DNA Barcoding of Tilapia Species (Pisces: Cichlidae) from North-Eastern Nigeria
Received: 02-Oct-2017 / Accepted Date: 14-Nov-2017 / Published Date: 21-Nov-2017 DOI: 10.4172/2155-952X.1000277
Abstract
The Tilapiine complexes are made up of diverse species, hence the need for proper identification. Samples of tilapia species belonging to the family of Cichlidae obtained from Upper Benue River and Lake Geriyo landing sites were used for this study. Fish species were morphologically identified in situ by visual inspection with the aid of field guide for taxonomic studies. DNA was extracted from fresh fish tissue and the barcode gene region cytochrome c oxidase subunit 1 (co1) was amplified using the FishF1 and FishR1 primer pair. Amplified products were visualized on 1% agarose gel electrophoresis, purified and sequenced using Genetic Analyzer (Applied Biosystems 3130 XL). Four species (Oreochromis niloticus, Sarotherodon galilaeus, Sarotherodon galilaeus boulengeri and Coptodon zillii) belonging to three genera (Oreochromis, Sarotherodon and Coptodon) were identified in the study area. From our sampling survey, Oreochromis niloticus was the most dominant species in both areas, followed by Coptodon zillii. Sarotherodon spp. was not present in Upper Benue River but present in Lake Geriyo (an ox-bow lake of Upper Benue River). A new sub-species of Sarotherodon galilaeus known as Sarotherodon galilaeus boulengeri was identified in this study from the sequencing result. This species has never been reported in any literature to be present in Nigeria, which further confirmed the efficacy of DNA barcoding in the identification of cryptic species. The phylogenetic relationship and genetic distance between generic groups obtained in this study was in concordance with the hypothesis of Trewavas that the mouth brooders Orechromis niloticus and Sarotherodon galilaeus are more closely related to each other than the substrate brooders (Tilapia zillii).
Keywords: CO1; DNA Barcoding; Phylogenetics; Tilapia
Introduction
The distinction of species within the tilapiine genera is “notoriously difficult” [1] and most published descriptions of the species are based on their morphological characteristics which “show considerable variation and broad interspecies overlaps” [2-7]. Often, geography has played a huge role in the identification of species [1]. Over the past 3 to 5 decade, species identification has become more difficult by the extensive introduction of alien species and transfer of native from outside of their natural ranges, for example in Lake Victoria [8,9]. This introduction has led to inter-specific hybridization, and the emergence of hybrid forms making morphological identification of species clearly impossible [10- 14]. Incorrect description and misidentification of species and possible hybrids using the conventional methods call for better molecular techniques.
Little is known about the relationships among the tilapiine species, despite the fact that they have become an important component of African Aquaculture [1]. DNA marker technologies have revolutionized the way aquaculture genetics research is conducted [15]. Due to the advances in the sequencing and computational technologies, DNA sequences have become the major source of new information for advance understanding of evolutionary and genetic relationships [16]. Species identification through barcoding is usually achieved by the retrieval of a short DNA sequence (the barcode) from a standard part of the genome (i.e., a specific gene region, cytochrome oxidase 1), from the specimen under investigation [16]. The barcode sequence from the unknown specimen is the compared with a library reference barcode sequences derived from individuals of unknown identity (http://www.barcodinglife.org). Hence, this study was aimed to identify tilapia species of family cichlidae from Upper River Benue and Lake Geriyo in the northeastern part of Nigeria using DNA barcoding.
Materials And Methods
Study area
The study areas used for this study are Lake Geriyo (Latitude 9.30°N and Longitude 12.40°E) and Upper River Benue fish landing site (Latitude 9.29°N and Longitude 12.47°E). DNA extraction, polymerase chain reaction and gel electrophoresis was carried out at the Chevron Biotechnology Centre Molecular Biology Laboratory, Modibbo Adama University of Technology, Yola.
