Female New-Born with Undifferentiated Sarcoma Defined by Bcor-Ccnb3 Fusion Transcript
Received: 15-Jul-2015 / Accepted Date: 03-Aug-2015 / Published Date: 05-Aug-2015 DOI: 10.4172/2161-0681.1000244
Abstract
A female new-born presented with a sacrococcygeal mass with spinal cord compression. A preliminary histologic diagnosis determined a small round blue cell tumor and immunohistochemical results discarded neuroblastoma, rhabdomyosarcoma, rhabdoid tumor, Ewing or peripheral neuroectodermal tumor (PNET). The results obtained by SNPa showed a chromosome Xp11.4 deletion of 0.9 Mb, where the BCOR gene is located. RT-PCR did not detect the ETV6-NTRK3 or EWSR1-FLI1 fusion, but did reveal the presence of the BCOR-CCNB3 fusion transcript, recently reported in some undifferentiated sarcomas, establishing the diagnosis of “Ewing-like” sarcoma. Analysis of CCNB3 expression by immunohistochemistry showed nuclear overexpression in 100% of tumor cells. The patient received aggressive chemotherapy which reduced the size of the tumor, but ultimately the patient died of liver failure secondary to infection with adenovirus 5 months after diagnosis. While the literature reports the predominance of these tumors in male children or young adults, we present the first case in a female new-born. This finding supports the need for further studies to look for the presence of the BCOR-CCNB3 fusion transcript in younger patients, to achieve an effective sub-classification within undifferentiated sarcomas.
Keywords: BCOR-CCNB3 sarcoma; Undifferentiated sarcoma; “Ewing-like” sarcoma; Small round cell tumor
320138Introduction
Small round cell sarcomas (SRCS) are the most aggressive tumors of children and young adults. They constitute a heterogeneous group of neoplasms that can be divided into different subtypes based on clinical, morphological, immunophenotypic and molecular features [1]. A subset of SRCS, however, lack specific characteristics and are included in a group of undifferentiated sarcomas [2,3]. Some of these tumors often resemble Ewing sarcoma morphologically, but lack its classical immunoprofile and genetic features, and are known as “Ewing-like” sarcomas.
Using next-generation sequencing techniques Pierron et al. identified a novel BCOR-CCNB3 fusion gene resulting from a X-chromosomal paracentric inversion [4]. In undifferentiated sarcomas, exon 15 of BCOR and exon 5 of CCNB3, which are located at Xp11.4 and Xp11.22 respectively, are linked. This fusion gene has been identified mainly in male adolescents and young adults SRCS of bone. This finding was confirmed in later studies where BCOR-CCNB3 fusion was associated with “Ewing-like” sarcomas [5-7].
Here, we describe the first case of a female new-born with undifferentiated sarcoma patient defined by the presence of the BCOR-CCNB3 fusion transcript.
Case Report
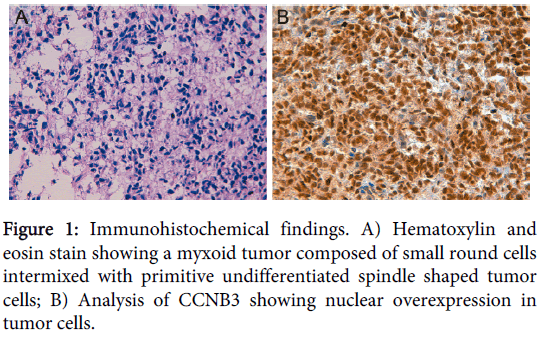
An 8-day-old Caucasian girl with history of prematurity presented with a sacrococcygeal mass with spinal cord compression, progressive loss of the right plantar reflex and the mobility in lower extremities, and anal sphincter dilatation. Electromyogram showed an axonal lesion in L3-S2 and magnetic resonance imaging suggested neuroblastic tumor. A Metaiodobenzylguanidine scan was negative and no urinary catecholamines were detected. Histopathological analysis showed a neoplasm with a diffuse growth pattern composed of small round cells intermixed with primitive undifferentiated spindle shaped tumor cells in a myxoid background (Figure 1). Atypical mitoses were evident. The patient was treated with a neuroblastoma protocol (Etoposide-Carboplatin) obtaining a 30% tumor reduction after the first cycle. Immunohistochemical stains were performed on different formalin fixed paraffin-embedded (FFPE) sections to categorize the unclassified sarcoma, including NB84, Tyrosine Hydroxylase, NCAM (CD56) and synaptophysin immunomarkers for neuroblastoma; myogenin, myoD-1, actin and desmin for rhabdomyosarcoma; WT1 for Wilms tumor and desmoplastic small round cell tumor, CD99 for Ewing/PNET; INI1 for rhabdoid tumor and vimentin as marker of mesenchymal cell. Immunohistochemistry showed negative staining for all tumor markers except for INI1 (negative in rhabdoid tumor) and vimentin. From these results, a final diagnosis of primitive myxoid mesenchymal undifferentiated sarcoma was established. After rapid tumor progression the therapy was changed to Ifosfamide-Vincristine-Anthracycline (IVA). As the tumor was non-resectable, the patient received aggressive chemotherapy for high-grade sarcomas with Cisplatin-Adriamycin and caffeine, with the first intra-arterial cycle resulting in tumor size reduction and abundant tumor necrosis, as well as improvement of leg paresis. Ultimately, the patient died of liver failure secondary to infection with adenovirus 5 months after diagnosis. Necropsy revealed that tumor necrosis was close to 100%.
