 h. KNO3 (0.04 g:20 g chemical/L) was added to a vial containing reconstituted immobilized beads. Effects on bioluminescence production were compared between sets of with and without stimulant, in the presence of inducer chemical. Following inducer chemicals were tested: 1 mM of m-MBA, toluene, three xylene isomers, and m-toluate).
Results and Discussion
Immobilization protocol of the recombinant strain
Protocols for immobilization and reconstitution of deep freezed immobilized strain KG1206 were investigated to conveniently use on site monitoring. KG1206 contains the intact TOL plasmid and a Pmlux fusion plasmid. Pm is the promoter of the lower or meta operon (13 genes) of the TOL catabolic plasmid, pWW0, and is responsible for producing bioluminescence in the presence of toluene analogs and their metabolites [6].
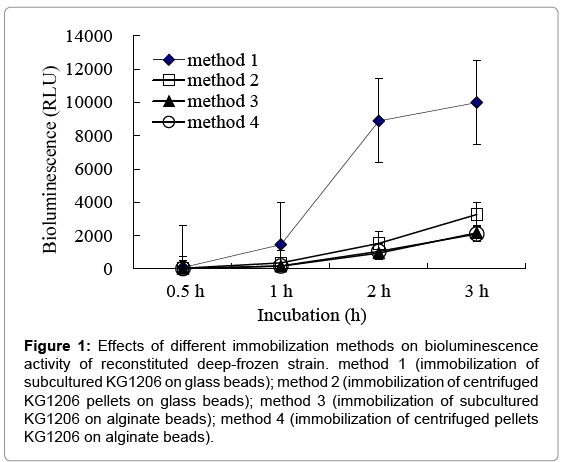
Bioluminescence activities of reconstituted strains, which were immobilized by different protocols, were measured in the presence of 1 mM m-toluate, and the results were compared. For deep-freezing process, Surcose, known as a good cryprotectant, was used. Before being reconstitution, the vials containing immobilized strain were stored in a deep-freezer at -70°C at least for 12 h. Effects of different immobilization methods on bioluminescence activity of reconstituted deep-frozen strain are shown in Figure 1. Method 1 and 2 (immobilized in glass beads) showed higher bioluminescent activity than that in alginate beads, indicating 1,536 ± 433.5 and 8,893 ± 340.8 RLU at 2 h incubation for method 2 and method 1 (1.5-8.4 times of sets with alginate beads), respectively. Moreover, reconstituted strain in method 1 (immobilization on glass beads with subcultured cells) showed great bioluminescence activity compared to others. Statistically significant differences were observed (p=0.0001) compared to others. This may be caused by the different immobilization characteristics between two beads: attached on the surface for glass beads (3 mm), entrapped inside for alginate beads (less than 1 mm). Entrapped cells in alginate beads may have difficulty obtaining proper growing conditions, such as nutrient or inducers, etc. Also, sets of immobilization with centrifuged pellets showed lower bioluminescence activities than ones with subcultured cells, where used directly on immobilization without centrifugation. This could be considered due to cell loss, disruption, or nutrient loss during centrifugation or resuspension processes, possibly loss of the exponential growth activity. Therefore, for all following investigations, equal mixtures of KG1206 culture (OD600=0.6; without centrifugation) and 24% sucrose were completely mixed and added to the vial (15 mL working volume of serum vials) containing glass beads, and continuously soaked over 20 min (method 1). After decanting the solution, the glass beads were stored at -70°C until needed.
Figure 1: Effects of different immobilization methods on bioluminescence activity of reconstituted deep-frozen strain. method 1 (immobilization of subcultured KG1206 on glass beads); method 2 (immobilization of centrifuged KG1206 pellets on glass beads); method 3 (immobilization of subcultured KG1206 on alginate beads); method 4 (immobilization of centrifuged pellets KG1206 on alginate beads).
Of the tested chemicals, m-toluate is known to directly activate XylS protein, while the other compounds are activators of XylR, and converted to lower pathway inducers during catabolism. As expected, the reconstituted strain clearly produced less bioluminescence (weak expressions) compared to those found with the subcultured strain, but fully detectable bioluminescence did appear. It is possible that this decreased activity was due to cell damage during the deep freezing and reconstitution processes [12]. The magnitude of bioluminescence of reconstituted strain appeared in the following order: p-xylene ≈ m-toluate>toluene ≈ m-MBA>m-xylene>o-xylene. Different patterns were also observed with the subcultured (not immobilized) strain. In case of subcultured strain, generally, toluene was strong inducer, while p-xylene weak inducer. In this investigation, p-xylene, the weakest inducer in subcultured strain, appeared as the strongest one in reconstituted immobilized strain, preserved by deep-freezing. The cause of this contrary result is not clear at this point, and the cause of this result has to be investigated more specifically on a molecular basis, possibly.