Sample collection and initial identification
All specimens were caught in the wild and morphologically identified in situ by visual inspection. Taxonomic classification was done with the aid of the field guide to Nigerian freshwater fish [17] and freshwater fishes of Nigeria [18]. Fresh/frozen muscle and pectoral fin was collected from the identified fishes.
DNA extraction
DNA extraction was carried out using Zymo Research Genomic DNA™-Tissue MiniPrep extraction kit, and the manufacturer protocol was followed. Briefly, in a clean 1.5 ml microcentrifuge tube, 25 mg of fish tissue was added. 95 μl of RNase free water, 95 μl of 2x digestion buffer and 10 μl proteinase k was added to the tube. The mixture was incubated in a water bath set at 55°C for 3 h. RNase treatment was performed by adding 5 μl of RNase enzyme and incubated at 37°C for 15 min.
Polymerase chain reaction (PCR) amplification
The CO1 gene located in the mitochondrial genome was amplified using the pair of primers FishF1: TCAACCAACCACAAAGACATTGGCAC and FishR1: TAGACTTCTGGGTGGCCAAAGAATCA [19]. PCR amplification was performed with Eppendorf Mastercycler personal in 50 μl total reaction volume using OneTaq Quick-Load 2X Master Mix with Standard Buffer (New England BioLabs Inc) ready-to-use kit as shown in Table 1. The PCR was carried out under the following conditions: an initial denaturation of 94°C for 30 s, followed by 35 cycles of 94°C for 30 s, 60°C for 1 min, 68°C for 1 min; followed by final extension of 68°C for 10 min and held at 20°C. The PCR products were visualized on 1% agarose gel electrophoresis stained with ethidium bromide.
| Component | 50 µl Reaction | Final Concentration |
|---|---|---|
| 2x Mastermix | 25 µl | 1x |
| 10 µM Forward Primer | 1 µl | 0.2 µM |
| 10 µM Reverse Primer | 1 µl | 0.2 µM |
| Template DNA | 3 µl | |
| Nuclease-Free Water | 20 µl |
Table 1: PCR setup.
Purification of PCR products and sequencing
Zymo Research DNA clean and concentrator™-5 was used to purify the PCR products. In a 1.5 ml microcentrifuge tube, DNA biding buffer was added to the PCR products in a ratio of 5:1. The mixture was transferred to a Zymo-spin™ column in a collection tube. The sample was centrifuged for 30 s at 10,000x g and the flow-through was discarded. To the spin column, 200 μl DNA wash buffer was added and centrifuge at 10,000x g for 30 s. The wash step was repeated. DNA elution buffer was added at a volume of 15 μl directly to the column matrix and incubated at room temperature for a minute. This was transferred to a 1.5 μl microcentrifuge tube and centrifuge for 30 s at 10,000x g to elute the DNA. The purified PCR products were sequenced using Genetic Analyzer (Applied Biosystems 3130 XL). Cycle sequencing was carried out using the Big Dye Terminator 3.1 Cycle Sequencing Kit (Applied Biosystems).
Sequence analysis
The sequencing was subjected to Barcode of Life Database Identification (BOLD-ID) and BLAST algorithm from the National Centre of Biotechnology Information, Bethesda, MD, USA (http://www. ncbi.nlm.nih.gov) to retrieve for homologous sequences in GenBank. Sequences were edited in BioEdit [20] and aligned using ClustalX 2.1 software [21]. Phylogenetic and analyses were conducted using MEGA 6 [22]. A Maximum Likelihood tree of K2P distances was created to provide a graphic pattern of divergence between the species [23]. A minimum of 1000 bootstrap replications was used for distance creation [22]. Sample identification based on sequence similarity approach was carried out using Barcoding of Life Database (BOLD) (http://www. barcodinglife.org) and GenBank (http://www.ncbi.nlm.nih.gov). All sequences were submitted in GenBank under the accession number KY784669-KY784688, a sequence of Citharinus citharus (GenBank: HQ927822) was used as an out-group.