Single Nucleotide Polymorphism array (SNPa) analysis was performed to determine the genomic profile as described elsewhere [8]. The SNPa platform used was HumanCytoSNP-12 DNA Analysis BeadChip (299,140 markers) from Illumina Inc (San Diego, CA). DNA was extracted from fresh tissue using a salting out procedure [9]. DNA amplification, tagging, and hybridization to Illumina chip were performed according to the manufacturer's protocol (http://www.illumina.com). Data were analyzed using GenomeStudio Genotyping and KaryoStudio software with standard settings. For exclusion of constitutional copy number polymorphisms, the Database of Genomic Variants was used (http://projects.tcaq.ca/variation). Genomic position annotations were based on the hg19 build of the human genome sequence (http://genome.ucsc.edu/).
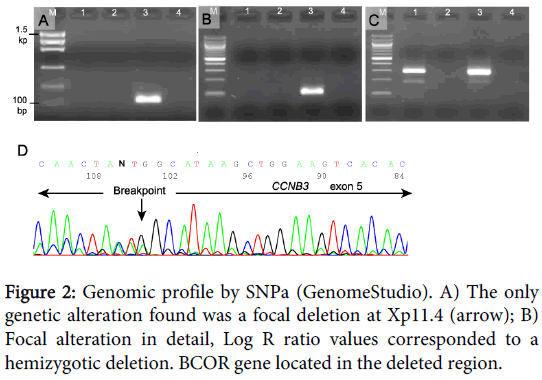
The results obtained by SNPa showed a genomic profile lacking any Segmental Chromosome Aberration (SCA) or Numerical Chromosome Aberration (NCA) but with a focal loss of small chromosomal region (FSCA) in Xp11.4 of 0.9 Mb approximately, involving the BCOR gene (Figure 2).
In an attempt to discard the ETV6-NTRK3 and EWSR1-FLI1 fusion transcript (congenital fibrosarcoma and Ewing sarcoma respectively), and to look for the BCOR-CCNB3 fusion, reverse-transcription PCR (RT-PCR) was performed using specific primers previously described [4,10,11]. Three 20-µm sections were cut from the representative FFPE tissue, deparaffinized with xylene, rinsed in ethanol, and digested by proteinase K. Total RNA was extracted using the RecoverAll Total Nucleic Acid (Ambion Ltd. Cambridgeshire, UK). RNA (1 µg) was then reverse-transcribed to cDNA using a high-capacity cDNA archive kit (Applied Biosystems, Foster City, CA). PCR amplification was performed using AmpliTaq Gold 360 Master Mix (Applied Biosystems). The reaction products were subjected to electrophoresis in 2% agarose gel and visualized by RedSafe staining (iNtRON Biotechnology). The assay was considered satisfactory for interpretation when the internal ß2-microglobulin control 136-bp PCR product was present, the external fusion-gene positive control was positive, and the water control showed no amplified product.
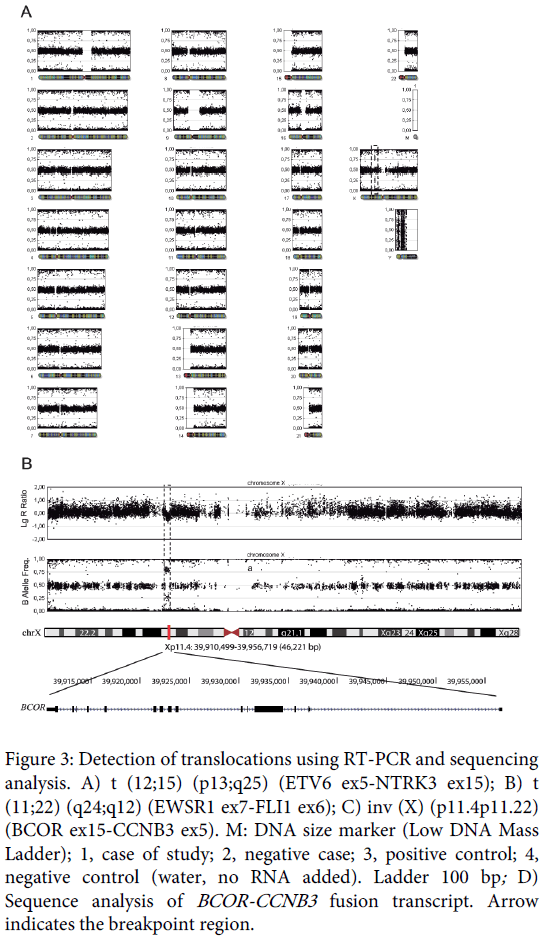
Only the BCOR-CCNB3 fusion was detected in the samples (Figure 3) and immunohistochemical analysis of CCNB3 revealed nuclear overexpression in 100% of tumor cells (Figure 1).