Development of reconstitution and immobilization protocols of the recombinant strain
Reconstitution process of immobilized strain, preserved by deepfreezing also can affect bioluminescence activity. Effects of different amount of reconstitution solution on bioluminescence activity were examined by adding 2 to 10 mL LB medium in vials containing immobilized strain, preserved by deep-freezing (Figure 2). As shown in Figure 2, vials amended with 2 and 5 mL, which showed similar results, produced considerably higher bioluminescence than that in 10 mL. At incubation time 2 h, sets amended with 2, 5, and 10 mL reconstituting medium produced bioluminescence 8,893 ± 340.8, 8,205 ± 2537.8, and 2,050 ± 912.9 RLU, respectively. Bioluminescence activities of sets with 2 or 5 mL reconstitution solution also showed statistically significant differences compared to set with 10 mL reconstitution solution (p value in the range of 0.002-0.017).
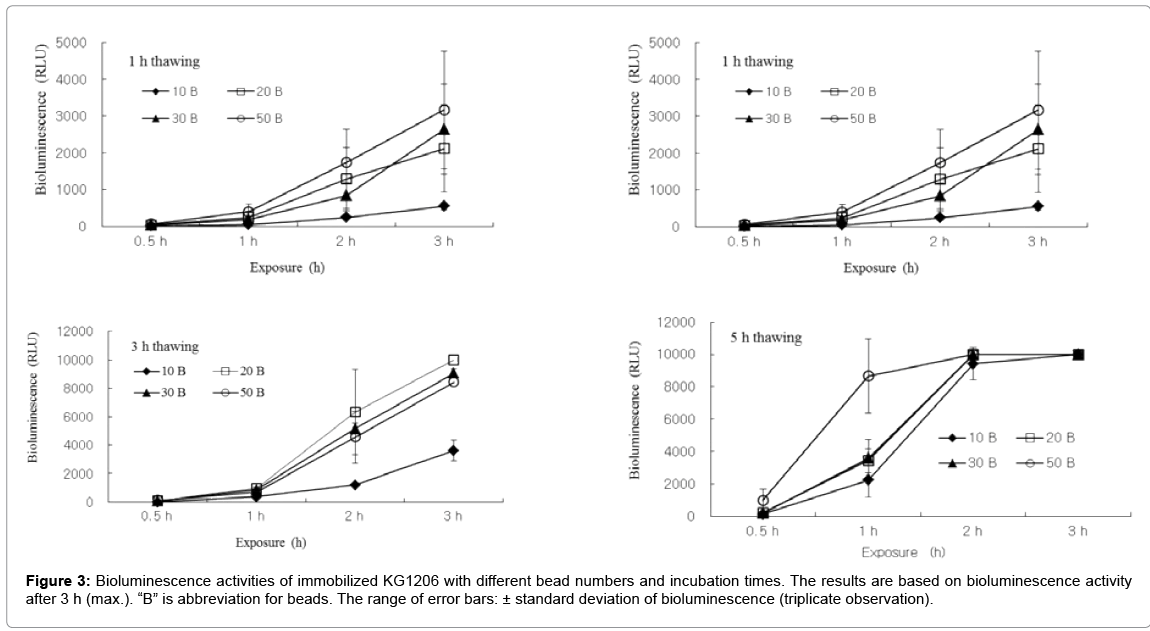
Appropriate beads number and reconstitution thawing time for test also were investigated at following conditions: bead number 10, 20, 30, and 50 glass beads, thawing time 0.5, 1, 3, and 5 h. As shown in Figure 3, sets with 10 beads showed low bioluminescence (<500 RLU), except 5 h thawing time. Sets with short thawing time (0.5 h) also produced low bioluminescence during incubation period compared to other conditions. In contrast, sets with 5 h thawing time produced too high bioluminescence to observed, showing >9,999 RLU (max. detection limit) after 2 h incubation time. Results indicated that sets with thawing time for 1 and 3 h with 20 to 30 beads provide high enough bioluminescence activity that is possible to be compared in different variations. For example, sets with 1 and 3 h thawing time with 20 beads showed 1,285 ± 858.2 and 6,322 ± 3003.0 RLU after 2 h incubation, respectively. Also, sets with 1 and 3 h thawing time with 30 beads showed 833 ± 347.7 and 5,159 ± 385.4 RLU after 2 h incubation, respectively. In this study, the greater activities were generally observed at longer reconstitution time. Reconstitution time around 2 h or 3 h seemed as being reasonable to obtain proper bioluminescence intensity. Following reconstitution, at least 2 h exposure on samples was required to observe proper bioluminescence activity.