Results
PCR and sequencing
Similarities between sequences were calculated using built in algorithms for local sequence alignments. Both GenBank and BOLDIDS databases revealed definitive identity matches in the range of 98%- 100% for consensus sequences of all the species. In each case, all queried sequence returned a top hit for the morphologically identified species. One of the species morphologically identified as Sarotherodon galilaeus returned a hit of 98% and 100% for Sarotherodon galilaeus boulengeri in GenBank and BOLD-IDS respectively.
Phylogenetic relationship and sequence divergence
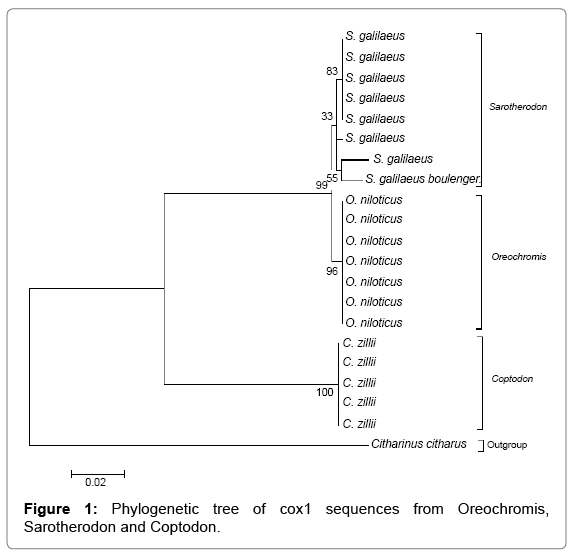
Multiple sequence alignment w carried out using ClustalX 2.1 [21] and subsequently converted to MEGA format using MEGA 6 before constructing the Maximum Likelihood tree with a bootstrap of 1000. The tree is drawn to scale, with branch lengths measured in the number of substitutions per site. From the phylogenetic tree shown in Figure 1 the species clustered independently within their corresponding genera (i.e., Sarotherodon, Oreochromis and Coptodon). The Coptodon zilli formed a more cohesive group sharing common ancestor with a bootstrap value of 100%, Oreochromis niloticus were held together by a bootstrap value of 96%. The Sarotherodon species formed three groups within a cluster and the sequences for this genera showed more variation than the other two (Coptodon and Oreochromis). The overall means distance of the CO1 sequences for the three generic groups is 0.081. A higher level of divergence was observed in Sarotheron spp. against Coptodon zillii (0.150), Oreochromis niloticus and Sarotherodon spp. showed the lowest level of divergence (0.010), while Oreochromis niloticus and Coptodon zillii showed a divergence of 0.145.
Discussion
This study has justified the use of mitochondrial DNA CO1 barcodes for identification of fish species and their relationship. All the sequences obtained showed a clear distinction of the three species studied. The result was in concordance with the work of Wu and Yang [24] where the CO1 gene was used to identify captive and wild tilapia species. Similarly, Hubert et al. [25] and Mohanty et al. [26] used the CO1 gene in distinguishing North American freshwater fishes and cultivable carps in India, respectively. In Nigeria, Nwani et al. [27] used the gene to distinguish fresh water fish species from the southern part of the country. The effectiveness of DNA barcoding technology has been validated for various animal groups and most investigated species (>94%) possess distinct barcode arrays, with low intraspecies variation and high divergences from closely allied taxa [16,19]. This clear sequence divergence between species, coupled with sequence conservation within species confirmed the barcode CO1 sequence as highly variable and specific.
The branch length within species is more clustered together than the branch length between species which are deeper. This confirms the findings of Meyer et al. [28] that branch length between species tends to be much deeper than between conspecific individuals leading to a gap in the distribution of the pairwise distance between conspecific individuals and between species that have been referred to the barcoding gap. The phylogenetic tree clearly separates the three tilapia species into three clustered groups based on their genetic distance. Ward et al. [19] in their study of Australian fish species demonstrated the efficacy of phylogenetic tree in distinguishing species by showing 14 different clusters from 14 species of flathead. Mohanty et al. [26] in their study of cultivable carp clearly distinguished different carp species into different clustered group. The result showed all individuals of a species clustered together.