Figure 3: Detection of translocations using RT-PCR and sequencing analysis. A) t (12;15) (p13;q25) (ETV6 ex5-NTRK3 ex15); B) t (11;22) (q24;q12) (EWSR1 ex7-FLI1 ex6); C) inv (X) (p11.4p11.22) (BCOR ex15-CCNB3 ex5). M: DNA size marker (Low DNA Mass Ladder); 1, case of study; 2, negative case; 3, positive control; 4, negative control (water, no RNA added). Ladder 100 bp; D) Sequence analysis of BCOR-CCNB3 fusion transcript. Arrow indicates the breakpoint region.
Sequencing of the RT-PCR products was performed using the Big-DyeTerminator cycle-sequencing kit on an ABI Prism 310 automated sequencer (Applied Biosystems). The fusion of BCOR exon 15 to CCNB3 exon 5 resulting in inv (X) (p.11.4p11.22) was confirmed (Figure 3).
Discussion
The emergence of next-generation sequencing technologies in recent years has revolutionized diagnosis in oncology. In sarcomas, the discovery of chromosomal rearrangements that produce previously unrecognized oncogenic fusion genes, has led to a better subclassification of round cell sarcomas [12].
Recently, a novel X-chromosomal paracentric inversion, resulting in a BCOR-CCNB3 fusion, has been described in a series of undifferentiated sarcomas [4]. The emergence of this fusion gene has lead scientists to screen for additional cases of the BCOR-CCNB3 fusion in translocation-negative undifferentiated unclassified sarcomas to further characterize their pathology and clinical course. Consequently various studies have reported similar results: undifferentiated sarcomas with the BCOR-CCNB3 fusion transcript presented remarkable clinical and pathological similarities with Ewing sarcoma tumors, most of them located in bone and occurring in adolescent or young adults, preferentially in males. This new group of tumors, originally called “Ewing-like” sarcomas, does not share the EWSR1-ETS expression signature, or the classical abnormalities present in Ewing sarcomas, and have a less consistent CD99 expression. Moreover, the most frequent copy-number abnormalities of Ewing sarcoma, particularly the gain at chromosome 8 and 12, the gain of 1q or the loss of 16q are not recurrent characteristics of BCOR-CCNB3-positive cases [5-7]. In our case, SNPa detected no alterations apart from the FSCA in Xp11.4, and RT-PCR results showed no expression of the EWSR1-FLI1 and ETV6-NTRK3 transcripts but did detect the presence of the BCOR-CCNB3 fusion transcript, thus we confirm previous genetic findings reported. CD99 immuno-staining was also negative. In contrast, the clinical characteristics of our female new-born patient were strongly different from the clinical features so far described for undifferentiated sarcomas with the BCOR-CCNB3 fusion transcript. Moreover, the tumor showed a mixture of small round cells and spindle cells in a myxoid matrix, which are not the usual histomorphologic features of conventional Ewing sarcoma, suggesting the diagnosis of undifferentiated myxoid sarcoma of new-born.
The pathogenic role of BCOR-CCNB3 fusions in sarcomas remains unclear. The BCOR gene encodes a nuclear corepressor of BCL6 and regulates mesenchymal stem cell function by epigenetic mechanisms [13] while CCNB3 encodes a meiotic cyclin that is expressed in the leptotene and zygotene phases of spermatogenesis [14]. The expression of the BCOR-CCNB3 fusion oncoprotein can be detected by CCNB3 immunostaining, which provide a simple and reliable immunohistochemical marker to diagnose this subset of tumors [4]. Given this expression pattern, it would be important to clarify the specific role of the different domains of BCOR as a key oncogenic event.