This investigation found out that the beads number and amount of reconstitution solution on each vial also affect on bioluminescence activity of the immobilized strain, stored by deep-freezing. Based on this experimental condition, numbers of 20-30 beads and 2-5 mL of reconstitution solution in 15 mL volume vials appeared as optimum settings to obtain proper bioluminescence activity for biomonitoring. Therefore, those parameters should be carefully determined depending on each experimental condition.
Effects of enhancer on bioluminescence activity of reconstituted beads
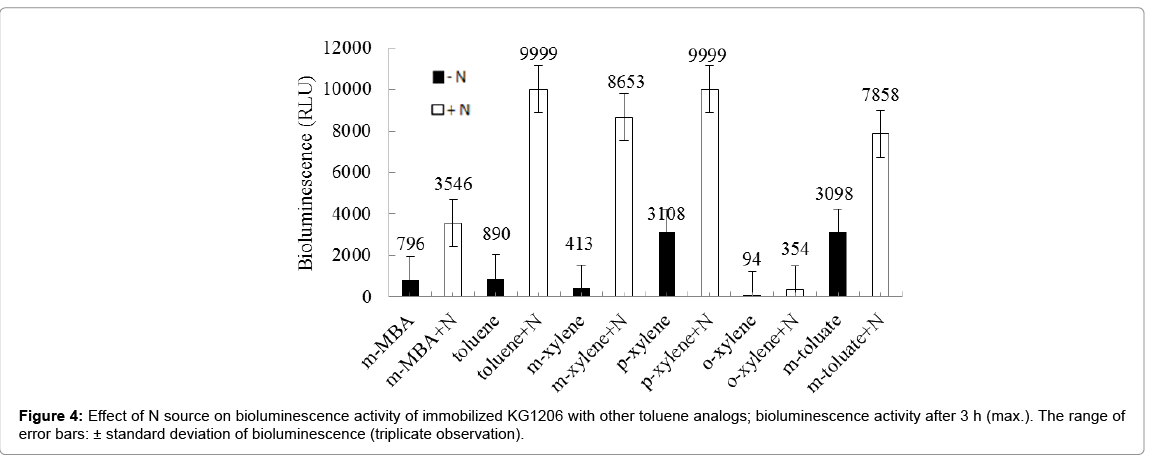
The stimulation of the reconstituted strain, showing low bioluminescence activity, is necessary to use on environmental monitoring. In previous author’s investigation with various nitrogen compounds, greater stimulation by nitrate forms, KNO3 and NH4NO3 was observed, while a complete inhibition was with KNO2. Also no significant effects were observed with other ammonium forms, such as (NH4)2SO4 and (NH4)2C2H4H2O (Kong 2010). In addition, better stimulation was mostly observed with KNO3 compared to other good stimulant, sodium lactate. Based on these results, the influence of addition of KNO3 on the reconstituted immobilized strain, stored by deep-freezing was investigated in the presence of various inducers. The reconstituted immobilized strain showed different bioluminescence activity, showing 796, 890, 94, 413, 3,108, and 3,098 RLU for m-MBA, toluene, o-, m-, p-xylenes, and m-toluate, respectively, at 3 h incubation time. Stimulations by KNO3 were within the range of 2.5- 21.0 (avg. 7.7 ± 7.22) times those of the control (Figure 4). Relative stimulation intensity varied depending on the inducer chemicals. The highest relative stimulation was observed with toluene and m-xylene, stimulated from 413 RLU and 8,653 (21 times) RLU at 3 h incubation time, respectively, but the highest bioluminescence activity was observed with p-xylene inducer, increased from 3,108 (control) to 9,999 (with KNO3) RLU. In contrast, the lowest relative stimulation was observed with m-toluate, stimulated from 3,098 to 7,858 RLU (2.5 times). Different order of bioluminescence intensity was appeared with addition of stimulant, showing in the order of p-xylene ≈ toluene>mxylene ≈ m-toluate>m-MBA>o-xylene.
The stimulating chemicals may have caused positive effects on the bioluminescence regulatory systems, such as the rpo-S gene or other unidentified regulatory systems [13]. A number of mechanisms could potentially bring about the stimulation and inhibition effects, ranging from changes in promoter DNA supercoiling, which may affect the gene expression in response to nutritional changes, to the specific RNA polymerase subunit, σ54, for transcription [14].