The close similarities between the mouth brooders: Sarotherodon species and Oreochromis niloticus as shown in the phylogenetic tree is in concordance with the hypothesis of Trewavas et al. [29] but in disagreement with the hypothesis of Peter et al. [30]. Similar studies using enzyme loci showed clustering patterns according to their genera, i.e., Oreochromis, Sarotherodon and Tilapia [31,32]. The study of Saad et al. [33], using microsatellite DNA markers showed a clustering pattern of Tilapia species according to their generic group. The genetic distance between the three generic groups in this study is less than the values obtained in the study of Gu et al. [34]. This could be attributed to the common source of the collection sites in this study. The efficacy of DNA barcode region (CO1), in the identification cryptic species was validated with the identification of Sarotherodon galilaeus boulongeri, a sub-species of Sarotherodon galilaeus which has never been reported in Nigeria.
Conclusion
Mitochondrial CO1 gene is an ideal region for fish species identification. DNA sequence from this gene will improve and aid the identification of fish species even where morphological characteristics needed for the identification of a particular species is not available as a result of predation or smaller samples that cannot be identified using morphological characteristic such as larvae.
Acknowledgement
The authors thank the Coordinator, Chevron Biotechnology Centre, Modibbo Adama University of Technology Yola, for granting us access to the Molecular Biology Laboratory, to carry out this research.
References
- Nagl S, Tichy H, Mayer WE, Samonte IE, McAndrew BJ, et al. (2001) Classification and phylogenetic relationships of African tilapiine fishes inferred from mitochondrial DNA sequences. Mol Phylogenet Evol 20: 361-374.
- Aad G, Abbott B, Abdallah J, Abdinov O, Aben R, et al. (2016) Measurement of the transverse momentum and distributions of Drell-Yan lepton pairs in proton-proton collisions at TeV with the ATLAS detector. Eur Phys J C Part Fields 76: 291.
- Trewavas E (1983) Tilapiine fishes of the genera Sarotherodon, Oreochromis and Danakilia. British Museum. British Museum, London.Â
- Wohlfarth GW, Hulata GI (1983) Applied genetics of tilapias. ICLARM Stud Rev 6: 1-26.
- van der Bank FH, Grant WS, Ferriera JT (1989) Electrotrophoretically detectable genetic data for fifteen Southern African cichlids. J Fish Biol 34: 465-483.
- van der Bank FH (1994) Molecular systematics of 15 old world cichlid species using allozyme data. Biochem Syst Ecol 22: 791-801.
- Rao C, Majumdar KC (1998) Multivariate map representation of phylogenetic relationships: Application to tilapiine fish. J Fish Biol 52: 1199-1217.
- Welcomme RL (1967) Observations on the biology of the introduced species of tilapia in Lake Victoria. Rev Zool Bot Afr 86: 249-279.
- Agnèse J, Ade´po-Goure`ne B, Owino J, Pouyard, L Aman R (1999) Genetic characterization of a pure relict population of Oreochromis esculentus, an endangered tilapia. J Fish Biol 54: 1119-1123.
- McAndrew BJ, Majumdar KC (1983) Tilapiine stock identification using electrophoretic markers. Aquaculture 30: 249-261.
- Thompson PJ (1984) Hybridization of indigenous Oreochromis species in Zimbabwe and production of all-male offspring. Aquaculture in South Africa 1984 joint symposium by the CSIR and SAAU, South Africa.
- Beattie IH (1986) Summary of a talk on ‘the development of pure strains. Department of National Parks and Wild Life Management, Harare, Zimbabwe.
- Bell-Cross G, Minshull JL (1988) The fishes of Zimbabwe. National Museums, Zimbabwe.
- Gregg RE, Howard JE, Shonhiwa F (1998) Introgressive hybridization of tilapias in Zimbabwe. J Fish Biol 52: 1-10.
- Liu ZJ, Cordes JF (2004) DNA marker technologies and their applications in aquaculture genetics. Aquaculture 238: 1-37.