On the basis of previous reports, as well as on our data, we emphasize that the clinical and pathological characteristics of this tumor do not completely fall within “Ewing-like” sarcomas; BCOR-CCNB3 undifferentiated sarcoma has been rarely described in female patients and infantile sarcoma or sarcoma developed since intra-uterine stage is an outstanding feature of this case from another reported BCOR-CCNB3 fusion tumors so may be considered a type of sarcoma unrelated to Ewing tumor [5,12,15].
Acknowledgements
The authors thank Olivier Delattre, from the Institute Curie (Unité de Génétique Somatique-Centre Hospitalier, Paris, France), for providing control material from BCOR-CCNB3 cases. The authors also thank David Harrison for English correction. This work was supported by FIS (contract PI14/01008) and RTICC (contract RD12/36/20), Instituto Carlos III, Madrid & ERDF, Spain.
References
- Lessnick SL, Dei Tos AP, Sorensen PH, Dileo P, Baker LH, et al. (2009) Small round cell sarcomas. SeminOncol 36:338-346.
- Fletcher CD (2008) Undifferentiated sarcomas: what to do? And does it matter? A surgical pathology perspective. UltrastructPathol 32:31-36.
- Alaggio R, Bisogno G, Rosato A, Ninfo V, Coffin CM (2009) Undifferentiated sarcoma: does it exist? A clinicopathologic study of 7 pediatric cases and review of literature. Hum Pathol 40:1600-1610.
- Pierron G, Tirode F, Lucchesi C, Reynaud S, Ballet S, et al. (2012) A new subtype of bone sarcoma defined by BCOR-CCNB3 gene fusion. Nat Genet 44:461-466.
- Puls F, Niblett A, Marland G, Gaston CL, Douis H, et al. (2014) BCOR-CCNB3 (Ewing-like) sarcoma: a clinicopathologic analysis of 10 cases, in comparison with conventional Ewing sarcoma. Am J SurgPathol 38:1307-1318.
- Cohen-Gogo S, Cellier C, Coindre JM, Mosseri V, Pierron G, et al. (2014) Ewing-like sarcomas with BCOR-CCNB3 fusion transcript: a clinical, radiological and pathological retrospective study from the SocieteFrancaise des Cancers de L'Enfant. Pediatr Blood Cancer 61:2191-2198.
- Peters TL, Kumar V, Polikepahad S, Lin FY, Sarabia SF, et al. (2015) BCOR-CCNB3 fusions are frequent in undifferentiated sarcomas of male children. Mod Pathol 28:575-586.
- Berbegall AP, Villamon E, Tadeo I, Martinsson T, Canete A, et al. (2014) Neuroblastoma after childhood: prognostic relevance of segmental chromosome aberrations, ATRX protein status, and immune cell infiltration. Neoplasia 16:471-480.
- Miller SA, Dykes DD, Polesky HF (1988) A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res 16:1215.
- Terrier-Lacombe MJ, Guillou L, Chibon F, Gallagher G, Benhattar J, et al. (2009) Superficial primitive Ewing's sarcoma: a clinicopathologic and molecular cytogenetic analysis of 14 cases. Mod Pathol 22:87-94.
- Alaggio R, Barisani D, Ninfo V, Rosolen A, Coffin CM (2008) Morphologic Overlap between Infantile Myofibromatosis and Infantile Fibrosarcoma: A Pitfall in Diagnosis. PediatrDevPathol 11:355-362.
- Marino-Enriquez A, Fletcher CD (2014) Round cell sarcomas - biologically important refinements in subclassification. Int J Biochem Cell Biol 53:493-504.
- Fan Z, Yamaza T, Lee JS, Yu J, Wang S, et al. (2009) BCOR regulates mesenchymal stem cell function by epigenetic mechanisms. Nat Cell Biol 11:1002-1009.
- Nguyen TB, Manova K, Capodieci P, Lindon C, Bottega S, et al. (2002) Characterization and expression of mammalian cyclin b3, a prepachytene meiotic cyclin. J BiolChem 277:41960-41969.
- Antonescu C (2014) Round cell sarcomas beyond Ewing: emerging entities. Histopathology 64:26-37.
Citation: Martín-Vañó S, Berbegall AP, Navarro L, Blanquer-Maceiras M, Beléndez-Bieler, et al. (2015) Female New-Born with Undifferentiated Sarcoma Defined by Bcor-Ccnb3 Fusion Transcript. J Clin Exp Pathol 5:244. DOI: 10.4172/2161-0681.1000244
Copyright: © 2015 Martín-Vañó S et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Select your language of interest to view the total content in your interested language
Share This Article
Recommended Journals
Open Access Journals
Article Tools
Article Usage
- Total views: 16186
- [From(publication date): 10-2015 - Aug 30, 2025]
- Breakdown by view type
- HTML page views: 11522
- PDF downloads: 4664