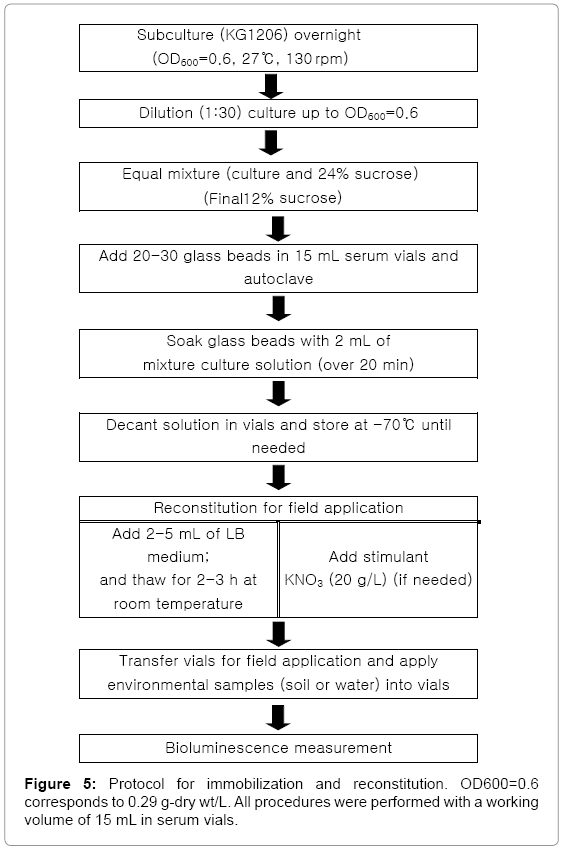
Based on all tested conditions, the following procedure was determined for future usages of the recombinant bioluminescence bacteria on in situ biomonitoring (Figure 5). These findings might be applicable to other recombinant bioluminescence strains for environmental monitoring. Overall, this investigation demonstrated the ability of immobilized bacteria on glass beads, preserved by deep freezing, for the monitoring of a specific group of environmental pollutants using a stimulant KNO3, suggesting the potential method for preliminary application in a field-ready portable bioassay.
References
- Liebeg EW, Cutright TJ (1999) The investigation of enhanced bioremediation through the addition of macro and micronutrients in a PAH contaminated soil. IntBiodetBiodeg 44: 55-64.
- Ting YP, Hu HL, Tan HM (1999) Bioremediation of petroleum hydrocarbons in soil microcosms. Resou Environ Biotechnol 2: 197-218.
- Vasudevan N, Rajaram P (2001) Bioremediation of oil sludge-contaminated soil. Environ Int 26: 409-411.
- Yerushalmi L, Rocheleau S, Cimpoia R, Sarrazin M, Sunahara G, et al. (2003) Enhanced biodegradation of petroleum hydrocarbons in contaminated soil. Bioremed J 7: 37-51.
- Kong IC (2010) Application of stimulating agents on the immobilized bioluminescence strain Pseudomonas putida mt-2 KG1206, preserved by deep-freezing, for the convenient monitoring. J Environ Sci 22: 1475-1480.
- Harayama S, Rekik M (1990) The meta cleavage operon of TOL degradative plasmid pWW0 comprises 13 genes. Mol Gen Genet 221: 113-120.
- Inouye S, Nakazawa A, Nakazawa T (1987) Overproduction of the xylS gene product and activation of the xylDLEGF operon on the TOL plasmid. J Bacteriol 169: 3587-3592.
- Tanaka A, Kawamoto T (1999) Cell and Enzyme Immobilization. In: Manual of industrial Microbiology and biotechnology, Demain AL and Davies JE (Eds.), ASM Press, USA.
- Cheetham PSJ, Bucke C (1984) Immobilization of microbial cells and their use in waste water treatment. In: Microbiological Methods for Environmental Biotechnology. Academic Press.
- Stormo KE, Crawford RL (1992) Preparation of encapsulated microbial cells for environmental applications. Appl Environ Microbiol 58: 727-730.
- Gu MB, Choi SH, Kim SW (2001) Some observations in freeze-drying of recombinant bioluminescent Escherichia coli for toxicity monitoring. J Biotechnol 88: 95-105.
- Choi SH, Gu MB (2003) Toxicity biomonitoring of degradation byproducts using freeze-dried recombinant bioluminescence bacteria. AnalytChimicaActa 481: 229-238.
- Miura K, Inouye S, Nakazawa A (1998) TherpoS gene regulates OP2, an operon for the lower pathway of xylene catabolism on the TOL plasmid, and the stress response in Pseudomonas putida mt-2. Mol Gen Genet 259: 72-78.
- Assinder SJ, Williams PA (1990) The TOL plasmids: determinants of the catabolism of toluene and the xylenes. AdvMicrobPhysiol 31: 1-69.
Relevant Topics
- Agricultural biotechnology
- Animal biotechnology
- Applied Biotechnology
- Biocatalysis
- Biofabrication
- Biomaterial implants
- Biomaterial-Based Drug Delivery Systems
- Bioprinting of Tissue Constructs
- Biotechnology applications
- Cardiovascular biomaterials
- CRISPR-Cas9 in Biotechnology
- Nano biotechnology
- Smart Biomaterials
- White/industrial biotechnology
Recommended Journals
Article Tools
Article Usage
- Total views: 11559
- [From(publication date):
December-2015 - Aug 29, 2025] - Breakdown by view type
- HTML page views : 10601
- PDF downloads : 958