- Hajibabaei M, Singer GAC, Hebert PDN, Hickey DA (2007) DNA barcoding: How it complements taxonomy, molecular phylogenetics and population genetics. Trends Genet 23: 167-172.
- Akinwande AA, Sogbesan AO, Moody FO, Ugwumba AA (2007) Piscicidal potential of mesocarp of Neem plant (Azadirachta indica L.) fruit on hybrid, "Heteroclarias". J Environ Biol 28: 533-536.
- Idodo-Umeh G (2003) Freshwater fishes of Nigeria (taxonomy, ecological notes, diet and utilization). Idodo Umeh Publishers Limited.
- Ward RD, Zemlak TS, Innes BH, Last PR, Hebert PDN (2005) DNA barcoding Australia’s fish species. Philosophical Transactions of the Royal Society of London Series B-Biological Sciences 360: 1847-1857.
- Hall TA (1999) BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl Acids Symp Ser 41: 95-98.
- Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, et al. (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23: 2947-2948.
- Kumar S, Tamura K, Nei M (2004) MEGA3: Integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5: 150-163.
- Saitou N, Nei M (1987) The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol Biol Evol 4: 406-425.
- Wu L, Yang J (2012) Identifications of captive and wild tilapia species existing in Hawaii by mitochondrial DNA control region sequence. PLoS ONE 7: e51731
- Hubert N, Hanner R, Holm E, Mandrak NE, Taylor E, et al. (2008) Identifying Canadian freshwater fishes through DNA barcodes. PLoS ONE 3: e2490.
- Mohanty M, Jayasankar P, Sahoo L, Das P (2015) A comparative study of COI and 16 S rRNA genes for DNA barcoding of cultivable carps in India. Mitochondrial DNA 26: 79-87.
- Nwani CD, Becker S, Braid HE, Udec EF, Okogwu OI, et al. (2011) DNA barcoding discriminates freshwater fishes from Southeastern Nigeria and provides river system - Level phylogeographic resolution within some species. Mitochondrial DNA 22: 43-51.
- Meyer CP, Paulay G (2005) DNA barcoding: Error rates based on comprehensive sampling. PLoS Biol 3: e422.
- Trewavas E (1980) Tilapia and sarotherodon. Buntbarsche Bulletin 81: 1-6.
- Peters HM, Berns S (1982) Die Maulbrutpflege der Cichliden. Untersuchungen zur Evolution eines Verhaltensmusters Z. Zool Syst Evolutionforsch 20: 18-52.
- McAndrew BJ, Majumdar KC (1984) Evolutionary relationships within three Tilapiine genera (Pisces: Cichlidae). Zoological Journal of the Linnean Society 80: 421-435.
- Pouyaud L, Agnèse JF (1995) Phylogenetic relationships between 21 species of three Tilapiine genera Tilapia, Sarotherodon and Oreochromis using allozyme data. J Fish Biol 47: 26-38.
- Saad YM, Rashed MA, Atta AH, Ahmed NE (2013) The efficiency of microsatellite DNA markers for estimating genetic polymorphism in some Tilapia species. Life Sci J 10: 2230-2234.
- Gu DE, Mu XD, Xu M, Luo D, Wei H, et al. (2016) Identification of wild tilapia species in the main rivers of south China using mitochondrial control region sequence and morphology. Biochm Sys Ecol 65: 100-107.
Citation: Sogbesan OA, Sanda MK, Ja’afar JN, Adedeji HA (2017) DNA Barcoding of Tilapia Species (Pisces: Cichlidae) from North-Eastern Nigeria. J Biotechnol Biomater 7: 277. DOI: 10.4172/2155-952X.1000277
Copyright: © 2017 Sogbesan OA, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited
Select your language of interest to view the total content in your interested language
Share This Article
Recommended Journals
Open Access Journals
Article Tools
Article Usage
- Total views: 7566
- [From(publication date): 0-2017 - Nov 08, 2025]
- Breakdown by view type
- HTML page views: 6507
- PDF downloads: 1059